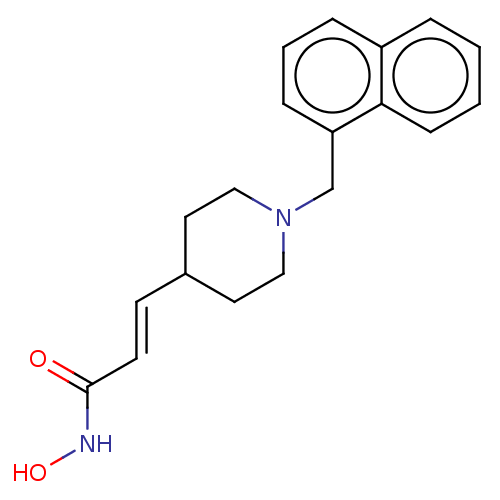
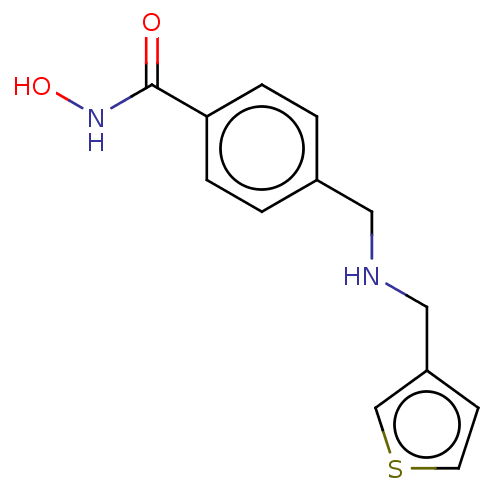
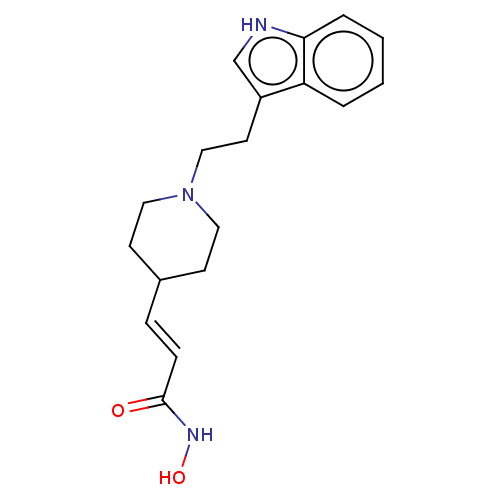
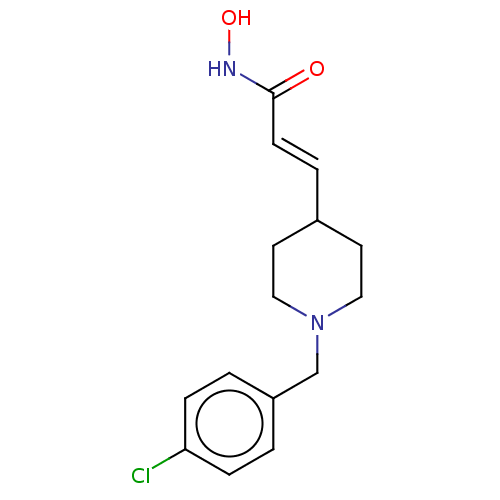
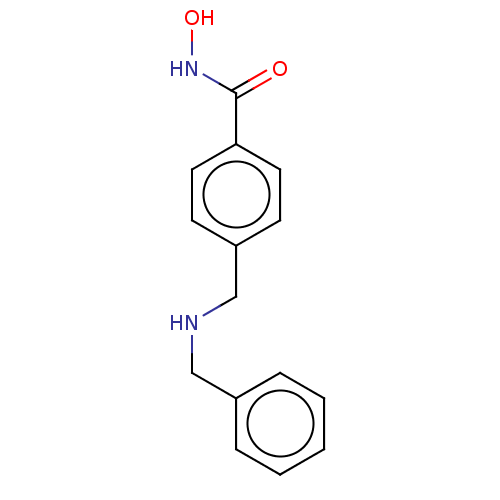
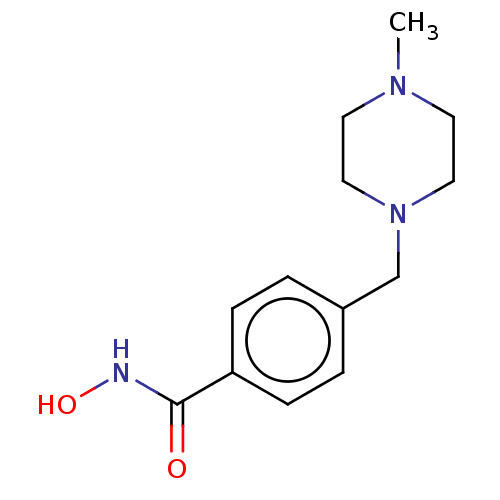
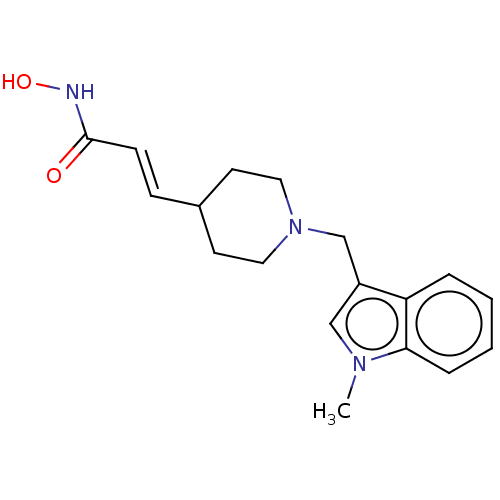
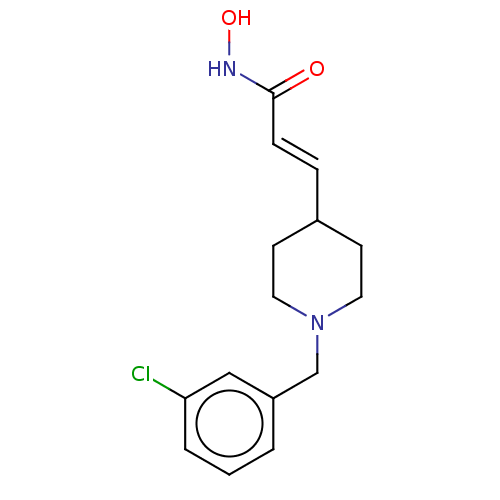
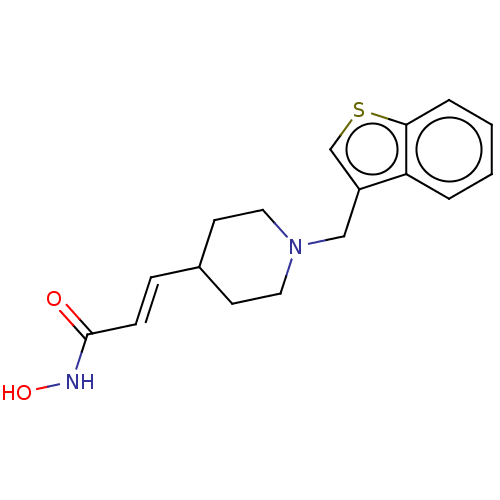
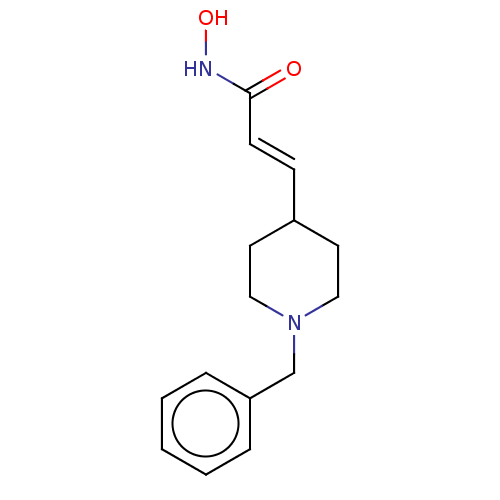
TargetPolyamine deacetylase HDAC10(Zebrafish)
Martin-Luther University of Halle-Wittenberg
Curated by ChEMBL
Martin-Luther University of Halle-Wittenberg
Curated by ChEMBL
Affinity DataIC50: 11nMAssay Description:Inhibition of zebra fish HDAC10 using Ac-spermidine-AMC as substrate incubated for 25 mins and measured by fluorescence assayMore data for this Ligand-Target Pair
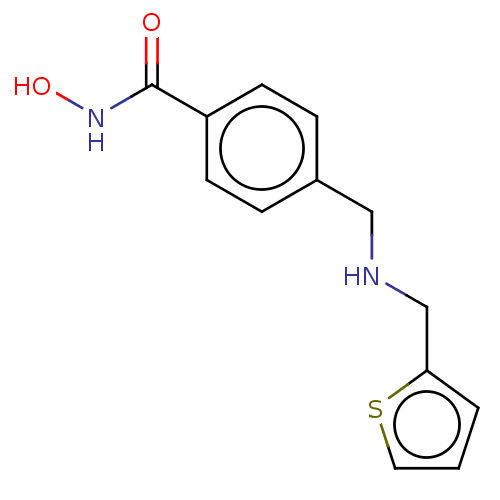
TargetPolyamine deacetylase HDAC10(Zebrafish)
Martin-Luther University of Halle-Wittenberg
Curated by ChEMBL
Martin-Luther University of Halle-Wittenberg
Curated by ChEMBL
Affinity DataIC50: 20nMAssay Description:Inhibition of zebra fish HDAC10 using Ac-spermidine-AMC as substrate incubated for 25 mins and measured by fluorescence assayMore data for this Ligand-Target Pair
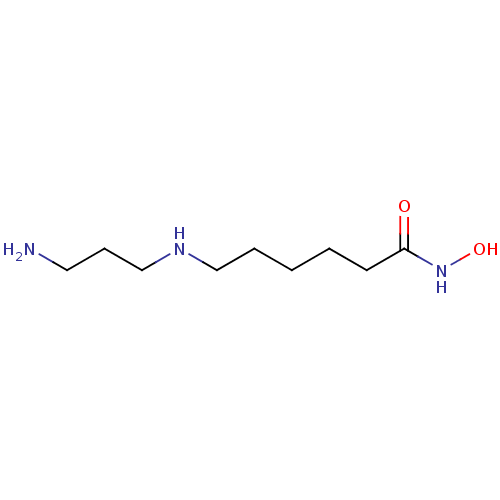
TargetPolyamine deacetylase HDAC10(Zebrafish)
Martin-Luther University of Halle-Wittenberg
Curated by ChEMBL
Martin-Luther University of Halle-Wittenberg
Curated by ChEMBL
Affinity DataIC50: 24nMAssay Description:Inhibition of zebra fish HDAC10 using Ac-spermidine-AMC as substrate incubated for 25 mins and measured by fluorescence assayMore data for this Ligand-Target Pair
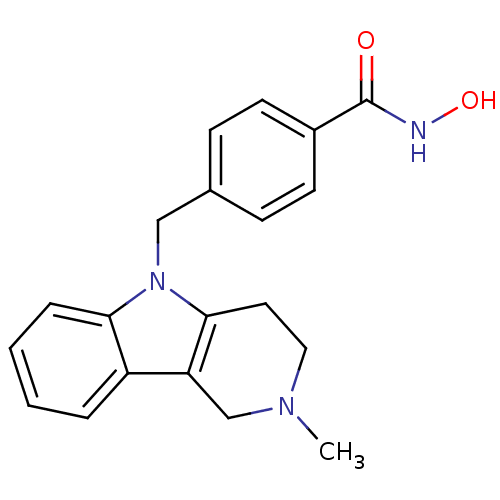
TargetPolyamine deacetylase HDAC10(Zebrafish)
Martin-Luther University of Halle-Wittenberg
Curated by ChEMBL
Martin-Luther University of Halle-Wittenberg
Curated by ChEMBL
Affinity DataIC50: 29nMAssay Description:Inhibition of zebra fish HDAC10 using Ac-spermidine-AMC as substrate incubated for 25 mins and measured by fluorescence assayMore data for this Ligand-Target Pair
TargetPolyamine deacetylase HDAC10(Zebrafish)
Martin-Luther University of Halle-Wittenberg
Curated by ChEMBL
Martin-Luther University of Halle-Wittenberg
Curated by ChEMBL
Affinity DataIC50: 33nMAssay Description:Inhibition of zebra fish HDAC10 using Ac-spermidine-AMC as substrate incubated for 25 mins and measured by fluorescence assayMore data for this Ligand-Target Pair
TargetPolyamine deacetylase HDAC10(Zebrafish)
Martin-Luther University of Halle-Wittenberg
Curated by ChEMBL
Martin-Luther University of Halle-Wittenberg
Curated by ChEMBL
Affinity DataIC50: 37nMAssay Description:Inhibition of zebra fish HDAC10 using Ac-spermidine-AMC as substrate incubated for 25 mins and measured by fluorescence assayMore data for this Ligand-Target Pair
TargetPolyamine deacetylase HDAC10(Zebrafish)
Martin-Luther University of Halle-Wittenberg
Curated by ChEMBL
Martin-Luther University of Halle-Wittenberg
Curated by ChEMBL
Affinity DataIC50: 43nMAssay Description:Inhibition of zebra fish HDAC10 using Ac-spermidine-AMC as substrate incubated for 25 mins and measured by fluorescence assayMore data for this Ligand-Target Pair
TargetPolyamine deacetylase HDAC10(Zebrafish)
Martin-Luther University of Halle-Wittenberg
Curated by ChEMBL
Martin-Luther University of Halle-Wittenberg
Curated by ChEMBL
Affinity DataIC50: 58nMAssay Description:Inhibition of zebra fish HDAC10 using Ac-spermidine-AMC as substrate incubated for 25 mins and measured by fluorescence assayMore data for this Ligand-Target Pair
TargetPolyamine deacetylase HDAC10(Zebrafish)
Martin-Luther University of Halle-Wittenberg
Curated by ChEMBL
Martin-Luther University of Halle-Wittenberg
Curated by ChEMBL
Affinity DataIC50: 60nMAssay Description:Inhibition of zebra fish HDAC10 using Ac-spermidine-AMC as substrate incubated for 25 mins and measured by fluorescence assayMore data for this Ligand-Target Pair
TargetPolyamine deacetylase HDAC10(Zebrafish)
Martin-Luther University of Halle-Wittenberg
Curated by ChEMBL
Martin-Luther University of Halle-Wittenberg
Curated by ChEMBL
Affinity DataIC50: 62nMAssay Description:Inhibition of zebra fish HDAC10 using Ac-spermidine-AMC as substrate incubated for 25 mins and measured by fluorescence assayMore data for this Ligand-Target Pair
TargetPolyamine deacetylase HDAC10(Zebrafish)
Martin-Luther University of Halle-Wittenberg
Curated by ChEMBL
Martin-Luther University of Halle-Wittenberg
Curated by ChEMBL
Affinity DataIC50: 106nMAssay Description:Inhibition of zebra fish HDAC10 using Ac-spermidine-AMC as substrate incubated for 25 mins and measured by fluorescence assayMore data for this Ligand-Target Pair
TargetPolyamine deacetylase HDAC10(Zebrafish)
Martin-Luther University of Halle-Wittenberg
Curated by ChEMBL
Martin-Luther University of Halle-Wittenberg
Curated by ChEMBL
Affinity DataIC50: 120nMAssay Description:Inhibition of zebrafish HDAC10 (2 to 675 residues) expressed in Escherichia coli BL21(DE3) cells using N-acetylputrescine as substrate preincubated f...More data for this Ligand-Target Pair
TargetPolyamine deacetylase HDAC10(Zebrafish)
Martin-Luther University of Halle-Wittenberg
Curated by ChEMBL
Martin-Luther University of Halle-Wittenberg
Curated by ChEMBL
Affinity DataIC50: 220nMAssay Description:Inhibition of zebra fish HDAC10 using Ac-spermidine-AMC as substrate incubated for 25 mins and measured by fluorescence assayMore data for this Ligand-Target Pair
TargetPolyamine deacetylase HDAC10(Zebrafish)
Martin-Luther University of Halle-Wittenberg
Curated by ChEMBL
Martin-Luther University of Halle-Wittenberg
Curated by ChEMBL
Affinity DataIC50: 530nMAssay Description:Inhibition of zebra fish HDAC10 using Ac-spermidine-AMC as substrate incubated for 25 mins and measured by fluorescence assayMore data for this Ligand-Target Pair
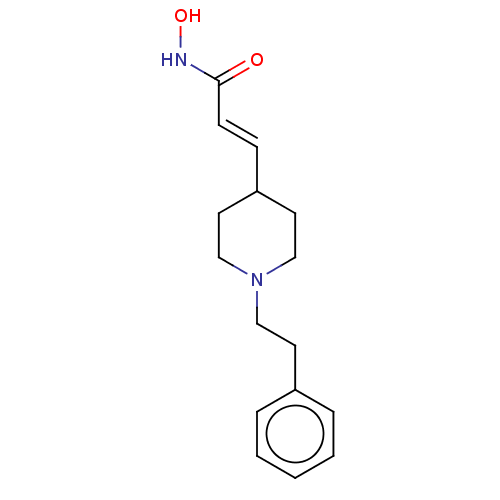
 3D Structure (crystal)
3D Structure (crystal)