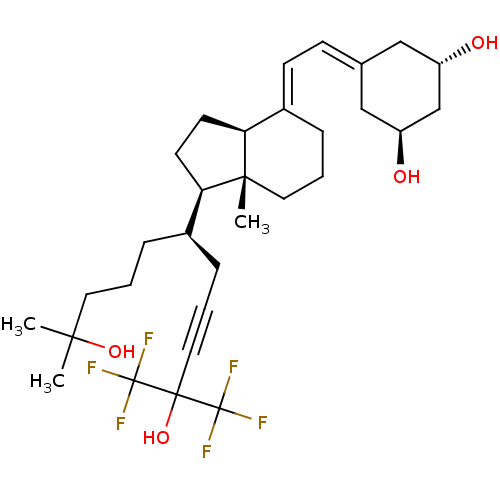
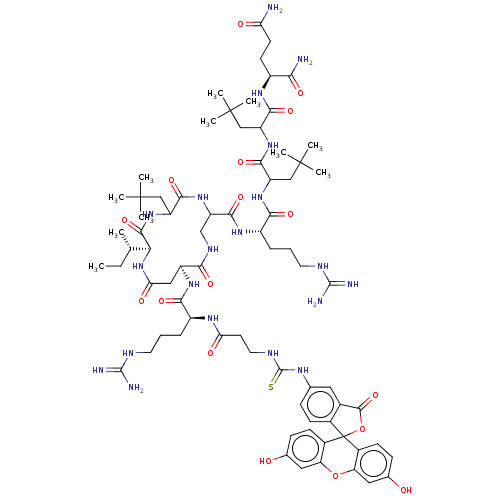
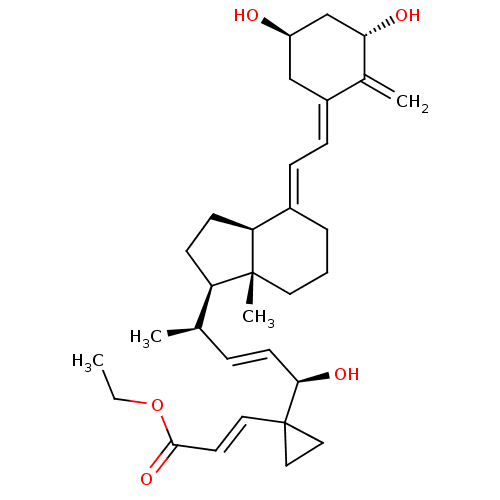
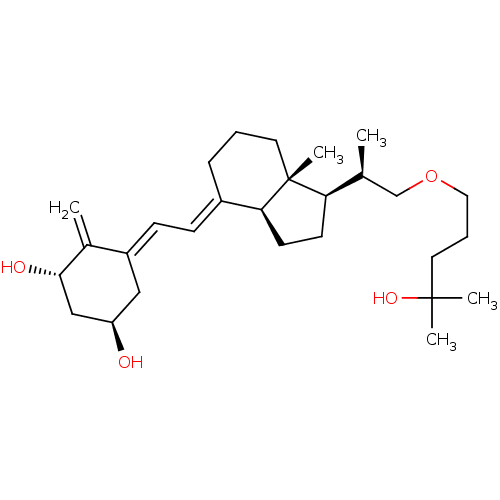
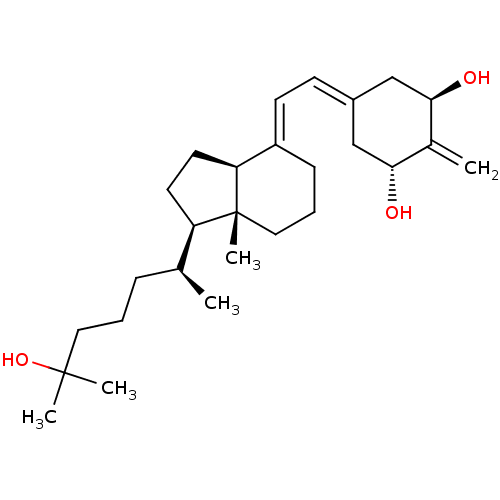
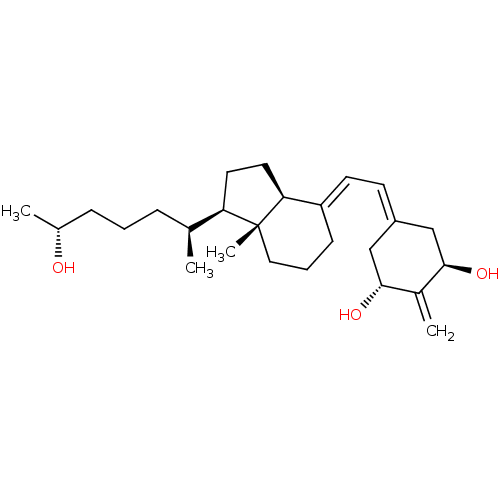
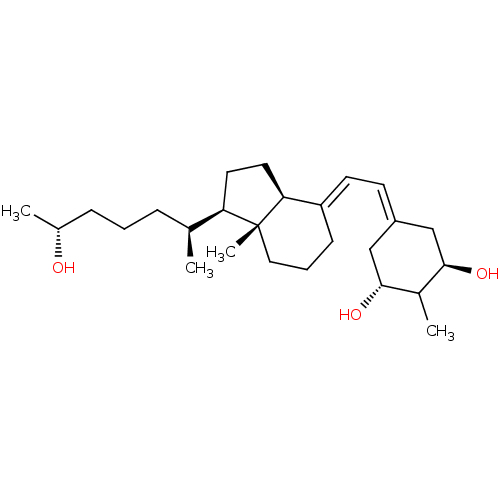
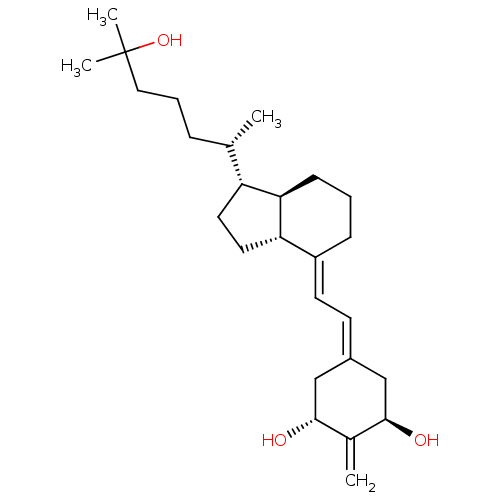
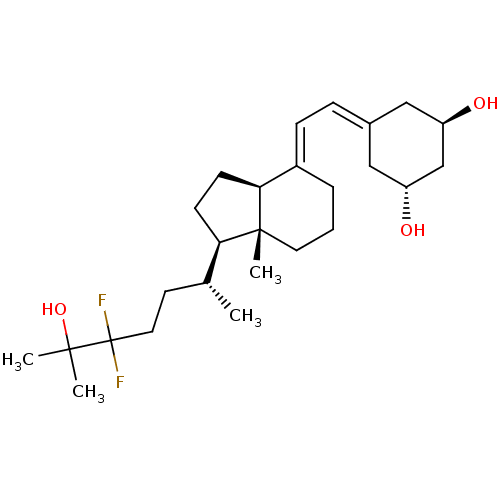
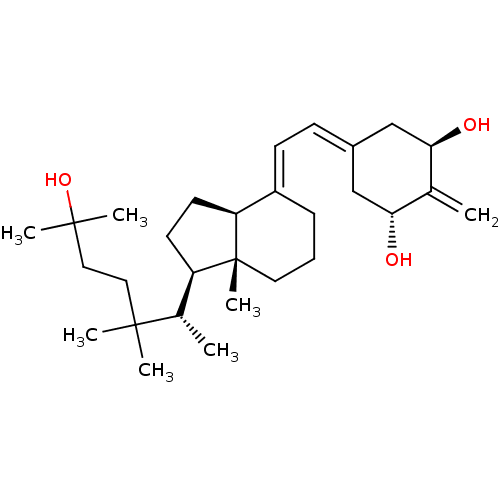
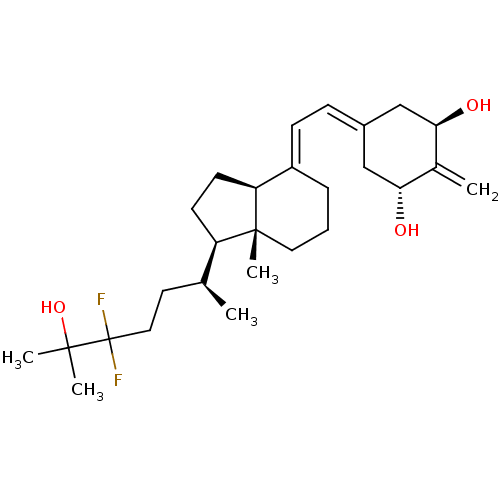
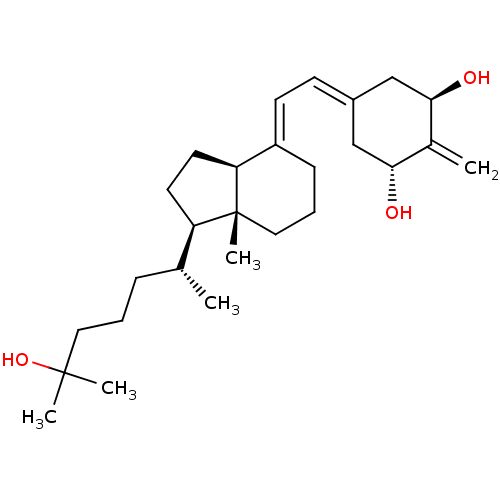
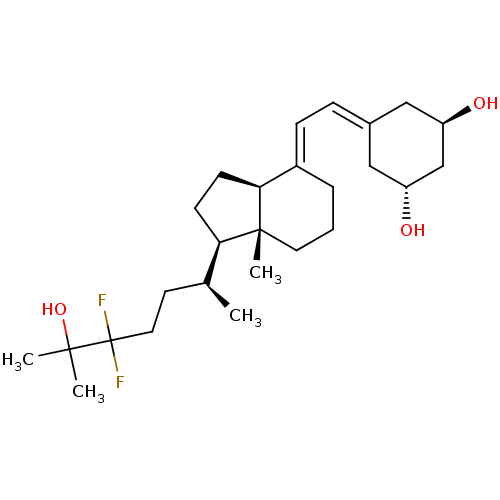
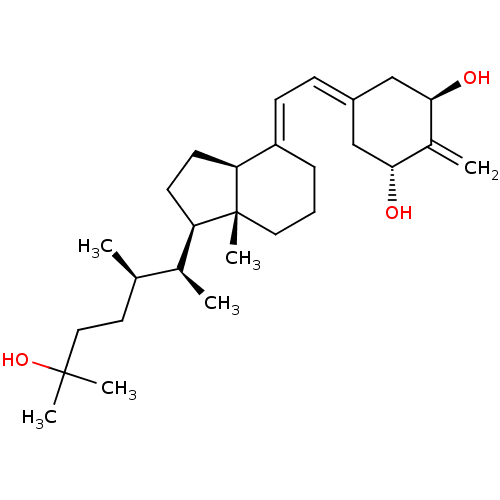
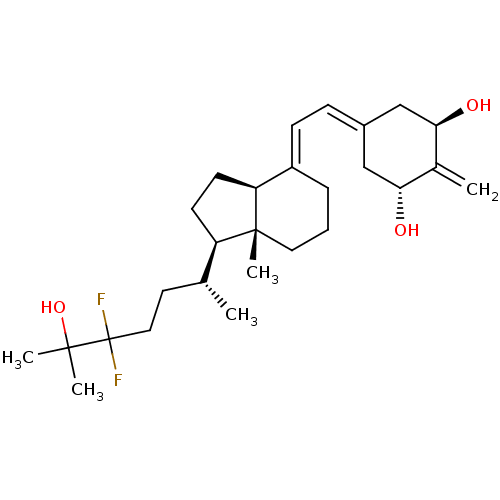
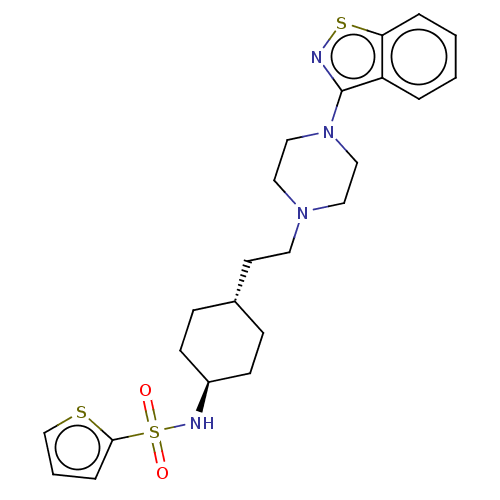
Affinity DataKd: 0.0650nMAssay Description:Binding affinity to VDR receptorMore data for this Ligand-Target Pair
Affinity DataKd: 0.0700nMAssay Description:Displacement of [3H]1-alpha,25-(OH)2D3 from VDR in pig intestinal mucosaMore data for this Ligand-Target Pair
Affinity DataKd: 0.0730nMAssay Description:Binding affinity to VDR receptorMore data for this Ligand-Target Pair
Affinity DataKd: 0.0890nMAssay Description:Binding affinity to VDR receptorMore data for this Ligand-Target Pair
Affinity DataKd: 0.310nMAssay Description:Binding affinity to VDR receptorMore data for this Ligand-Target Pair
Affinity DataKd: 142nMAssay Description:Binding affinity to recombinant human GST-tagged VDR LBD (156 to 453 residues) expressed in Echerichia coli BL21 star (DE3) after 40 mins in presence...More data for this Ligand-Target Pair
Affinity DataKd: 197nMAssay Description:Binding affinity to recombinant human GST-tagged VDR LBD (156 to 453 residues) expressed in Echerichia coli BL21 star (DE3) after 40 mins in presence...More data for this Ligand-Target Pair
Affinity DataKd: 330nMAssay Description:Binding affinity to N-terminal His-tagged human VDR LBD canonical site (118 to 427) by direct isothermal titration calorimetric analysisMore data for this Ligand-Target Pair
TargetVitamin D3 receptor A(Danio rerio)
Institut De G£N£Tique Et De Biologie Mol£Culaire Et Cellulaire (Igbmc)
Curated by ChEMBL
Institut De G£N£Tique Et De Biologie Mol£Culaire Et Cellulaire (Igbmc)
Curated by ChEMBL
Affinity DataKd: 1.20E+3nMAssay Description:Binding affinity to zebrafish VDR LBD assessed as Kd for fluorescein-labeled SRC1 NR2 peptide binding by micro-scale thermophoresis methodMore data for this Ligand-Target Pair
TargetVitamin D3 receptor A(Danio rerio)
Institut De G£N£Tique Et De Biologie Mol£Culaire Et Cellulaire (Igbmc)
Curated by ChEMBL
Institut De G£N£Tique Et De Biologie Mol£Culaire Et Cellulaire (Igbmc)
Curated by ChEMBL
Affinity DataKd: 1.80E+3nMAssay Description:Agonist activity at zebrafish VDR LBD assessed as binding affinity to TAMRA-labeled SRC-1 peptide by fluorescence anisotropy assayMore data for this Ligand-Target Pair
TargetVitamin D3 receptor A(Danio rerio)
Institut De G£N£Tique Et De Biologie Mol£Culaire Et Cellulaire (Igbmc)
Curated by ChEMBL
Institut De G£N£Tique Et De Biologie Mol£Culaire Et Cellulaire (Igbmc)
Curated by ChEMBL
Affinity DataKd: 1.80E+3nMAssay Description:Agonist activity at zebrafish VDR LBD assessed as binding affinity to TAMRA-labeled SRC-1 peptide by fluorescence anisotropy assayMore data for this Ligand-Target Pair
Affinity DataKd: >2.00E+3nMAssay Description:Inhibition of the binding of [3H]C18-Platelet activating factor to human platelet membrane preparationMore data for this Ligand-Target Pair
TargetVitamin D3 receptor A(Danio rerio)
Institut De G£N£Tique Et De Biologie Mol£Culaire Et Cellulaire (Igbmc)
Curated by ChEMBL
Institut De G£N£Tique Et De Biologie Mol£Culaire Et Cellulaire (Igbmc)
Curated by ChEMBL
Affinity DataKd: 3.30E+3nMAssay Description:Agonist activity at zebrafish VDR LBD assessed as binding affinity to TAMRA-labeled SRC-1 peptide by fluorescence anisotropy assayMore data for this Ligand-Target Pair
Affinity DataKd: 9.52E+3nMAssay Description:Binding affinity to N-terminal His-tagged human VDR LBD low-affinity site (118 to 427) by direct isothermal titration calorimetric analysisMore data for this Ligand-Target Pair
Affinity DataKd: >1.00E+4nMAssay Description:Activity at VDR expressed in human HeLa-derived HDLN6 cells assessed as inhibition of transcriptional activity after 18 hrs by luminescence-based luc...More data for this Ligand-Target Pair
Affinity DataKd: >1.00E+4nMAssay Description:Activity at 24 hydroxylase-fused VDR in human HG5LN cells after 18 hrs by luciferase reporter gene assayMore data for this Ligand-Target Pair
Affinity DataKd: 1.89E+4nMAssay Description:Binding affinity to N-terminal His-tagged human VDR LBD (118 to 427) by reverse isothermal titration calorimetric analysisMore data for this Ligand-Target Pair
TargetVitamin D3 receptor A(Danio rerio)
Institut De G£N£Tique Et De Biologie Mol£Culaire Et Cellulaire (Igbmc)
Curated by ChEMBL
Institut De G£N£Tique Et De Biologie Mol£Culaire Et Cellulaire (Igbmc)
Curated by ChEMBL
Affinity DataKd: >3.00E+4nMAssay Description:Binding affinity to zebrafish VDR LBD assessed as Kd for fluorescein-labeled SRC1 NR2 peptide binding by micro-scale thermophoresis methodMore data for this Ligand-Target Pair
Affinity DataKi: <0.00100nMAssay Description:Displacement from vitamin D receptor in chick intestine: 50% displacementMore data for this Ligand-Target Pair
Affinity DataKi: 0.00700nMAssay Description:Activity at rat recombinant full length VDR expressed in rat ROS 17/2.8 cells transfected with 24-hydroxylase gene promoter assessed as transcription...More data for this Ligand-Target Pair
Affinity DataKi: 0.0100nMAssay Description:Activity at rat recombinant full length VDR expressed in rat ROS 17/2.8 cells transfected with 24-hydroxylase gene promoter assessed as transcription...More data for this Ligand-Target Pair
Affinity DataKi: 0.0200nMAssay Description:Activity at rat recombinant full length VDR expressed in rat ROS 17/2.8 cells transfected with 24-hydroxylase gene promoter assessed as transcription...More data for this Ligand-Target Pair
Affinity DataKi: 0.0200nMAssay Description:Displacement of radiolabeled 1alpha,25-(OH)2D3 from rat recombinant full length VDRMore data for this Ligand-Target Pair
Affinity DataKi: 0.0200nMAssay Description:Displacement of [3H]1alpha,25-(OH)2D3 from full-length recombinant rat VDR by scintillation counterMore data for this Ligand-Target Pair
Affinity DataKi: 0.0200nMAssay Description:Displacement of [3H]-1alpha,25(OH)2D3 from recombinant rat VDR after overnight incubation by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 0.0250nMAssay Description:Displacement from vitamin D receptor in chick intestine : 50% displacementMore data for this Ligand-Target Pair
Affinity DataKi: 0.0290nMAssay Description:The experiment is carried out by the method according to Journal of Pharmacology and Experimental Therapeutics 2010, 333(1): 328. With [3H]methyl-spi...More data for this Ligand-Target Pair
Affinity DataKi: 0.0290nMAssay Description:The experiment is carried out by the method according to Journal of Pharmacology and Experimental Therapeutics 2010, 333(1): 328. With [3H]methyl-spi...More data for this Ligand-Target Pair
Affinity DataKi: 0.0300nMAssay Description:Displacement of [3H]1alpha,25-(OH)2D3 from full-length recombinant rat VDR by scintillation counterMore data for this Ligand-Target Pair
Affinity DataKi: 0.0300nMAssay Description:Displacement of [3H]1alpha,25-(OH)2D3 from full-length recombinant rat VDR by scintillation counterMore data for this Ligand-Target Pair
Affinity DataKi: 0.0300nMAssay Description:Displacement of [3H]1alpha,25-(OH)2D3 from full-length recombinant rat VDR by scintillation counterMore data for this Ligand-Target Pair
Affinity DataKi: 0.0350nMAssay Description:The experiment is carried out by the method according to Journal of Pharmacology and Experimental Therapeutics 2010, 333(1): 328. With [3H]methyl-spi...More data for this Ligand-Target Pair
Affinity DataKi: 0.0350nMAssay Description:The experiment is carried out by the method according to Journal of Pharmacology and Experimental Therapeutics 2010, 333(1): 328. With [3H]methyl-spi...More data for this Ligand-Target Pair
Affinity DataKi: 0.0400nMAssay Description:Displacement of [3H]1alpha,25-(OH)2D3 from full-length recombinant rat VDR by scintillation counterMore data for this Ligand-Target Pair
Affinity DataKi: 0.0400nMAssay Description:Displacement of [3H]-1alpha,25(OH)2D3 from recombinant rat VDR after overnight incubation by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 0.0400nMAssay Description:Displacement of [3H]1alpha,25-(OH)2D3 from full-length recombinant rat VDR by scintillation counterMore data for this Ligand-Target Pair
Affinity DataKi: 0.0430nMAssay Description:The experiment is carried out by the method according to Journal of Pharmacology and Experimental Therapeutics 2010, 333(1): 328. With [3H]methyl-spi...More data for this Ligand-Target Pair
Affinity DataKi: 0.0430nMAssay Description:The experiment is carried out by the method according to Journal of Pharmacology and Experimental Therapeutics 2010, 333(1): 328. With [3H]methyl-spi...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:Displacement of [3H]1alpha,25-(OH)2D3 from rat recombinant full length VDRMore data for this Ligand-Target Pair
Affinity DataKi: 0.0560nMAssay Description:The experiment is carried out by the method according to Journal of Pharmacology and Experimental Therapeutics 2010, 333(1): 328. With [3H]methyl-spi...More data for this Ligand-Target Pair
Affinity DataKi: 0.0560nMAssay Description:The experiment is carried out by the method according to Journal of Pharmacology and Experimental Therapeutics 2010, 333(1): 328. With [3H]methyl-spi...More data for this Ligand-Target Pair
Affinity DataKi: 0.0580nMAssay Description:The experiment is carried out by the method according to Journal of Pharmacology and Experimental Therapeutics 2010, 333(1): 328. With [3H]methyl-spi...More data for this Ligand-Target Pair
Affinity DataKi: 0.0580nMAssay Description:The experiment is carried out by the method according to Journal of Pharmacology and Experimental Therapeutics 2010, 333(1): 328. With [3H]methyl-spi...More data for this Ligand-Target Pair
Affinity DataKi: 0.0600nMAssay Description:Displacement of [3H]-1alpha,25(OH)2D3 from recombinant rat VDR after overnight incubation by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 0.0600nMAssay Description:Activity at rat recombinant full length VDR expressed in rat ROS 17/2.8 cells transfected with 24-hydroxylase gene promoter assessed as transcription...More data for this Ligand-Target Pair
Affinity DataKi: 0.0600nMAssay Description:Displacement of [3H]1alpha,25-(OH)2D3 from rat recombinant full length VDRMore data for this Ligand-Target Pair
Affinity DataKi: 0.0610nMAssay Description:The experiment is carried out by the method according to Journal of Pharmacology and Experimental Therapeutics 2010, 333(1): 328. With [3H]methyl-spi...More data for this Ligand-Target Pair
Affinity DataKi: 0.0610nMAssay Description:The experiment is carried out by the method according to Journal of Pharmacology and Experimental Therapeutics 2010, 333(1): 328. With [3H]methyl-spi...More data for this Ligand-Target Pair
Affinity DataKi: 0.0620nMAssay Description:The experiment is carried out by the method according to Journal of Pharmacology and Experimental Therapeutics 2010, 333(1): 328. With [3H]methyl-spi...More data for this Ligand-Target Pair
Affinity DataKi: 0.0620nMAssay Description:The experiment is carried out by the method according to Journal of Pharmacology and Experimental Therapeutics 2010, 333(1): 328. With [3H]methyl-spi...More data for this Ligand-Target Pair

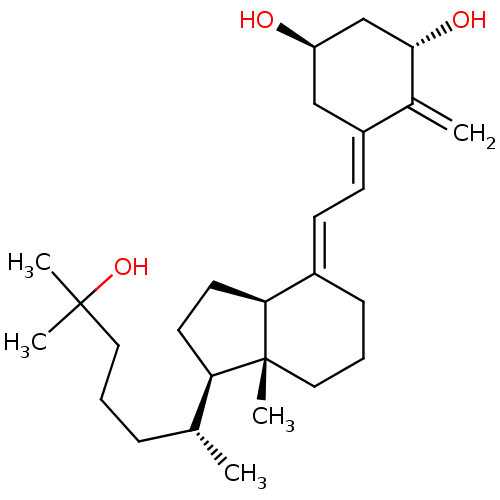


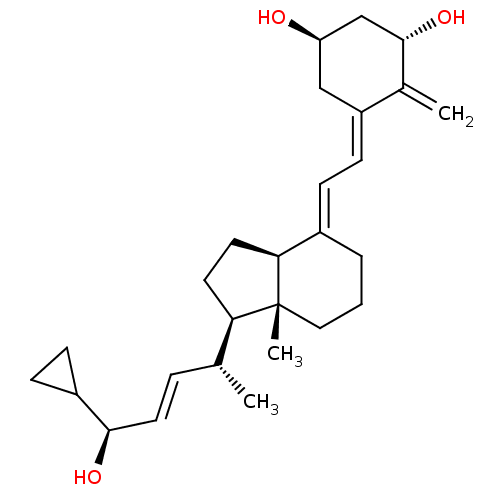
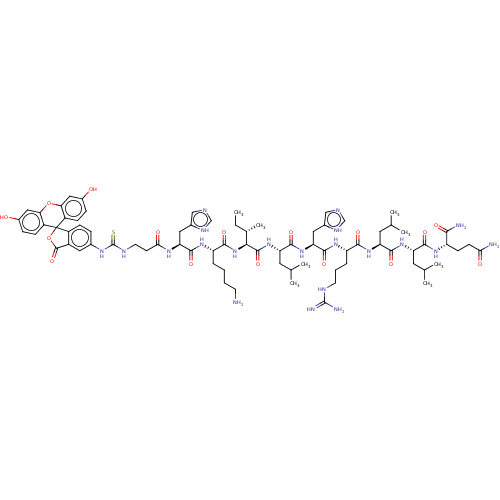
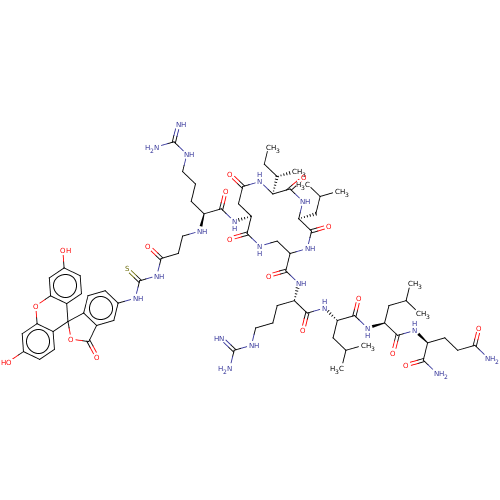
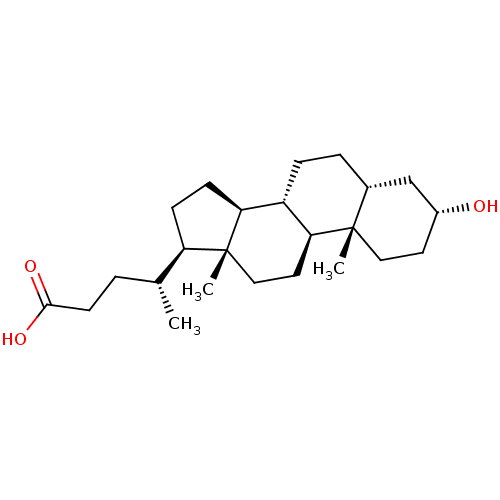
 3D Structure (crystal)
3D Structure (crystal)