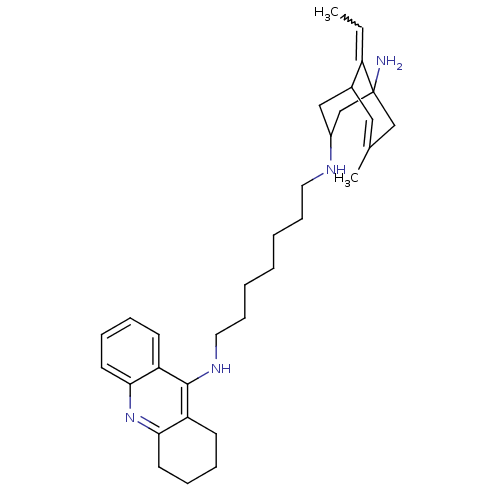
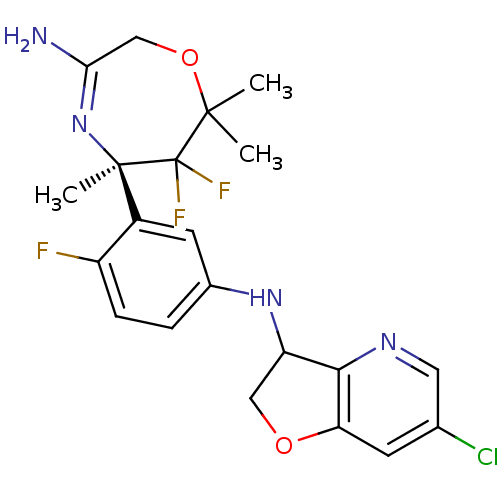
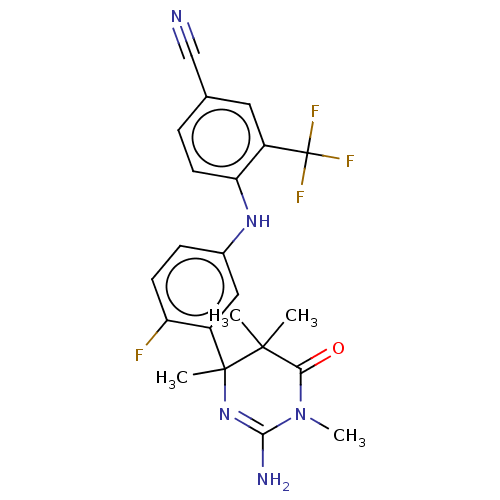
Affinity DataKi: 0.0260nM ΔG°: -60.4kJ/molepH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by E llman. Inhibition of enzyme activity was measured over a substrate c...More data for this Ligand-Target Pair
Affinity DataKi: 6.40nM ΔG°: -46.8kJ/molepH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by E llman. Inhibition of enzyme activity was measured over a substrate c...More data for this Ligand-Target Pair
Affinity DataKi: 7nM ΔG°: -46.5kJ/molepH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by E llman. Inhibition of enzyme activity was measured over a substrate c...More data for this Ligand-Target Pair
Affinity DataKi: 15.7nM ΔG°: -44.5kJ/molepH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by E llman. Inhibition of enzyme activity was measured over a substrate c...More data for this Ligand-Target Pair
Affinity DataKi: 16.5nM ΔG°: -44.4kJ/molepH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by E llman. Inhibition of enzyme activity was measured over a substrate c...More data for this Ligand-Target Pair
Affinity DataKi: 19.5nM ΔG°: -44.0kJ/molepH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by E llman. Inhibition of enzyme activity was measured over a substrate c...More data for this Ligand-Target Pair
Affinity DataKi: 20.8nM ΔG°: -43.8kJ/molepH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by E llman. Inhibition of enzyme activity was measured over a substrate c...More data for this Ligand-Target Pair
Affinity DataKi: 30.8nM ΔG°: -42.9kJ/molepH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by E llman. Inhibition of enzyme activity was measured over a substrate c...More data for this Ligand-Target Pair
Affinity DataKi: 47nM ΔG°: -41.8kJ/molepH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by E llman. Inhibition of enzyme activity was measured over a substrate c...More data for this Ligand-Target Pair
Affinity DataKi: 120nM ΔG°: -39.5kJ/molepH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by E llman. Inhibition of enzyme activity was measured over a substrate c...More data for this Ligand-Target Pair
Affinity DataKi: 137nM ΔG°: -39.2kJ/molepH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by E llman. Inhibition of enzyme activity was measured over a substrate c...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+3nM ΔG°: >-34.2kJ/molepH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by E llman. Inhibition of enzyme activity was measured over a substrate c...More data for this Ligand-Target Pair
Affinity DataIC50: 0.25nMAssay Description:Inhibition of rat MAGLMore data for this Ligand-Target Pair
Affinity DataIC50: 1.40nMAssay Description:Inhibition of recombinant C-terminal His6-tagged human MAGL expressed in Escherichia coli using 2-arachidonoyl-[3H]-glycerol as substrate incubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 3.10nMAssay Description:Displacement of [3H]PI-2620 from Tau in human brain homogenate incubated for 60 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.5nMAssay Description:Displacement of [3H]PI-2620 from Tau in human brain homogenate incubated for 60 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4.60nMAssay Description:Inhibition of recombinant C-terminal His6-tagged human MAGL expressed in Escherichia coli using 2-arachidonoyl-[3H]-glycerol as substrate incubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 4.60nMAssay Description:Inhibition of recombinant C-terminal His6-tagged human MAGL expressed in Escherichia coli using 2-arachidonoyl-[3H]-glycerol as substrate incubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 4.90nMAssay Description:Displacement of [3H]PI-2620 from Tau in human brain homogenate incubated for 60 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5nMpH: 4.5Assay Description:The FRET assay was performed essentially as described in Gruninger-Leitch et al., Journal of Biological Chemistry (2002) 277(7) 4687-93 (Substrate an...More data for this Ligand-Target Pair
Affinity DataIC50: 7.80nMAssay Description:Displacement of [18F]-FEH from MAO-A in mouse brain homogenate incubated for 60 mins by imaging analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:The assay uses the principle of inhibition of human TMEM27 cleavage by endogenous cellular BACE2 in the Ins1e rat cell line and shedding from the cel...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Inhibition of MAGL in rat brain membranes preincubated for 20 mins followed by fluorophosphonate-rhodamine addition measured after 30 mins by competi...More data for this Ligand-Target Pair
Affinity DataIC50: 11nMpH: 4.5Assay Description:The FRET assay was performed essentially as described in Gruninger-Leitch et al., Journal of Biological Chemistry (2002) 277(7) 4687-93 (Substrate an...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMT: 2°CAssay Description:BACE2 enzyme ectodomain (derived from plasmid pET17b-T7-hu proBACE2) was prepared as described in Ostermann et al., Crystal Structure of Human BACE2 ...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:Inhibition of recombinant C-terminal His6-tagged human MAGL expressed in Escherichia coli using 2-arachidonoyl-[3H]-glycerol as substrate incubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 14nMT: 2°CAssay Description:BACE2 enzyme ectodomain (derived from plasmid pET17b-T7-hu proBACE2) was prepared as described in Ostermann et al., Crystal Structure of Human BACE2 ...More data for this Ligand-Target Pair
Affinity DataIC50: 16nMAssay Description:Displacement of [3H]PI-2620 from Tau in human brain homogenate incubated for 60 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 17nMAssay Description:DNA of the human APP wt gene (APP695) were used to assess the potency of the compounds in a cellular assay. The cells were seeded in 96-well microtit...More data for this Ligand-Target Pair
Affinity DataIC50: 18nMAssay Description:The assay uses the principle of inhibition of human TMEM27 cleavage by endogenous cellular BACE2 in the Ins1e rat cell line and shedding from the cel...More data for this Ligand-Target Pair
Affinity DataIC50: 18nMAssay Description:Displacement of [3H]PI-2620 from Tau in human brain homogenate incubated for 60 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 21nMT: 2°CAssay Description:BACE2 enzyme ectodomain (derived from plasmid pET17b-T7-hu proBACE2) was prepared as described in Ostermann et al., Crystal Structure of Human BACE2 ...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Homo sapiens (Human))
University Of Siena
Curated by ChEMBL
University Of Siena
Curated by ChEMBL
Affinity DataIC50: 22nMAssay Description:Inhibition of human ERG expressed in rat after 2 to 3 days by patch clamp assayMore data for this Ligand-Target Pair
Affinity DataIC50: 23nMAssay Description:Inhibition of recombinant C-terminal His6-tagged human MAGL expressed in Escherichia coli using 2-arachidonoyl-[3H]-glycerol as substrate incubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 24nMpH: 4.5Assay Description:The FRET assay was performed essentially as described in Gruninger-Leitch et al., Journal of Biological Chemistry (2002) 277(7) 4687-93 (Substrate an...More data for this Ligand-Target Pair
Affinity DataIC50: 28nMpH: 4.5Assay Description:The FRET assay was performed essentially as described in Gruninger-Leitch et al., Journal of Biological Chemistry (2002) 277(7) 4687-93 (Substrate an...More data for this Ligand-Target Pair
Affinity DataIC50: 30nMpH: 4.5Assay Description:The FRET assay was performed essentially as described in Gruninger-Leitch et al., Journal of Biological Chemistry (2002) 277(7) 4687-93 (Substrate an...More data for this Ligand-Target Pair
Affinity DataIC50: 30nMpH: 4.5Assay Description:The FRET assay was performed essentially as described in Gruninger-Leitch et al., Journal of Biological Chemistry (2002) 277(7) 4687-93 (Substrate an...More data for this Ligand-Target Pair
Affinity DataIC50: 31nMAssay Description:Inhibition of recombinant C-terminal His6-tagged human MAGL expressed in Escherichia coli using 2-arachidonoyl-[3H]-glycerol as substrate incubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 35nMT: 2°CAssay Description:BACE2 enzyme ectodomain (derived from plasmid pET17b-T7-hu proBACE2) was prepared as described in Ostermann et al., Crystal Structure of Human BACE2 ...More data for this Ligand-Target Pair
Affinity DataIC50: 35nMpH: 4.5Assay Description:The FRET assay was performed essentially as described in Gruninger-Leitch et al., Journal of Biological Chemistry (2002) 277(7) 4687-93 (Substrate an...More data for this Ligand-Target Pair
Affinity DataIC50: 36nMAssay Description:Displacement of [3H]PI-2620 from Tau in human brain homogenate incubated for 60 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 39nMT: 2°CAssay Description:BACE2 enzyme ectodomain (derived from plasmid pET17b-T7-hu proBACE2) was prepared as described in Ostermann et al., Crystal Structure of Human BACE2 ...More data for this Ligand-Target Pair
Affinity DataIC50: 40nMpH: 4.5Assay Description:The FRET assay was performed essentially as described in Gruninger-Leitch et al., Journal of Biological Chemistry (2002) 277(7) 4687-93 (Substrate an...More data for this Ligand-Target Pair
Affinity DataIC50: 40nMpH: 4.5Assay Description:The FRET assay was performed essentially as described in Gruninger-Leitch et al., Journal of Biological Chemistry (2002) 277(7) 4687-93 (Substrate an...More data for this Ligand-Target Pair
Affinity DataIC50: 44nMAssay Description:DNA of the human APP wt gene (APP695) were used to assess the potency of the compounds in a cellular assay. The cells were seeded in 96-well microtit...More data for this Ligand-Target Pair
Affinity DataIC50: 51nMAssay Description:Displacement of [18F]-FEH from MAO-A in mouse brain homogenate incubated for 60 mins by imaging analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 58nMpH: 4.5Assay Description:The FRET assay was performed essentially as described in Gruninger-Leitch et al., Journal of Biological Chemistry (2002) 277(7) 4687-93 (Substrate an...More data for this Ligand-Target Pair
Affinity DataIC50: 61nMAssay Description:Inhibition of recombinant C-terminal His6-tagged human MAGL expressed in Escherichia coli using 2-arachidonoyl-[3H]-glycerol as substrate incubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 62nMT: 2°CAssay Description:BACE2 enzyme ectodomain (derived from plasmid pET17b-T7-hu proBACE2) was prepared as described in Ostermann et al., Crystal Structure of Human BACE2 ...More data for this Ligand-Target Pair

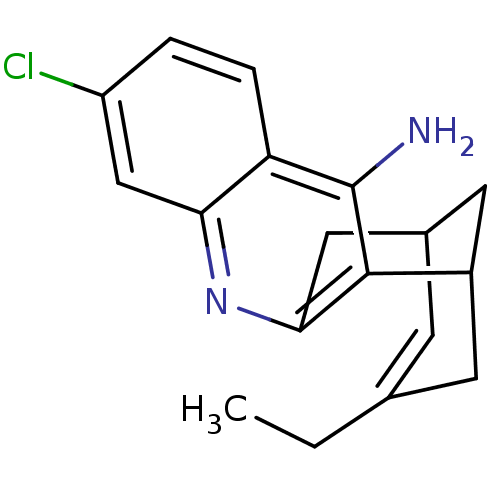
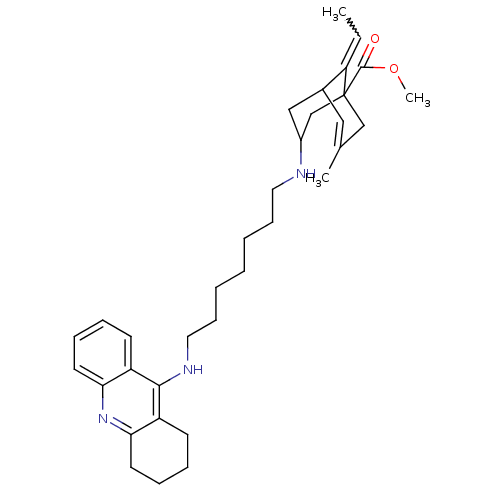
 3D Structure (crystal)
3D Structure (crystal)