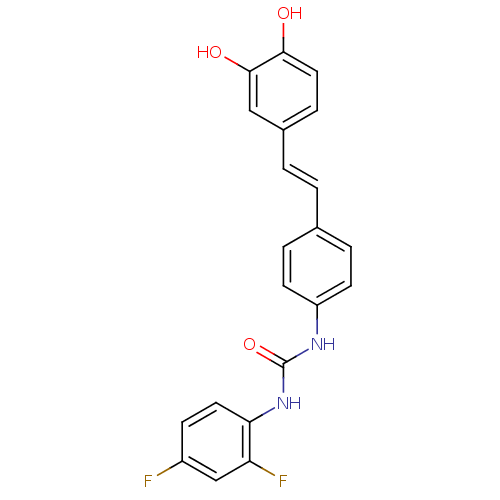
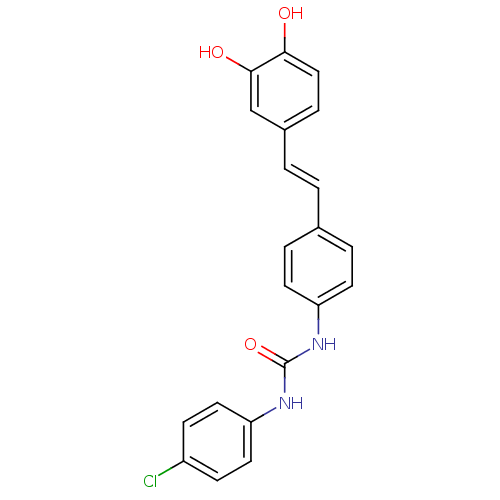
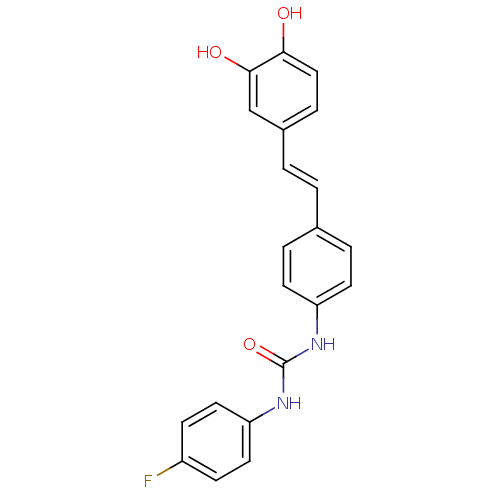
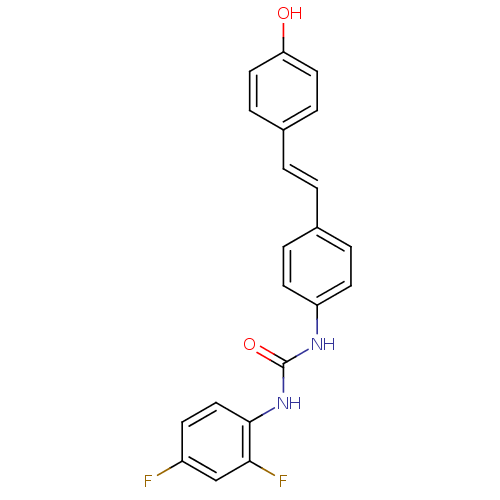
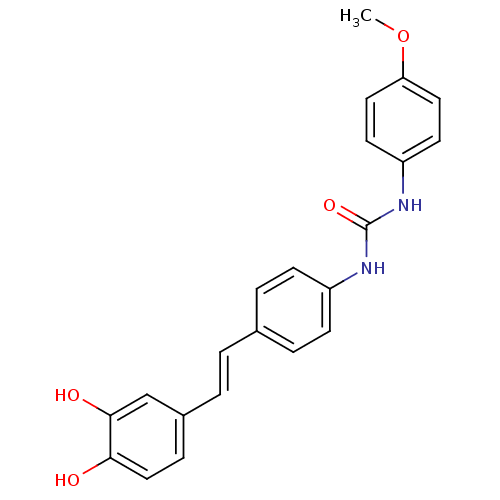
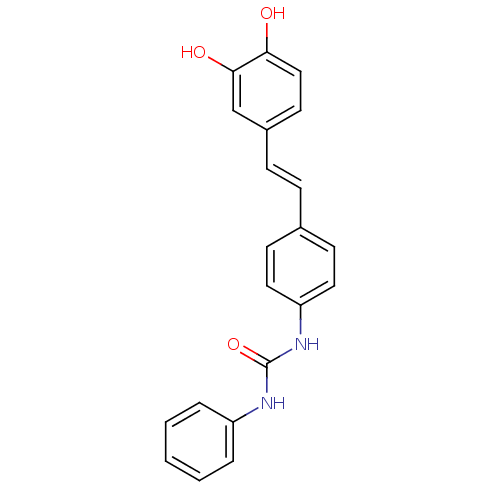
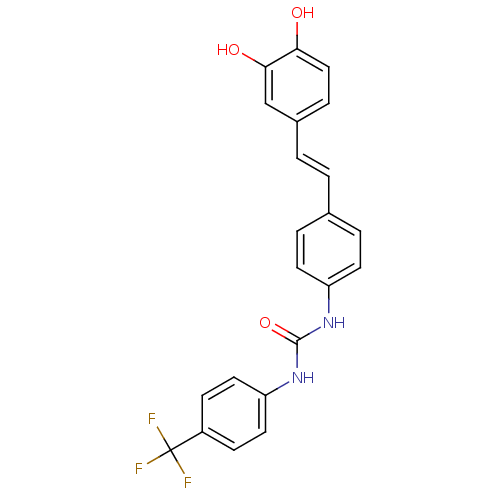
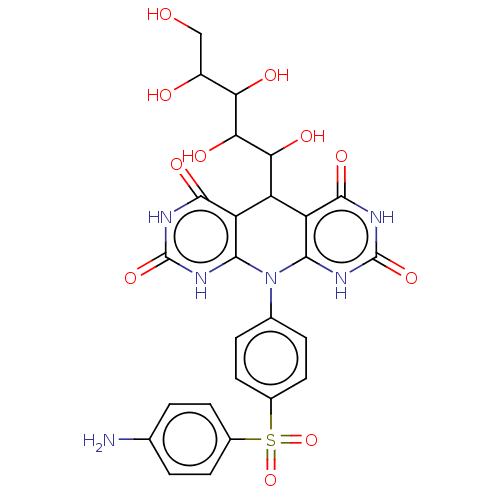
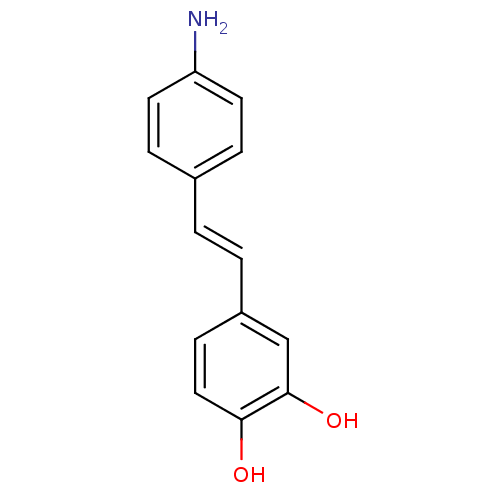
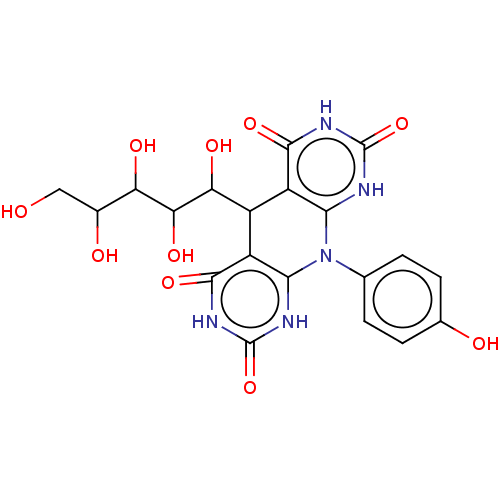
Affinity DataKi: 3.20E+3nM IC50: 8.40E+3nMAssay Description:All enzymatic activities were determined by using the appropriate substrate (p-nitrophenyl-alpha-D-glucopyranoside, p-nitrophenyl-beta-D-gulcopyranos...More data for this Ligand-Target Pair
Affinity DataKi: 4.60E+3nM IC50: 1.43E+4nMAssay Description:All enzymatic activities were determined by using the appropriate substrate (p-nitrophenyl-alpha-D-glucopyranoside, p-nitrophenyl-beta-D-gulcopyranos...More data for this Ligand-Target Pair
Affinity DataKi: 7.20E+3nM IC50: 1.98E+4nMAssay Description:All enzymatic activities were determined by using the appropriate substrate (p-nitrophenyl-alpha-D-glucopyranoside, p-nitrophenyl-beta-D-gulcopyranos...More data for this Ligand-Target Pair
Affinity DataKi: 9.40E+3nM IC50: 1.85E+4nMAssay Description:All enzymatic activities were determined by using the appropriate substrate (p-nitrophenyl-alpha-D-glucopyranoside, p-nitrophenyl-beta-D-gulcopyranos...More data for this Ligand-Target Pair
Affinity DataKi: 1.05E+4nM IC50: 1.95E+4nMAssay Description:All enzymatic activities were determined by using the appropriate substrate (p-nitrophenyl-alpha-D-glucopyranoside, p-nitrophenyl-beta-D-gulcopyranos...More data for this Ligand-Target Pair
Affinity DataKi: 1.06E+4nM IC50: 4.21E+4nMAssay Description:All enzymatic activities were determined by using the appropriate substrate (p-nitrophenyl-alpha-D-glucopyranoside, p-nitrophenyl-beta-D-gulcopyranos...More data for this Ligand-Target Pair
Affinity DataKi: 1.21E+4nM IC50: 2.88E+4nMAssay Description:All enzymatic activities were determined by using the appropriate substrate (p-nitrophenyl-alpha-D-glucopyranoside, p-nitrophenyl-beta-D-gulcopyranos...More data for this Ligand-Target Pair
Affinity DataKi: 1.40E+4nMpH: 7.0Assay Description:In this study, the inhibition assay of yeast enzyme was performed in 100 mM phosphate buffer pH 7.0 at 25�C with minor changes, according to the meth...More data for this Ligand-Target Pair
Affinity DataKi: 1.70E+4nM IC50: 2.91E+4nMAssay Description:All enzymatic activities were determined by using the appropriate substrate (p-nitrophenyl-alpha-D-glucopyranoside, p-nitrophenyl-beta-D-gulcopyranos...More data for this Ligand-Target Pair
Affinity DataKi: 1.80E+4nM IC50: 3.95E+4nMAssay Description:All enzymatic activities were determined by using the appropriate substrate (p-nitrophenyl-alpha-D-glucopyranoside, p-nitrophenyl-beta-D-gulcopyranos...More data for this Ligand-Target Pair
Affinity DataKi: 2.15E+4nM IC50: 2.64E+4nMAssay Description:All enzymatic activities were determined by using the appropriate substrate (p-nitrophenyl-alpha-D-glucopyranoside, p-nitrophenyl-beta-D-gulcopyranos...More data for this Ligand-Target Pair
Affinity DataKi: 3.00E+4nMpH: 7.0Assay Description:In this study, the inhibition assay of yeast enzyme was performed in 100 mM phosphate buffer pH 7.0 at 25�C with minor changes, according to the meth...More data for this Ligand-Target Pair
Affinity DataKi: 3.51E+4nM IC50: 4.64E+4nMAssay Description:All enzymatic activities were determined by using the appropriate substrate (p-nitrophenyl-alpha-D-glucopyranoside, p-nitrophenyl-beta-D-gulcopyranos...More data for this Ligand-Target Pair
Affinity DataKi: 7.00E+4nMpH: 7.0Assay Description:In this study, the inhibition assay of yeast enzyme was performed in 100 mM phosphate buffer pH 7.0 at 25�C with minor changes, according to the meth...More data for this Ligand-Target Pair
Affinity DataKi: 1.30E+5nMpH: 7.0Assay Description:In this study, the inhibition assay of yeast enzyme was performed in 100 mM phosphate buffer pH 7.0 at 25�C with minor changes, according to the meth...More data for this Ligand-Target Pair
Affinity DataKi: 1.70E+5nMpH: 7.0Assay Description:In this study, the inhibition assay of yeast enzyme was performed in 100 mM phosphate buffer pH 7.0 at 25�C with minor changes, according to the meth...More data for this Ligand-Target Pair