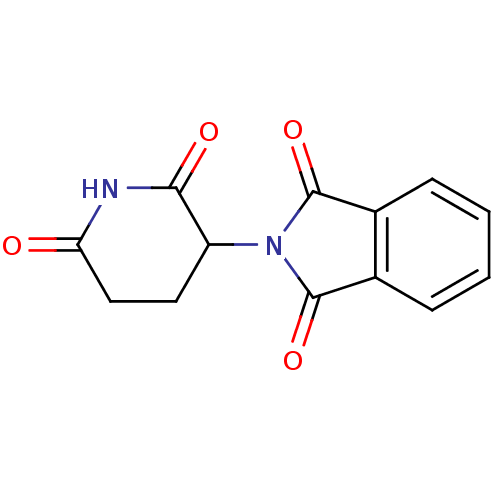
BDBM50070114 (+/-)-thalidomide::2-(2,6-Dioxo-piperidin-3-yl)-isoindole-1,3-dione::2-(2,6-dioxopiperidin-3-yl)isoindoline-1,3-dione::CHEMBL468::K-17::THALIDOMIDE::Thalomid::US11059801, Compound D-62::[(R,S)-2-(2,6-dioxo-3-piperidinyl)-1H-isoindole-1,3(2H)-dione
SMILES O=C1N(C2CCC(=O)NC2=O)C(=O)c2ccccc12
InChI Key InChIKey=UEJJHQNACJXSKW-UHFFFAOYSA-N
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 35 hits for monomerid = 50070114
Found 35 hits for monomerid = 50070114
Affinity DataKd: 250nMAssay Description:Binding affinity to CRBN (unknown origin) assessed as dissociation constant by TR-FRET assayMore data for this Ligand-Target Pair
Affinity DataKd: 250nMAssay Description:Binding affinity to CRBN (unknown origin) assessed as dissociation constantMore data for this Ligand-Target Pair
Affinity DataIC50: 268nMAssay Description:A thalidomide competition AlphaScreen assay was employed to measure the binding affinity (IC50) of thalidomide conjugates and novel IMiDs to CRBN-DDB...More data for this Ligand-Target Pair
Affinity DataIC50: 301nMAssay Description:A thalidomide competition AlphaScreen assay was employed to measure the binding affinity (IC50) of thalidomide conjugates and novel IMiDs to CRBN-DDB...More data for this Ligand-Target Pair
Affinity DataEC50: 330nMAssay Description:Induction of ePL tagged Aiolos degradation in human DF15 cells incubated for 4 hrs by luminescence based assayMore data for this Ligand-Target Pair
Affinity DataIC50: 370nMAssay Description:Inhibitory activity (RA1) against Prostaglandin G/H synthase 1 was calculated relative to aspirinMore data for this Ligand-Target Pair
Affinity DataIC50: 500nMAssay Description:Inhibitory activity (RA2) against Prostaglandin G/H synthase 2 was calculated relative to aspirinMore data for this Ligand-Target Pair
Affinity DataKd: >1.00E+3nMAssay Description:The binding affinity of the described test compounds was determined using RED-tris-NTA His-tag labeled Cereblon ligand binding domain. The labeling o...More data for this Ligand-Target Pair
Affinity DataIC50: 1.49E+3nMAssay Description:Displacement of fluorescent tracer from NanoLuc-tagged CRBN (unknown origin) by NanoBRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.80E+3nMAssay Description:Binding affinity to CRBN (unknown origin) assessed as inhibition constantMore data for this Ligand-Target Pair
Affinity DataKi: 1.80E+3nMAssay Description:Binding affinity to GST-tagged wild-type human CRBN incubated for 3 hrs by TR-FRET assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.70E+3nMAssay Description:Inhibition of biotinylated thalidomide binding to CRBN (40 to 442 residues) (unknown origin) incubated for 1 hrs by HTRF assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.90E+3nMAssay Description:Binding affinity to GST-tagged wild type human CRBN incubated for 3 hrs by TR-FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 4.40E+3nMAssay Description:Binding affinity to CRBN (unknown origin) assessed as inhibition constant by FRET assayMore data for this Ligand-Target Pair
TargetCereblon isoform 4(Magnetospirillum gryphiswaldense)
Max Planck Institute For Developmental Biology
Curated by ChEMBL
Max Planck Institute For Developmental Biology
Curated by ChEMBL
Affinity DataKi: 4.40E+3nMAssay Description:Inhibition of MANT-uracil binding to wild type Magnetospirillum gryphiswaldense cereblon isoform 4 by Cheng-Prusoff equation analysisMore data for this Ligand-Target Pair
TargetCereblon isoform 4(Magnetospirillum gryphiswaldense)
Max Planck Institute For Developmental Biology
Curated by ChEMBL
Max Planck Institute For Developmental Biology
Curated by ChEMBL
Affinity DataKi: 4.40E+3nMAssay Description:Inhibition of MANT-uracil binding to wild-type Magnetospirillum gryphiswaldense CRBN isoform 4 by FRET assayMore data for this Ligand-Target Pair
TargetCereblon isoform 4(Magnetospirillum gryphiswaldense)
Max Planck Institute For Developmental Biology
Curated by ChEMBL
Max Planck Institute For Developmental Biology
Curated by ChEMBL
Affinity DataIC50: 7.80E+3nMAssay Description:Inhibition of MANT-uracil binding to wild type Magnetospirillum gryphiswaldense cereblon isoform 4 by FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 8.50E+3nMAssay Description:Binding affinity to human CRBN thalidomide binding domain assessed as inhibition constant by microscale thermophoresis analysisMore data for this Ligand-Target Pair
Affinity DataKi: 8.50E+3nMAssay Description:Binding affinity to human CRBN TBD assessed as inhibition constant using BODIPY-uracil as substrate by Microscale thermophoresis assayMore data for this Ligand-Target Pair
Affinity DataKi: 8.51E+3nMAssay Description:Binding affinity to human CRBN-thalidomide binding domain expressed in Escherichia coli by measuring baseline corrected normalized fluorescence in pr...More data for this Ligand-Target Pair
Affinity DataKi: 8.55E+3nMAssay Description:Binding affinity to human CRBN TBD (319 to 425 residues) assessed as inhibition constant by MST assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMT: 2°CAssay Description:The Fix buffer I was warmed up to 37° C. in an incubator or water bath prior to use. The Perm Buffer III was chilled in a 20° C. freezer prio...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMT: 2°CAssay Description:The Fix buffer I was warmed up to 37° C. in an incubator or water bath prior to use. The Perm Buffer III was chilled in a 20° C. freezer prio...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMT: 2°CAssay Description:The Fix buffer I was warmed up to 37° C. in an incubator or water bath prior to use. The Perm Buffer III was chilled in a 20° C. freezer prio...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMT: 2°CAssay Description:The Fix buffer I was warmed up to 37° C. in an incubator or water bath prior to use. The Perm Buffer III was chilled in a 20° C. freezer prio...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMT: 2°CAssay Description:The Fix buffer I was warmed up to 37° C. in an incubator or water bath prior to use. The Perm Buffer III was chilled in a 20° C. freezer prio...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMT: 2°CAssay Description:The Fix buffer I was warmed up to 37° C. in an incubator or water bath prior to use. The Perm Buffer III was chilled in a 20° C. freezer prio...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMT: 2°CAssay Description:The Fix buffer I was warmed up to 37° C. in an incubator or water bath prior to use. The Perm Buffer III was chilled in a 20° C. freezer prio...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMT: 2°CAssay Description:The Fix buffer I was warmed up to 37° C. in an incubator or water bath prior to use. The Perm Buffer III was chilled in a 20° C. freezer prio...More data for this Ligand-Target Pair
Affinity DataKi: 2.27E+4nMAssay Description:Inhibition of MANT-uracil binding to human CRBN (delta 1 to 315) by Cheng-Prusoff equation analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 2.29E+4nMAssay Description:Binding affinity to human CRBN-thalidomide binding domain expressed in Escherichia coli by measuring baseline corrected normalized fluorescence in pr...More data for this Ligand-Target Pair
Affinity DataIC50: 2.99E+4nMAssay Description:Inhibition of MANT-uracil binding to human CRBN (delta 1 to 315) by FRET assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+4nMpH: 8.0 T: 2°CAssay Description:Thermal stabilities of CRBN-DDB1 in the presence or absence of phthalimide, thalidomide, lenalidomide and pomalidomide were done in the presence of S...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+5nMAssay Description:Inhibition of PDE4 enzyme from U937 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+6nMAssay Description:Inhibition of human BSEP expressed in baculovirus transfected fall armyworm Sf21 cell membranes vesicles assessed as reduction in ATP-dependent [3H]-...More data for this Ligand-Target Pair