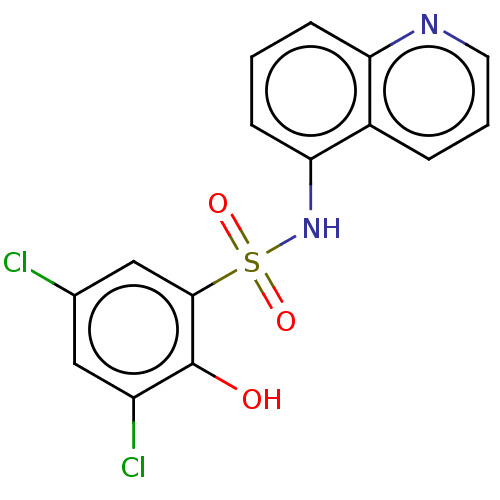
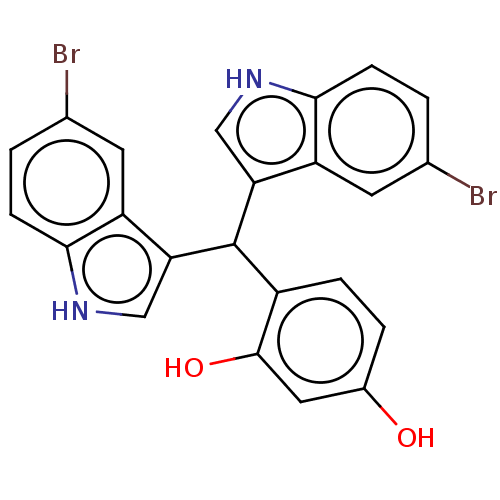
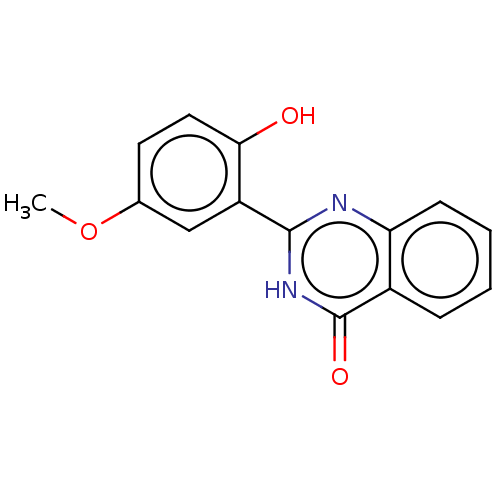
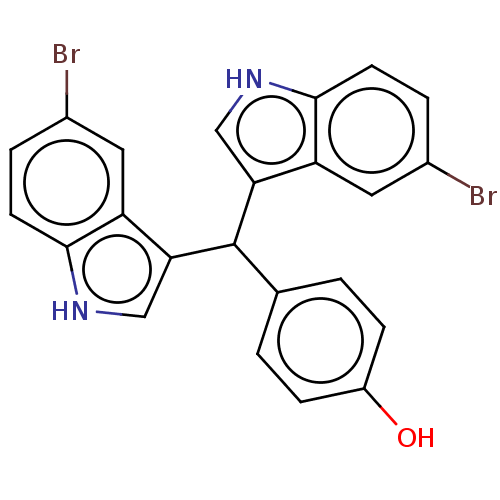
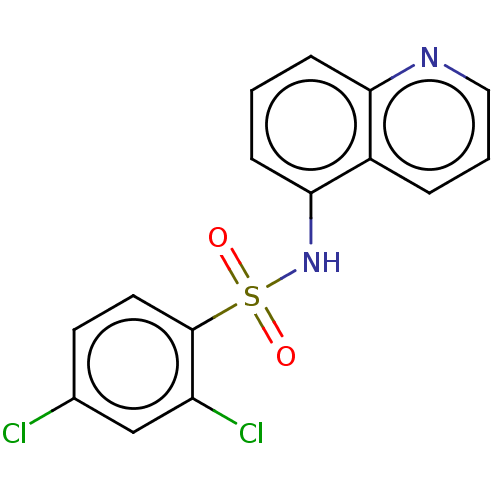
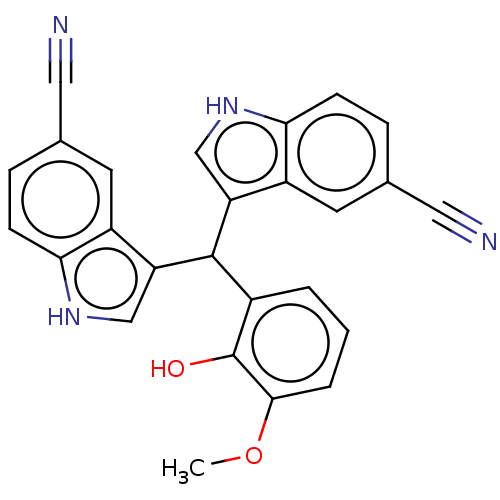
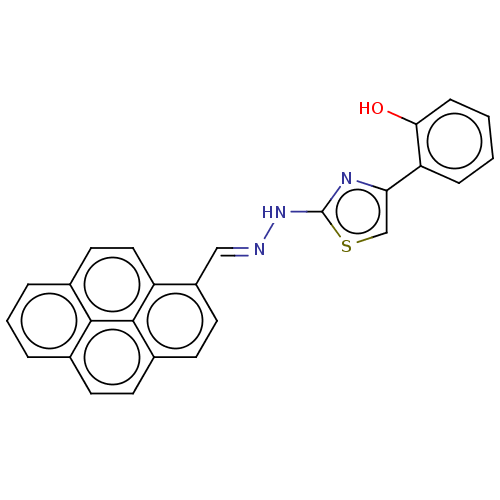
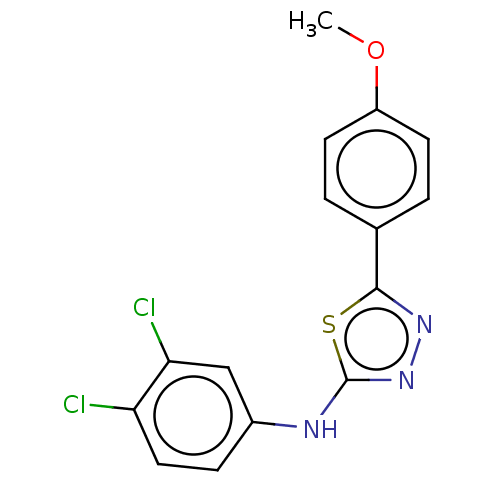
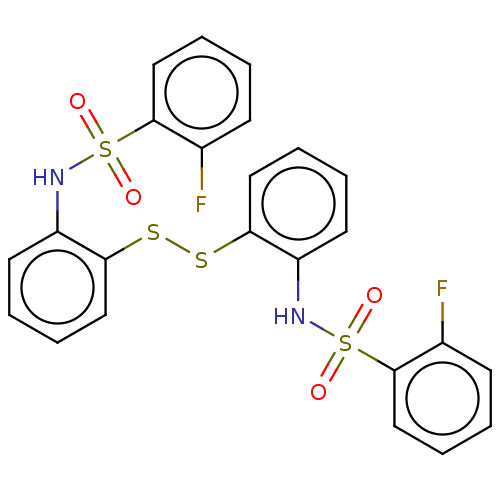
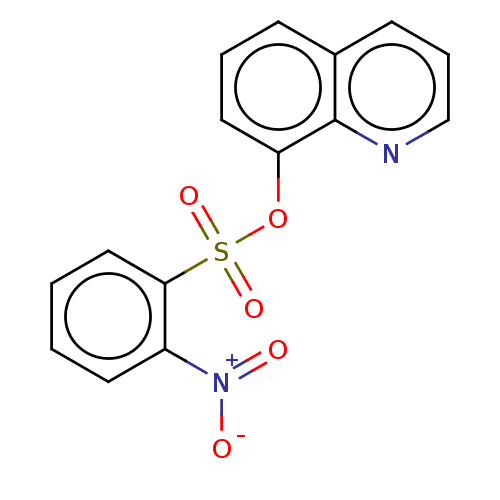
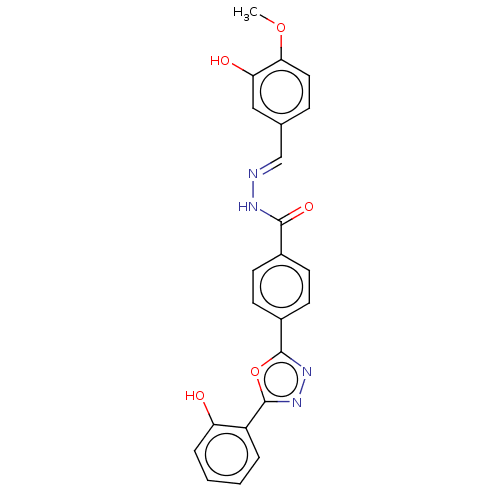
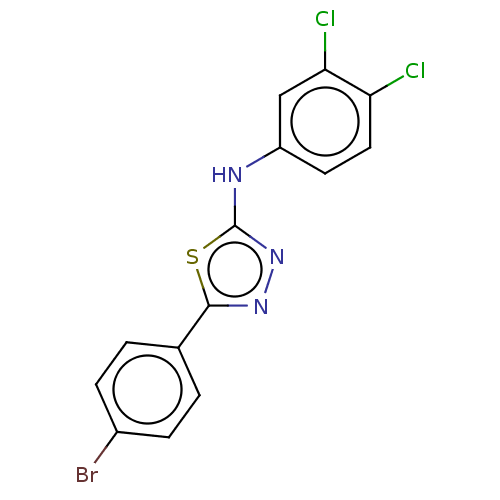
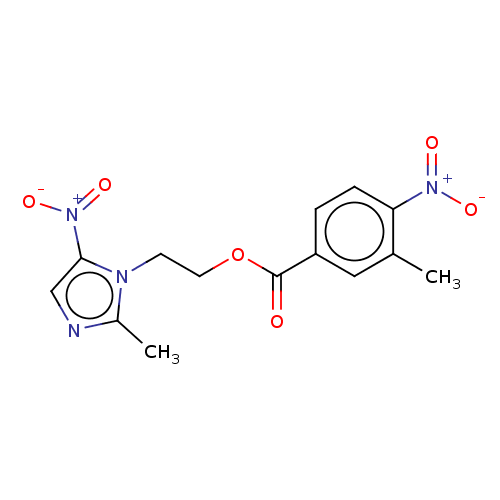
Affinity DataKi: 12nMAssay Description:Inhibition of bovine GUS pre-incubated for 5 to 30 mins before 4-methylumbelliferone beta-D-glucuronide substrate addition at pH 6.7More data for this Ligand-Target Pair
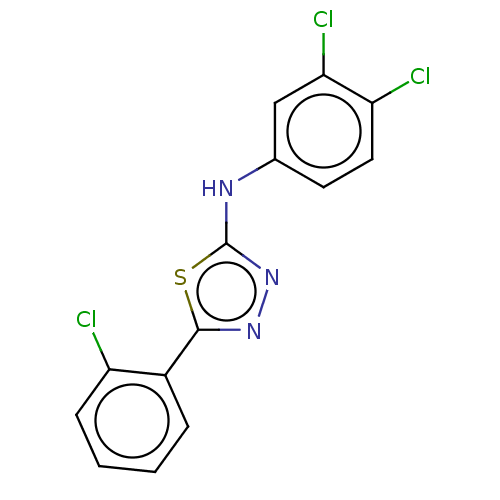
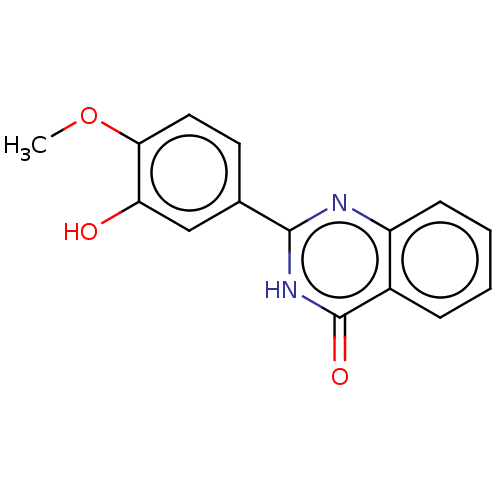
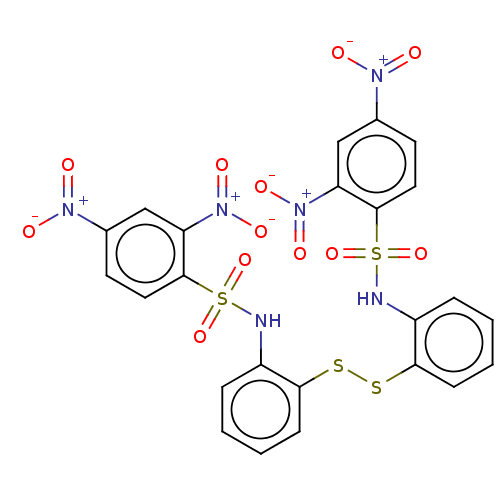
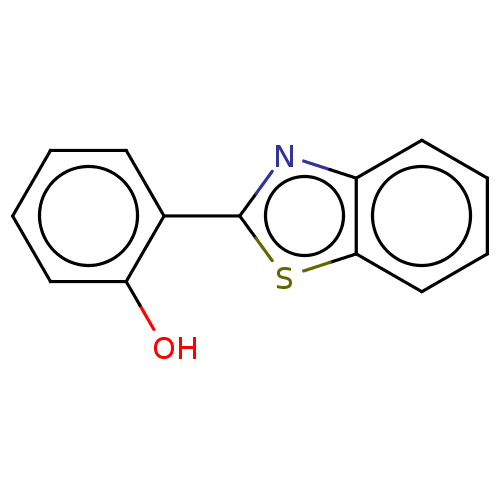
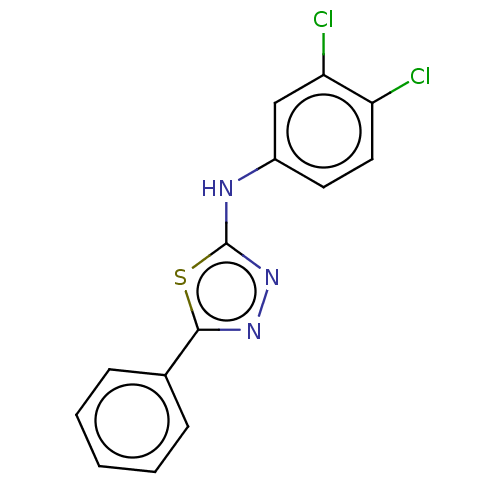
Affinity DataKi: 48nMAssay Description:Inhibition of bovine GUS pre-incubated for 5 to 30 mins before 4-methylumbelliferone beta-D-glucuronide substrate addition at pH 4.5More data for this Ligand-Target Pair
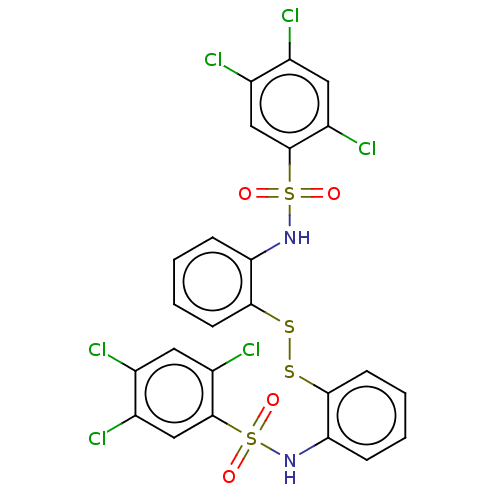
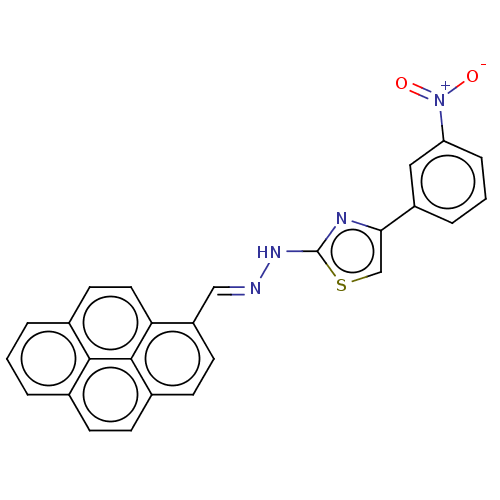
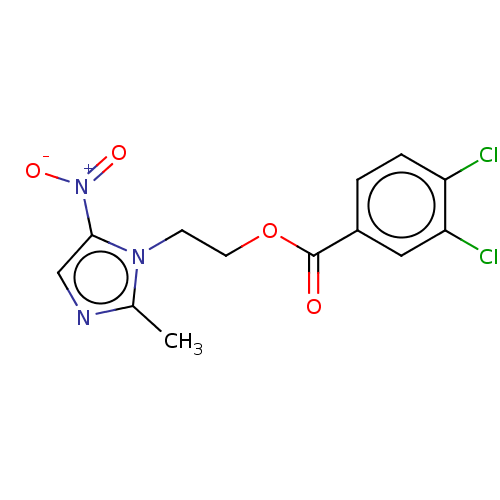
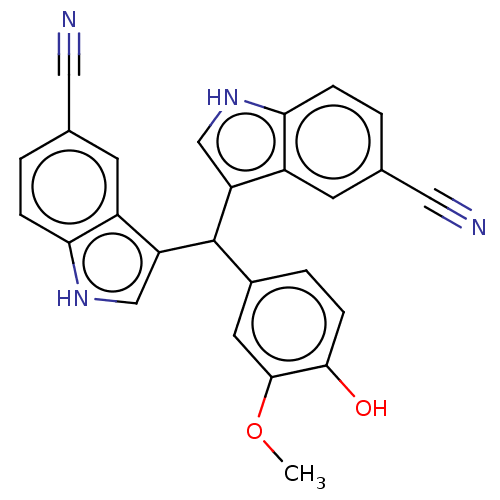
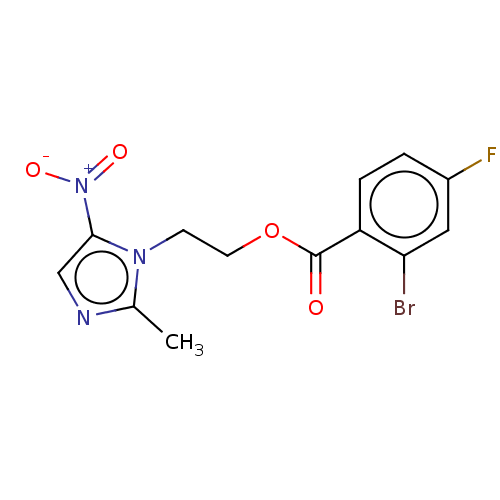
Affinity DataKi: 132nMAssay Description:Inhibition of bovine GUS pre-incubated for 5 to 30 mins before 4-methylumbelliferone beta-D-glucuronide substrate addition at pH 4.5More data for this Ligand-Target Pair
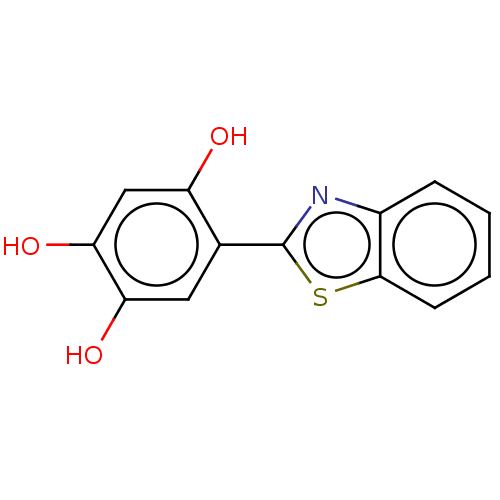
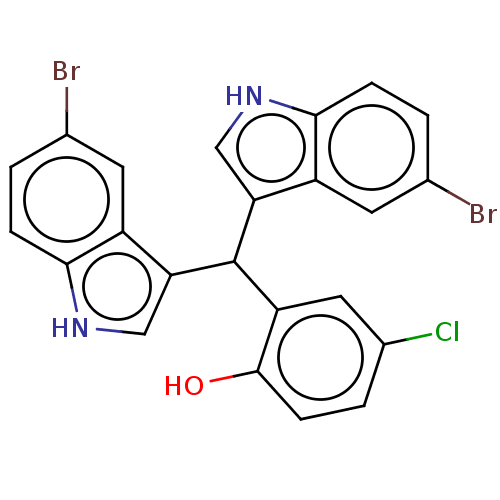
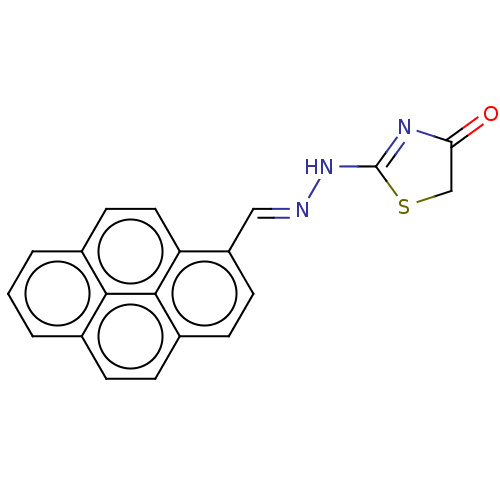
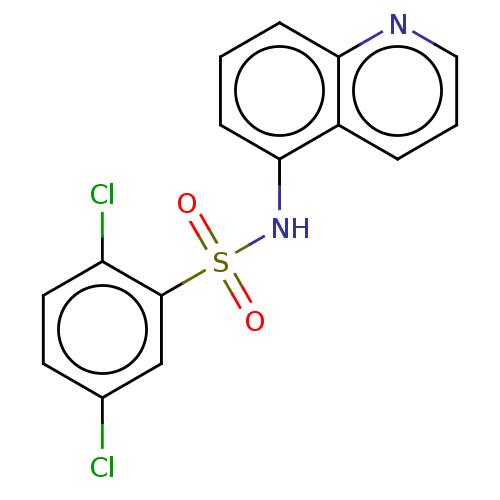
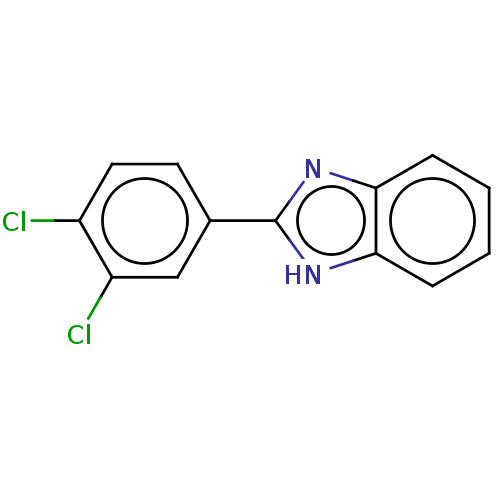
Affinity DataIC50: 700nMAssay Description:Inhibition of bovine liver beta-glucuronidase using p-nitrophenyl-beta-D-glucuronide as substrate preincubated for 30 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+3nMAssay Description:Inhibition of bovine liver beta-glucuronidase using p-nitrophenyl-beta-D-glucuronide as substrate preincubated for 30 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+3nMAssay Description:Inhibition of bovine liver beta glucuronidase using p-nitrophenyl-beta-D-glucuronide as substrate preincubated for 30 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+3nMAssay Description:Inhibition of bovine liver beta-glucuronidase using p-nitrophenyl-beta-D-glucuronide as substrate preincubated for 30 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 1.62E+3nMAssay Description:Inhibition of bovine liver beta-glucuronidase using p-nitrophenyl-beta-D-glucuronide as substrate after 30 mins by spectrophotometryMore data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+3nMAssay Description:Inhibition of bovine liver beta-glucuronidase using p-nitrophenyl-beta-D-glucuronide as substrate preincubated for 30 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 1.86E+3nMAssay Description:Inhibition of bovine liver beta-glucuronidase using p-nitrophenyl-beta-D-glucuronide as substrate after 30 mins by spectrophotometryMore data for this Ligand-Target Pair
Affinity DataIC50: 2.10E+3nMAssay Description:Inhibition of bovine liver beta glucuronidase using p-nitrophenyl-beta-D-glucuronide as substrate preincubated for 30 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 2.10E+3nMAssay Description:Inhibition of bovine liver beta-glucuronidase using p-nitrophenyl-beta-D-glucuronide as substrate preincubated for 30 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 2.14E+3nMAssay Description:Inhibition of bovine beta-glucuronidase assessed as p-nitrophenol formation preincubated for 30 mins followed by addition of p-nitrophenyl-beta-D-glu...More data for this Ligand-Target Pair
Affinity DataIC50: 2.16E+3nMAssay Description:Inhibition of bovine beta-glucuronidase assessed as p-nitrophenol formation preincubated for 30 mins followed by addition of p-nitrophenyl-beta-D-glu...More data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+3nMAssay Description:β-glucuronidase activity was determined by measuring absorbance at 405 nm of p-nitrophenol formed substrate by spectrophotometric method. 250 µL...More data for this Ligand-Target Pair
Affinity DataIC50: 2.26E+3nMAssay Description:Inhibition of bovine liver beta glucuronidase assessed as p-nitrophenol formation after 30 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 2.80E+3nMAssay Description:Inhibition of bovine liver beta-glucuronidase using p-nitrophenyl-beta-D-glucuronide as substrate after 30 mins by spectrophotometryMore data for this Ligand-Target Pair
Affinity DataIC50: 2.80E+3nMAssay Description:Inhibition of bovine liver beta-glucuronidase using p-nitrophenyl-beta-D-glucuronide as substrate preincubated for 30 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 3.10E+3nMAssay Description:β-Glucuronidase inhibitory activity was measured by quantifying the absorbance of para-nitrophenol which was formed from the substrate by the sp...More data for this Ligand-Target Pair
Affinity DataIC50: 3.10E+3nMAssay Description:Inhibition of bovine liver beta-glucuronidase using p-nitrophenyl-beta-D-glucuronide as substrate after 30 mins by spectrophotometryMore data for this Ligand-Target Pair
Affinity DataIC50: 3.20E+3nMAssay Description:β-Glucuronidase inhibitory activity was measured by quantifying the absorbance of para-nitrophenol which was formed from the substrate by the sp...More data for this Ligand-Target Pair
Affinity DataIC50: 3.34E+3nMAssay Description:β-glucuronidase activity was determined by measuring absorbance at 405 nm of p-nitrophenol formed substrate by spectrophotometric method. 250 µL...More data for this Ligand-Target Pair
Affinity DataIC50: 3.60E+3nMAssay Description:Inhibition of bovine liver beta glucuronidase using p-nitrophenyl-beta-D-glucuronide as substrate preincubated for 30 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 4.10E+3nMAssay Description:β-Glucuronidase inhibitory activity was measured by quantifying the absorbance of para-nitrophenol which was formed from the substrate by the sp...More data for this Ligand-Target Pair
Affinity DataIC50: 4.16E+3nMAssay Description:Inhibition of bovine beta-glucuronidase assessed as p-nitrophenol formation preincubated for 30 mins followed by addition of p-nitrophenyl-beta-D-glu...More data for this Ligand-Target Pair
Affinity DataIC50: 4.23E+3nMAssay Description:Inhibition of bovine liver beta glucuronidase assessed as p-nitrophenol formation after 30 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 4.30E+3nMAssay Description:Inhibition of bovine liver beta-glucuronidase using p-nitrophenyl-beta-D-glucuronide as substrate after 30 mins by spectrophotometryMore data for this Ligand-Target Pair
Affinity DataIC50: 4.48E+3nMAssay Description:β-Glucuronidase activity was determined by measuring the absorbance at 405 nm of p-nitrophenol formed from the substrate by the spectrophotometr...More data for this Ligand-Target Pair
Affinity DataIC50: 4.50E+3nMAssay Description:Inhibition of bovine liver beta-glucuronidase using p-nitrophenyl-beta-D-glucuronide as substrate preincubated for 30 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataKi: 4.87E+3nMAssay Description:Inhibition of bovine GUS pre-incubated for 5 to 30 mins before 4-methylumbelliferone beta-D-glucuronide substrate addition at pH 4.5More data for this Ligand-Target Pair
Affinity DataIC50: 4.88E+3nMAssay Description:Inhibition of bovine beta-glucuronidase assessed as p-nitrophenol formation preincubated for 30 mins followed by addition of p-nitrophenyl-beta-D-glu...More data for this Ligand-Target Pair
Affinity DataIC50: 5.50E+3nMAssay Description:Inhibition of bovine liver beta-glucuronidase using p-nitrophenyl-beta-D-glucuronide as substrate preincubated for 30 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 5.50E+3nMAssay Description:β-Glucuronidase inhibitory activity was measured by quantifying the absorbance of para-nitrophenol which was formed from the substrate by the sp...More data for this Ligand-Target Pair
Affinity DataKi: 5.87E+3nMAssay Description:Inhibition of bovine GUS pre-incubated for 5 to 30 mins before 4-methylumbelliferone beta-D-glucuronide substrate addition at pH 6.7More data for this Ligand-Target Pair
Affinity DataIC50: 6.10E+3nMAssay Description:Inhibition of bovine liver beta-glucuronidase using p-nitrophenyl-beta-D-glucuronide as substrate preincubated for 30 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 6.20E+3nMAssay Description:Inhibition of bovine liver beta glucuronidase using p-nitrophenyl-beta-D-glucuronide as substrate preincubated for 30 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 6.20E+3nMAssay Description:β-Glucuronidase inhibitory activity was measured by quantifying the absorbance of para-nitrophenol which was formed from the substrate by the sp...More data for this Ligand-Target Pair
Affinity DataIC50: 6.23E+3nMAssay Description:β-glucuronidase activity was determined by measuring absorbance at 405 nm of p-nitrophenol formed substrate by spectrophotometric method. 250 µL...More data for this Ligand-Target Pair
Affinity DataIC50: 6.28E+3nMAssay Description:Inhibition of bovine beta-glucuronidase assessed as p-nitrophenol formation preincubated for 30 mins followed by addition of p-nitrophenyl-beta-D-glu...More data for this Ligand-Target Pair
Affinity DataIC50: 6.30E+3nMAssay Description:Inhibition of bovine liver beta glucuronidase using p-nitrophenyl-beta-D-glucuronide as substrate preincubated for 30 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 6.33E+3nMAssay Description:Inhibition of bovine liver beta glucuronidase using p-nitrophenyl-beta-D-glucuronide as substrate preincubated for 30 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 6.40E+3nMAssay Description:Inhibition of bovine liver beta-glucuronidase using p-nitrophenyl-beta-D-glucuronide as substrate preincubated for 30 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 6.80E+3nMAssay Description:Inhibition of bovine beta-glucuronidaseMore data for this Ligand-Target Pair
Affinity DataIC50: 7.14E+3nMAssay Description:Inhibition of bovine liver beta-glucuronidase using p-nitrophenyl-beta-D-glucuronide as substrate assessed as p-nitrophenol formation after 30 mins b...More data for this Ligand-Target Pair
Affinity DataIC50: 7.16E+3nMAssay Description:Inhibition of bovine beta-glucuronidase assessed as p-nitrophenol formation preincubated for 30 mins followed by addition of p-nitrophenyl-beta-D-glu...More data for this Ligand-Target Pair
Affinity DataIC50: 7.30E+3nMAssay Description:Inhibition of bovine liver beta glucuronidase using p-nitrophenyl-beta-D-glucuronide as substrate preincubated for 30 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 8.10E+3nMAssay Description:Inhibition of bovine liver beta-glucuronidase using p-nitrophenyl-beta-D-glucuronide as substrate preincubated for 30 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 8.10E+3nMAssay Description:Inhibition of bovine liver beta glucuronidase using p-nitrophenyl-beta-D-glucuronide as substrate preincubated for 30 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 8.18E+3nMAssay Description:β-glucuronidase activity was determined by measuring absorbance at 405 nm of p-nitrophenol formed substrate by spectrophotometric method. 250 µL...More data for this Ligand-Target Pair
Affinity DataIC50: 8.40E+3nMAssay Description:Inhibition of bovine liver beta glucuronidase using p-nitrophenyl-beta-D-glucuronide as substrate preincubated for 30 mins followed by substrate addi...More data for this Ligand-Target Pair