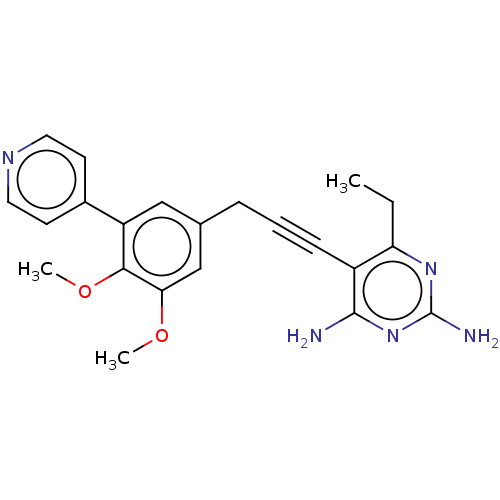
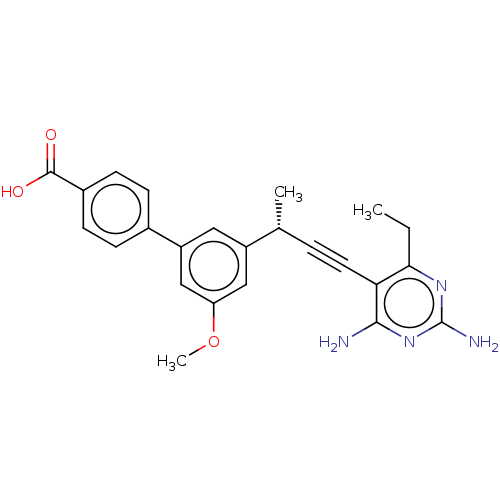
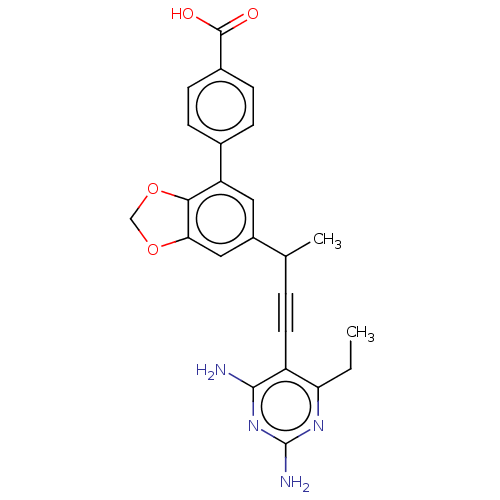
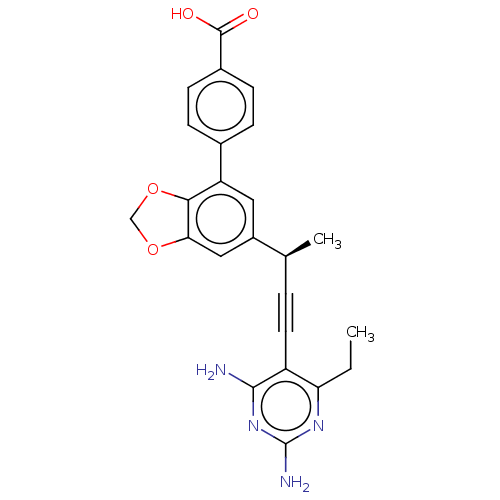
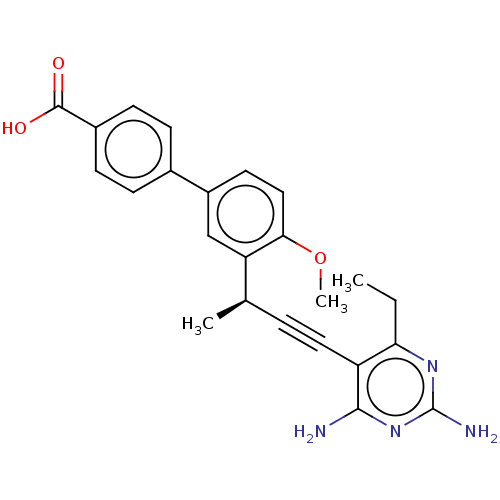
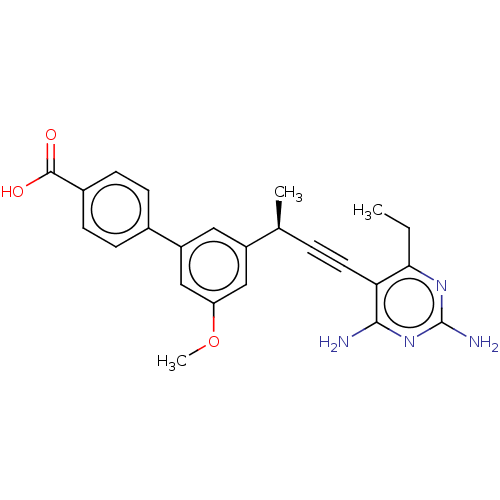
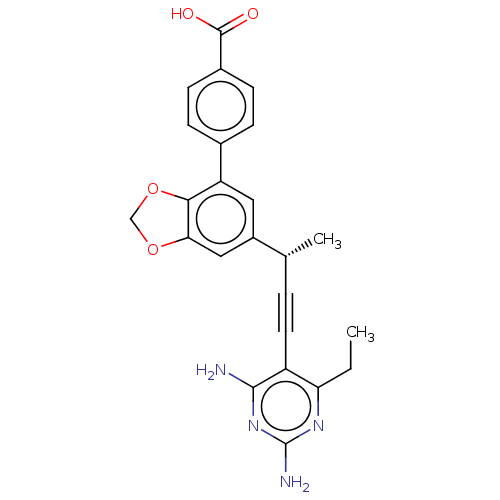
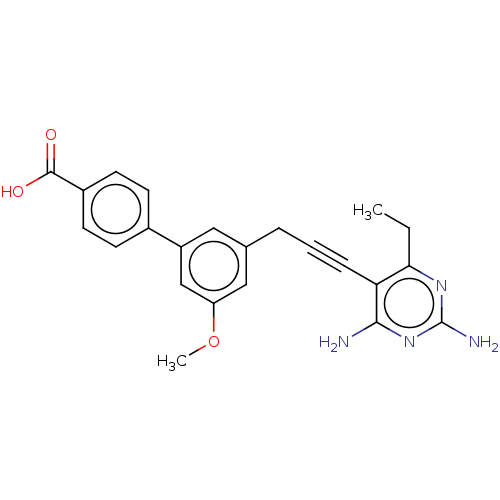
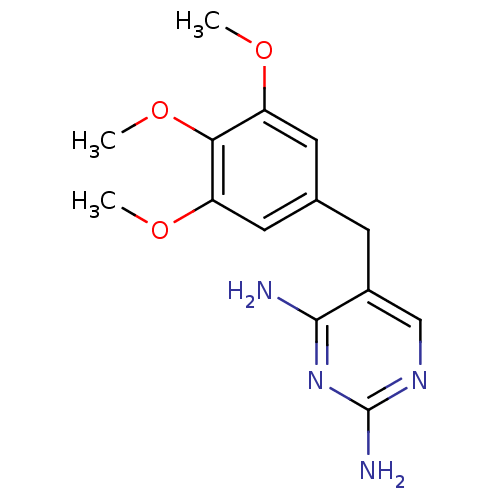
Affinity DataIC50: 22nMAssay Description:IC50 values were determined following a standard method that has been described previously (Reeve et al., 2014, 2016).More data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:IC50 values were determined following a standard method that has been described previously (Reeve et al., 2014, 2016).More data for this Ligand-Target Pair
Affinity DataIC50: 41nMAssay Description:IC50 values were determined following a standard method that has been described previously (Reeve et al., 2014, 2016).More data for this Ligand-Target Pair
Affinity DataIC50: 54nMAssay Description:IC50 values were determined following a standard method that has been described previously (Reeve et al., 2014, 2016).More data for this Ligand-Target Pair
Affinity DataIC50: 73nMAssay Description:IC50 values were determined following a standard method that has been described previously (Reeve et al., 2014, 2016).More data for this Ligand-Target Pair
Affinity DataIC50: 91nMAssay Description:IC50 values were determined following a standard method that has been described previously (Reeve et al., 2014, 2016).More data for this Ligand-Target Pair
Affinity DataIC50: 100nMAssay Description:IC50 values were determined following a standard method that has been described previously (Reeve et al., 2014, 2016).More data for this Ligand-Target Pair
Affinity DataIC50: 170nMAssay Description:IC50 values were determined following a standard method that has been described previously (Reeve et al., 2014, 2016).More data for this Ligand-Target Pair
Affinity DataIC50: 4.30E+4nMAssay Description:IC50 values were determined following a standard method that has been described previously (Reeve et al., 2014, 2016).More data for this Ligand-Target Pair