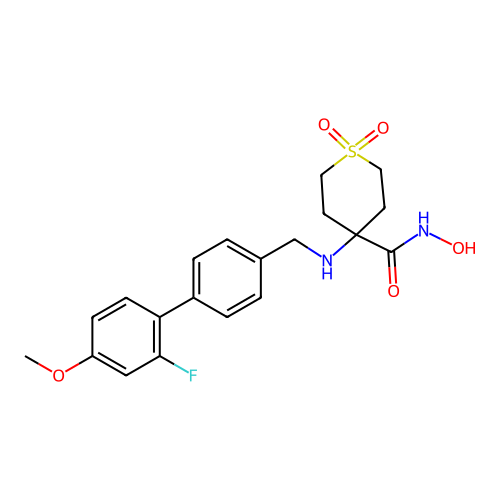
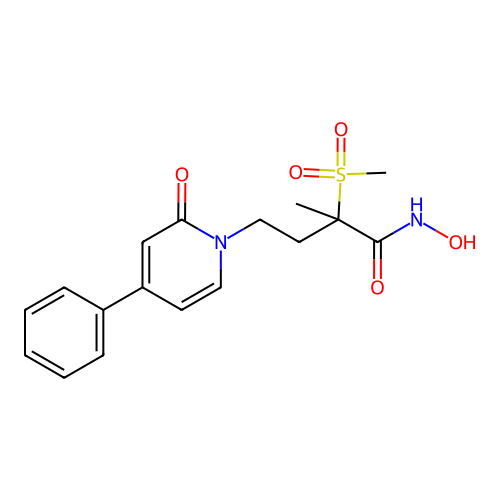
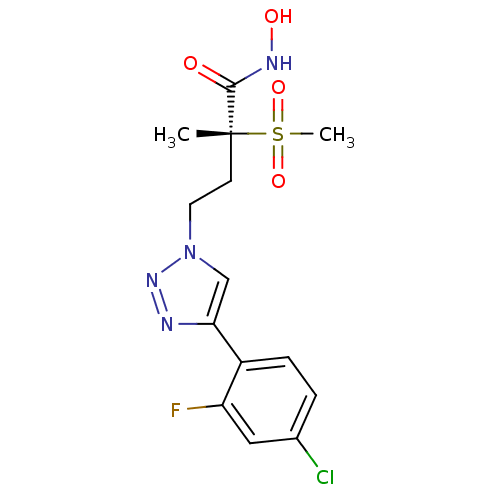
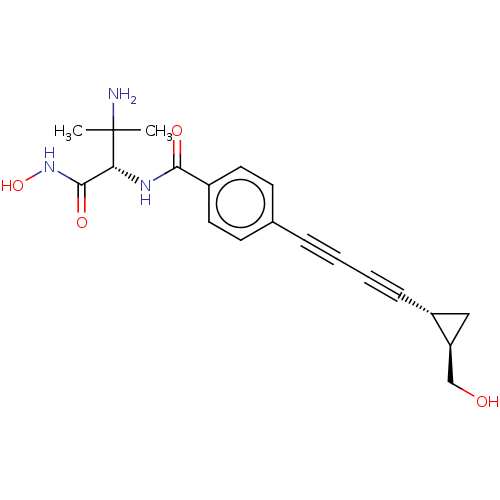
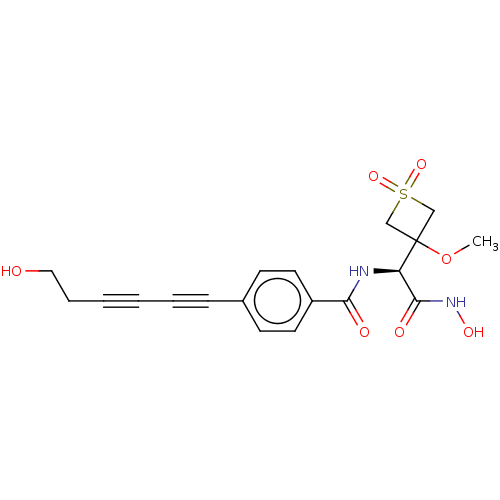
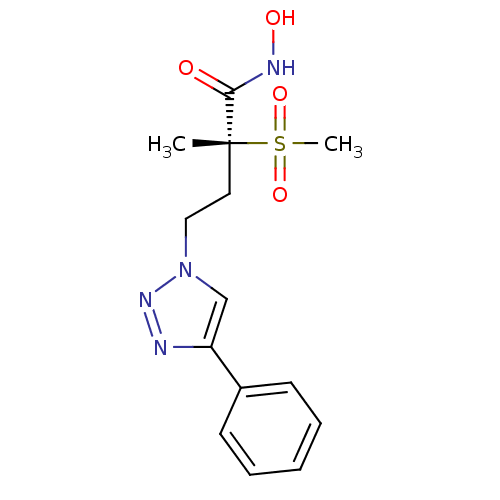
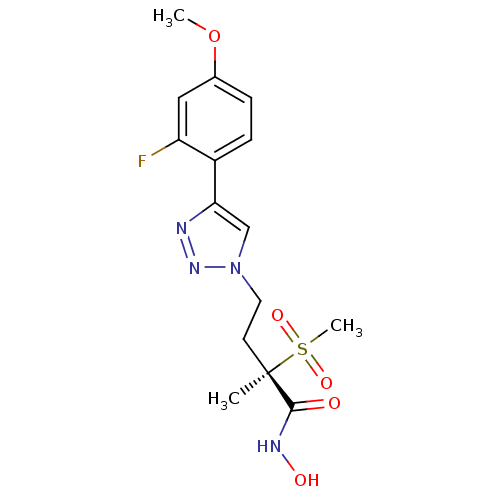
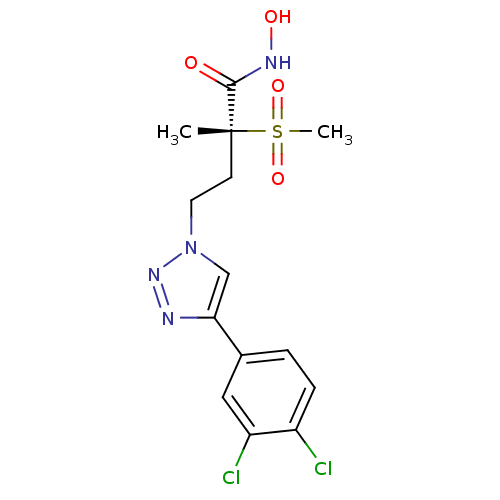
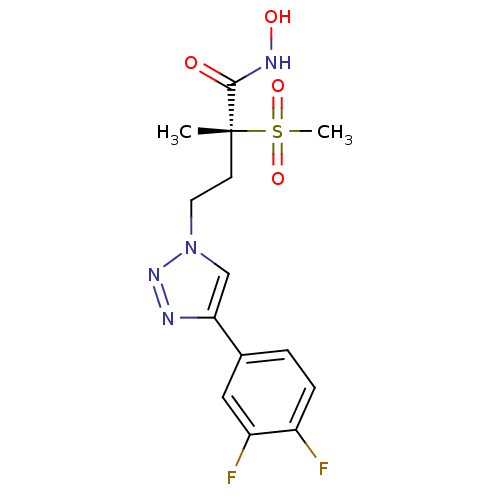
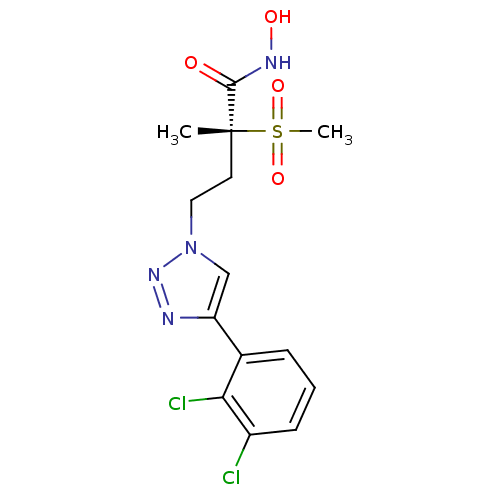
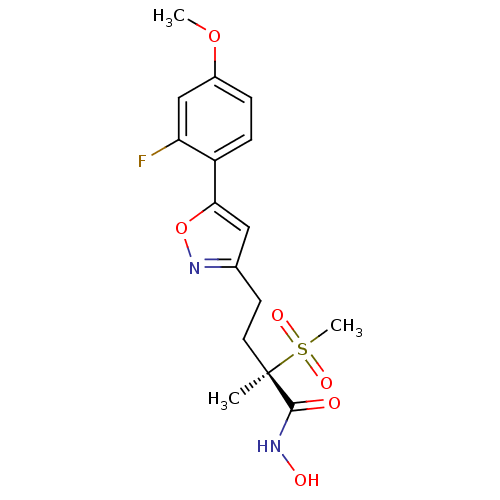
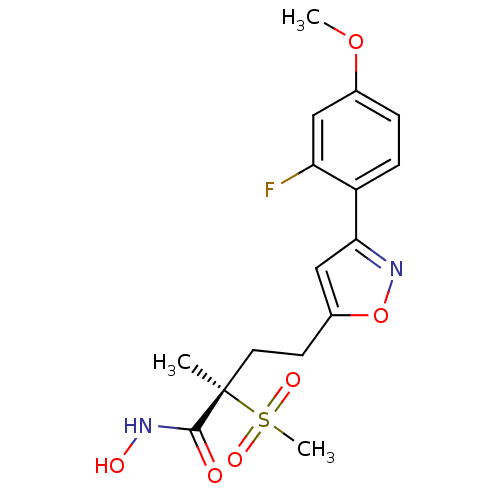
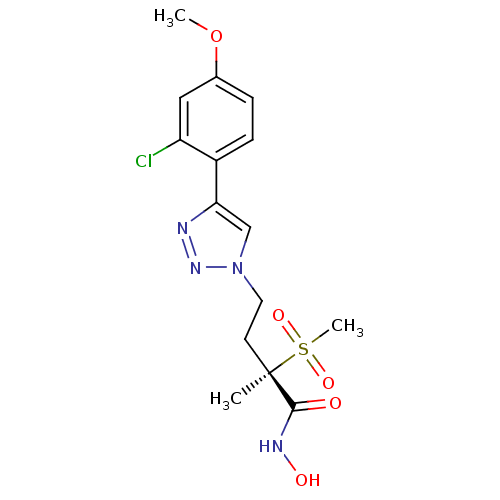
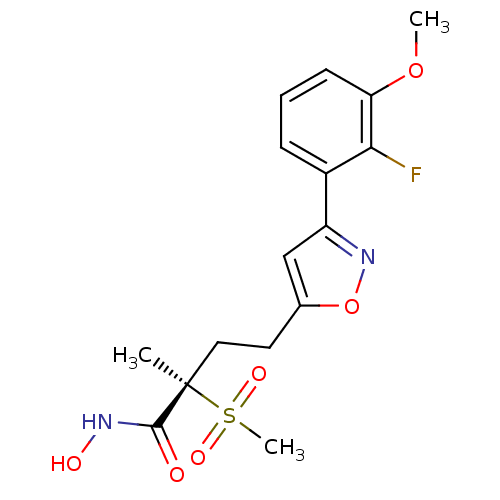
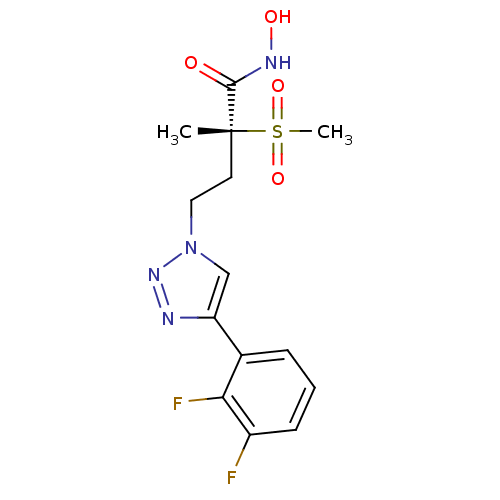
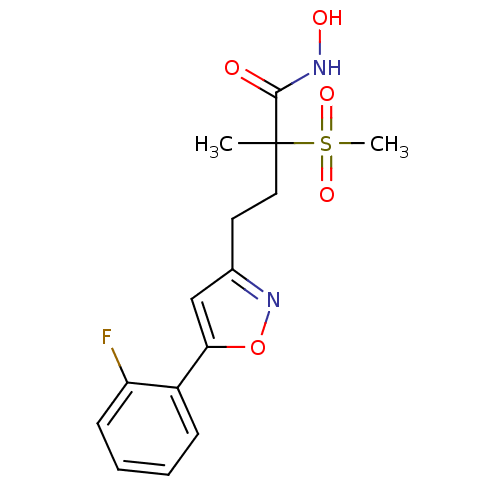
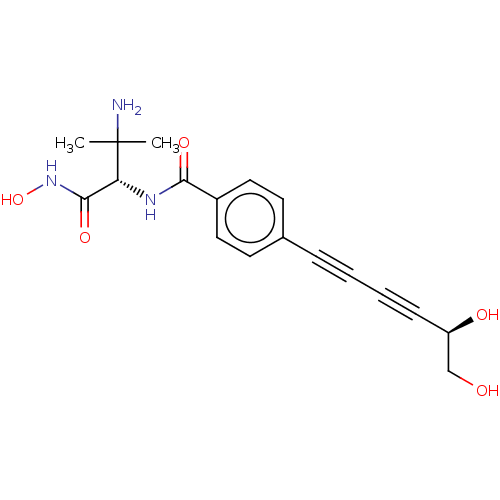
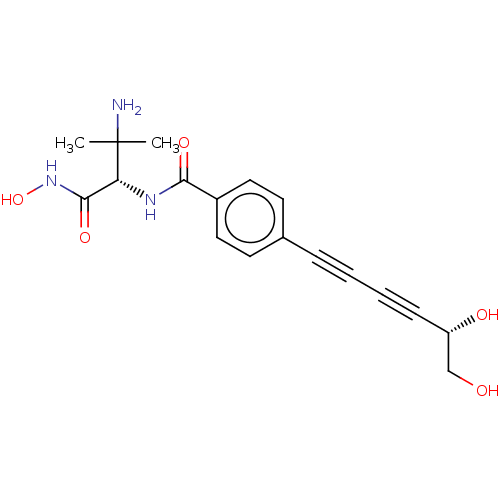
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataKi: 0.00210nMAssay Description:A fluorescence-based competition assay was used to determine the kinetic parameters for binding of the inhibitors to paLpxC. This assay is based on t...More data for this Ligand-Target Pair
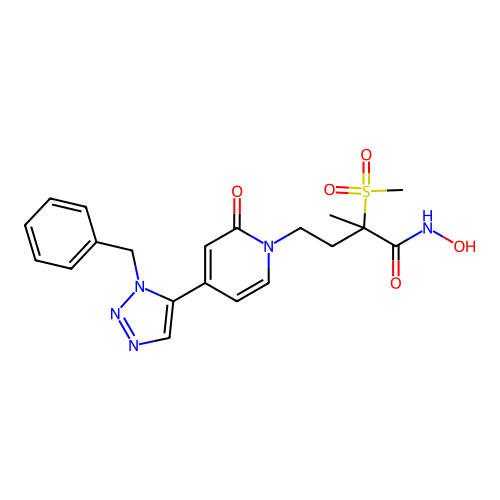
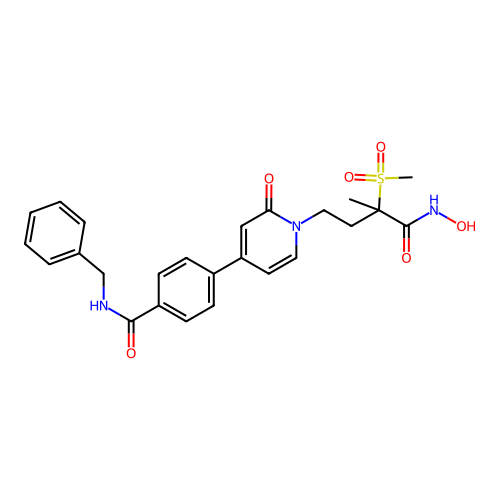
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataKi: 0.00320nMAssay Description:A fluorescence-based competition assay was used to determine the kinetic parameters for binding of the inhibitors to paLpxC. This assay is based on t...More data for this Ligand-Target Pair
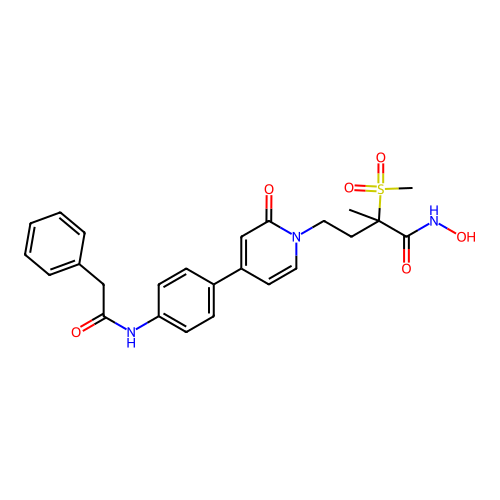
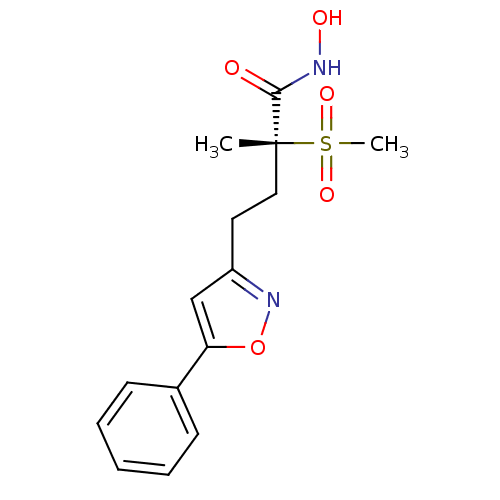
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataKi: 0.00370nMAssay Description:A fluorescence-based competition assay was used to determine the kinetic parameters for binding of the inhibitors to paLpxC. This assay is based on t...More data for this Ligand-Target Pair
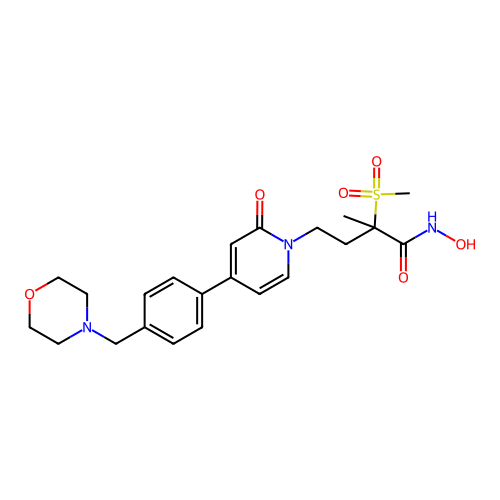
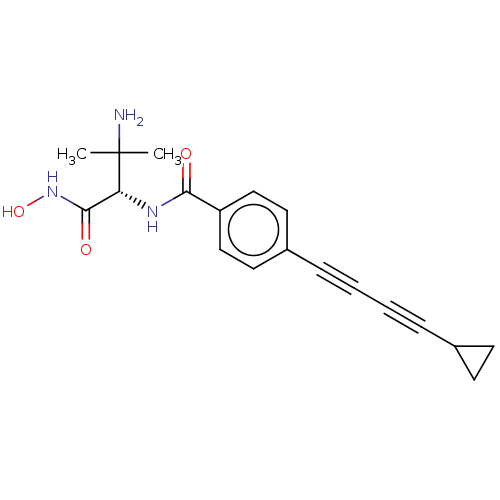
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataKi: 0.00400nMAssay Description:A fluorescence-based competition assay was used to determine the kinetic parameters for binding of the inhibitors to paLpxC. This assay is based on t...More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataKi: 0.00420nMAssay Description:A fluorescence-based competition assay was used to determine the kinetic parameters for binding of the inhibitors to paLpxC. This assay is based on t...More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataKi: 0.00550nMAssay Description:A fluorescence-based competition assay was used to determine the kinetic parameters for binding of the inhibitors to paLpxC. This assay is based on t...More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataKi: 0.00550nMAssay Description:A fluorescence-based competition assay was used to determine the kinetic parameters for binding of the inhibitors to paLpxC. This assay is based on t...More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataKi: 0.00610nMAssay Description:A fluorescence-based competition assay was used to determine the kinetic parameters for binding of the inhibitors to paLpxC. This assay is based on t...More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataKi: 0.00700nMAssay Description:A fluorescence-based competition assay was used to determine the kinetic parameters for binding of the inhibitors to paLpxC. This assay is based on t...More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataKi: 0.00800nMAssay Description:A fluorescence-based competition assay was used to determine the kinetic parameters for binding of the inhibitors to paLpxC. This assay is based on t...More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataKi: 0.0100nMAssay Description:A fluorescence-based competition assay was used to determine the kinetic parameters for binding of the inhibitors to paLpxC. This assay is based on t...More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataKi: 0.0102nMAssay Description:A fluorescence-based competition assay was used to determine the kinetic parameters for binding of the inhibitors to paLpxC. This assay is based on t...More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataKi: 0.0106nMAssay Description:A fluorescence-based competition assay was used to determine the kinetic parameters for binding of the inhibitors to paLpxC. This assay is based on t...More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataKi: 0.0126nMAssay Description:A fluorescence-based competition assay was used to determine the kinetic parameters for binding of the inhibitors to paLpxC. This assay is based on t...More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataKi: 0.0144nMAssay Description:A fluorescence-based competition assay was used to determine the kinetic parameters for binding of the inhibitors to paLpxC. This assay is based on t...More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataKi: 0.0150nMAssay Description:A fluorescence-based competition assay was used to determine the kinetic parameters for binding of the inhibitors to paLpxC. This assay is based on t...More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataKi: 0.0379nMAssay Description:A fluorescence-based competition assay was used to determine the kinetic parameters for binding of the inhibitors to paLpxC. This assay is based on t...More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataKi: 0.0677nMAssay Description:A fluorescence-based competition assay was used to determine the kinetic parameters for binding of the inhibitors to paLpxC. This assay is based on t...More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataKi: 0.0954nMAssay Description:A fluorescence-based competition assay was used to determine the kinetic parameters for binding of the inhibitors to paLpxC. This assay is based on t...More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataKi: 0.0968nMAssay Description:A fluorescence-based competition assay was used to determine the kinetic parameters for binding of the inhibitors to paLpxC. This assay is based on t...More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataKi: 0.213nMAssay Description:A fluorescence-based competition assay was used to determine the kinetic parameters for binding of the inhibitors to paLpxC. This assay is based on t...More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataKi: 0.274nMAssay Description:A fluorescence-based competition assay was used to determine the kinetic parameters for binding of the inhibitors to paLpxC. This assay is based on t...More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataIC50: 0.511nMAssay Description:Inhibition of Pseudomonas aeruginosa LpxCMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataIC50: 0.620nMAssay Description:Inhibition of recombinant Pseudomonas aeruginosa LpxC expressed in Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataIC50: 0.657nMAssay Description:Inhibition of Pseudomonas aeruginosa LpxCMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataIC50: 0.680nMAssay Description:Inhibition of recombinant Pseudomonas aeruginosa LpxC expressed in Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataIC50: 0.710nMAssay Description:Inhibition of recombinant Pseudomonas aeruginosa LpxC expressed in Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataIC50: 0.850nMAssay Description:Inhibition of Pseudomonas aeruginosa LpxCMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataIC50: 0.912nMAssay Description:Inhibition of Pseudomonas aeruginosa LpxCMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataIC50: 1.20nMAssay Description:Inhibition of Pseudomonas aeruginosa LpxCMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataIC50: 1.39nMAssay Description:Inhibition of Pseudomonas aeruginosa LpxCMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataIC50: 1.61nMAssay Description:Inhibition of Pseudomonas aeruginosa LpxCMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataIC50: 1.67nMAssay Description:Inhibition of Pseudomonas aeruginosa LpxCMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataIC50: 1.87nMAssay Description:Inhibition of Pseudomonas aeruginosa LpxCMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataIC50: 1.95nMAssay Description:Inhibition of Pseudomonas aeruginosa LpxCMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataIC50: 2.32nMAssay Description:Inhibition of Pseudomonas aeruginosa LpxCMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataIC50: 2.66nMAssay Description:Inhibition of Pseudomonas aeruginosa LpxCMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataIC50: 2.93nMAssay Description:Inhibition of Pseudomonas aeruginosa LpxCMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataIC50: 3nMAssay Description:Inhibition of Pseudomonas aeruginosa LpxCMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataIC50: 3.02nMAssay Description:Inhibition of Pseudomonas aeruginosa LpxCMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataIC50: 4.40nMAssay Description:Inhibition of Pseudomonas aeruginosa LpxCMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataIC50: 4.71nMAssay Description:Inhibition of Pseudomonas aeruginosa LpxCMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataIC50: 5.32nMAssay Description:Inhibition of Pseudomonas aeruginosa LpxCMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataIC50: 6.75nMAssay Description:Inhibition of Pseudomonas aeruginosa LpxCMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataIC50: 6.81nMAssay Description:Inhibition of Pseudomonas aeruginosa LpxCMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataIC50: 8.08nMAssay Description:Inhibition of Pseudomonas aeruginosa LpxCMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataIC50: 8.31nMAssay Description:Inhibition of Pseudomonas aeruginosa LpxCMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataIC50: 12nMAssay Description:Inhibition of recombinant Pseudomonas aeruginosa LpxC expressed in Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataIC50: 13.5nMAssay Description:Inhibition of Pseudomonas aeruginosa LpxCMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
The State University of New York
US Patent
The State University of New York
US Patent
Affinity DataIC50: 19nMAssay Description:Inhibition of recombinant Pseudomonas aeruginosa LpxC expressed in Escherichia coliMore data for this Ligand-Target Pair
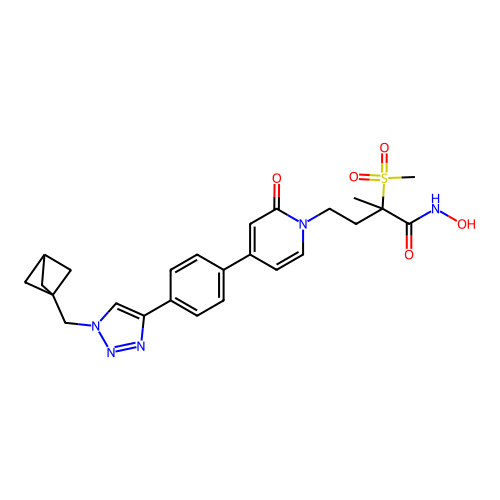
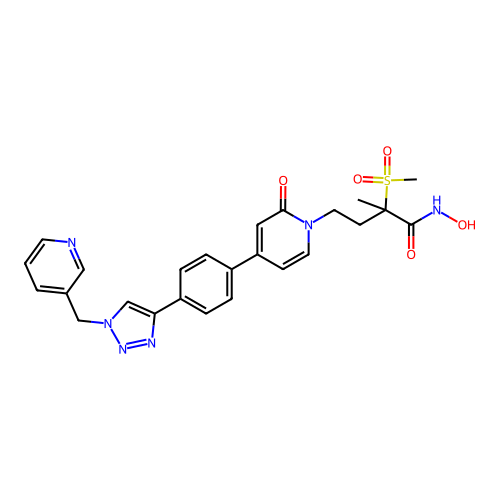
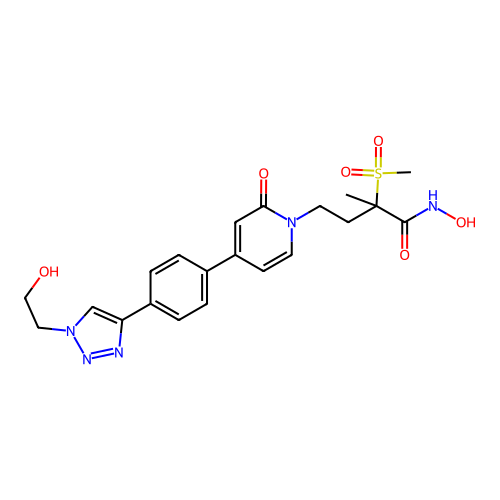
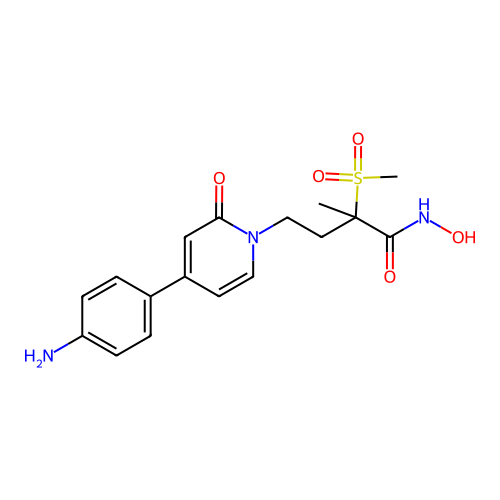
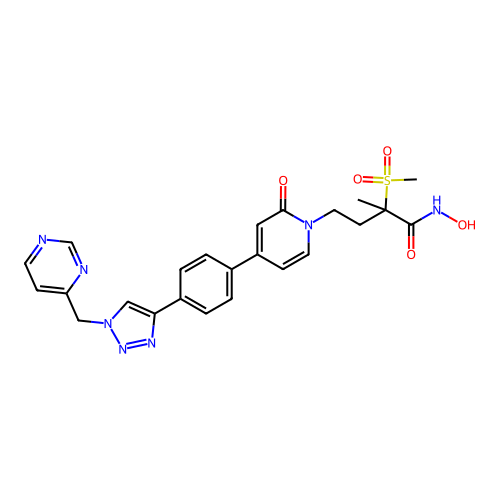
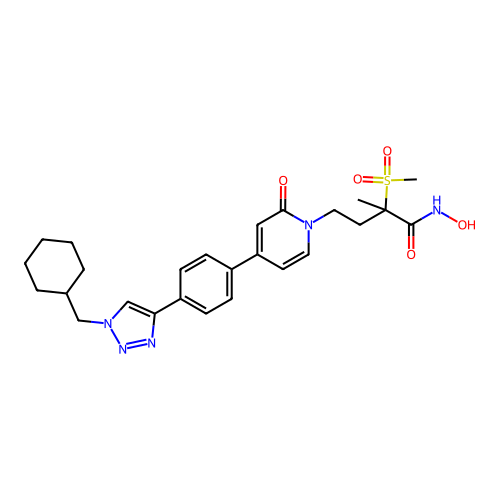
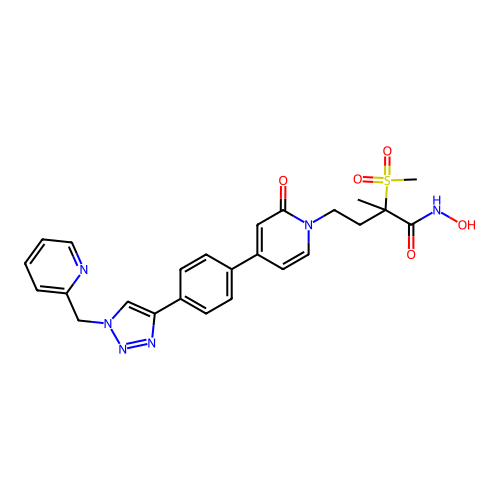
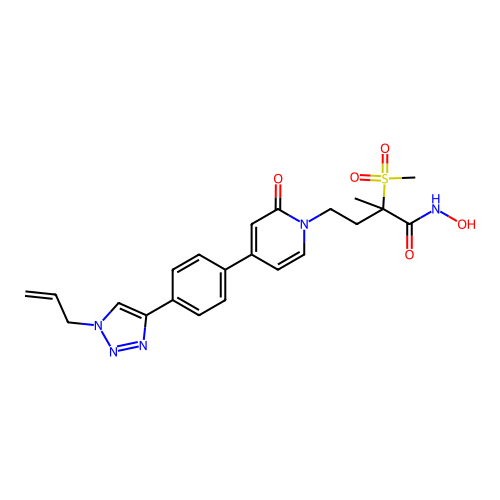
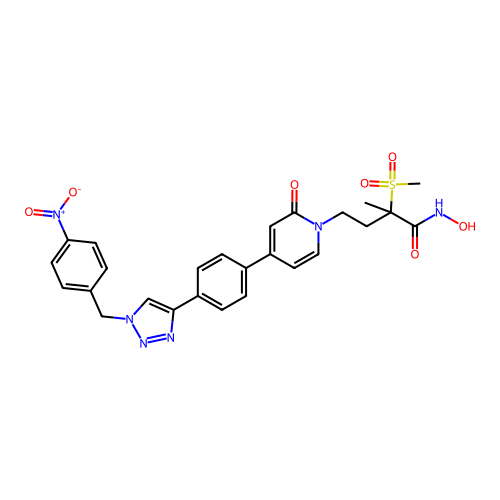
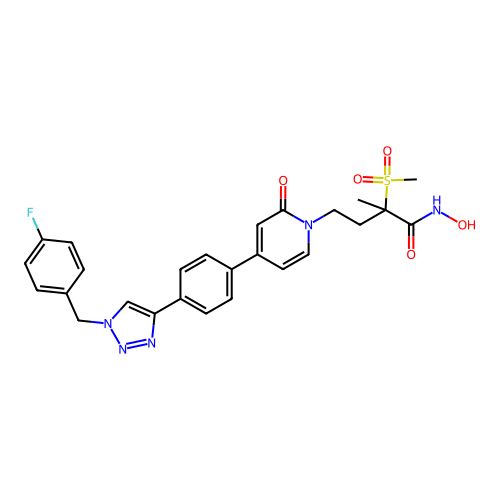
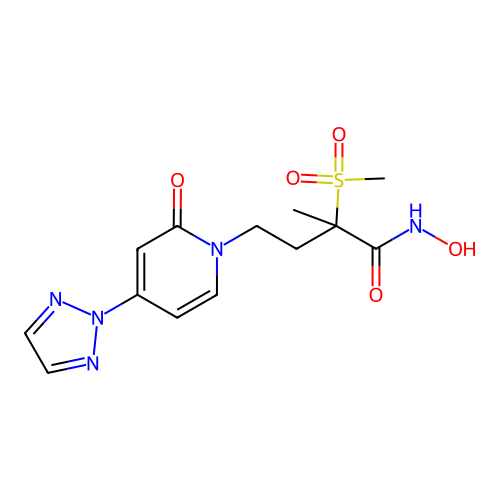
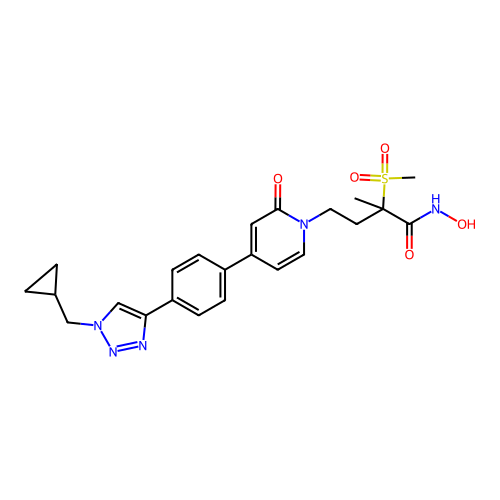
 3D Structure (crystal)
3D Structure (crystal)