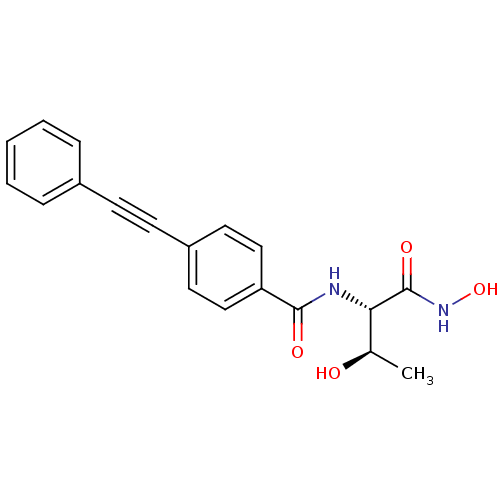
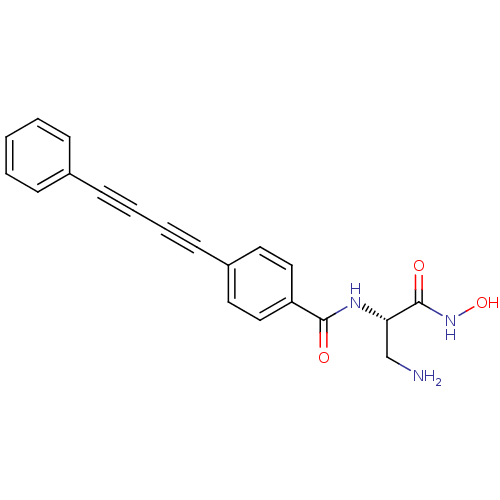
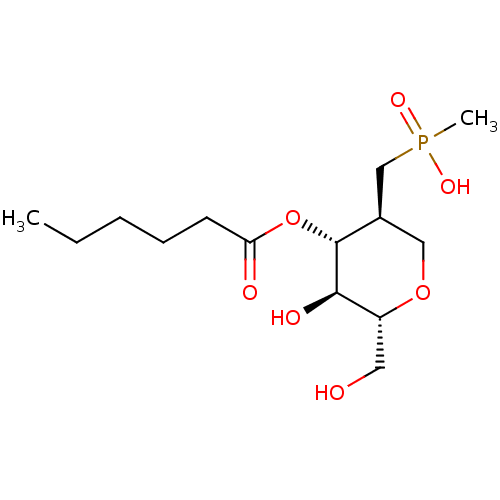
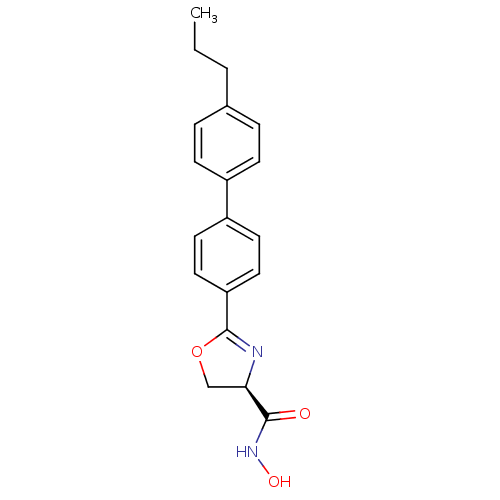
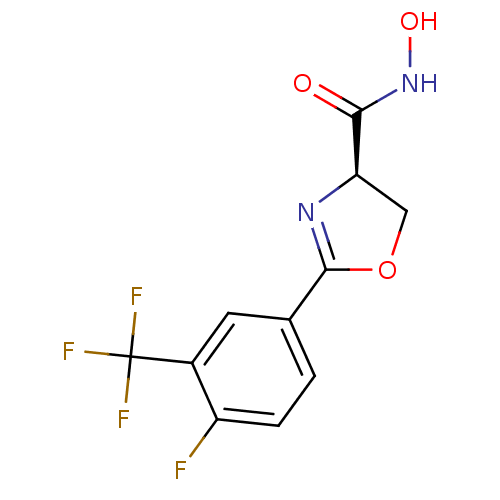
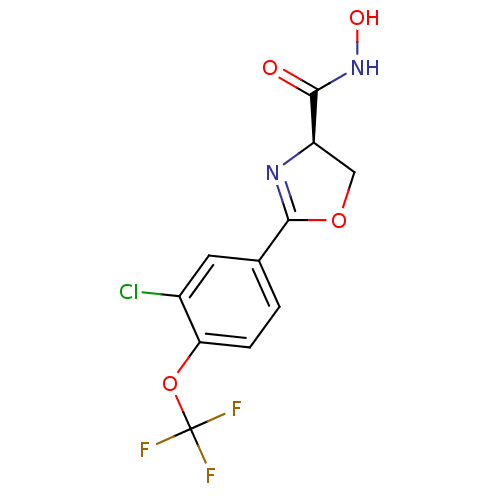
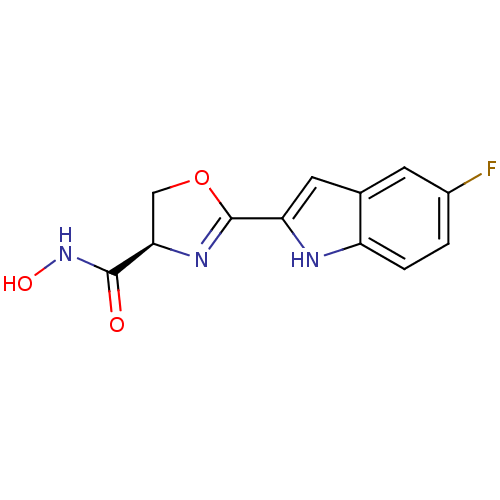
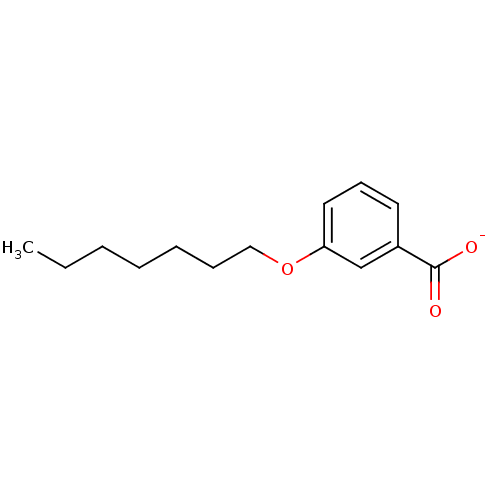
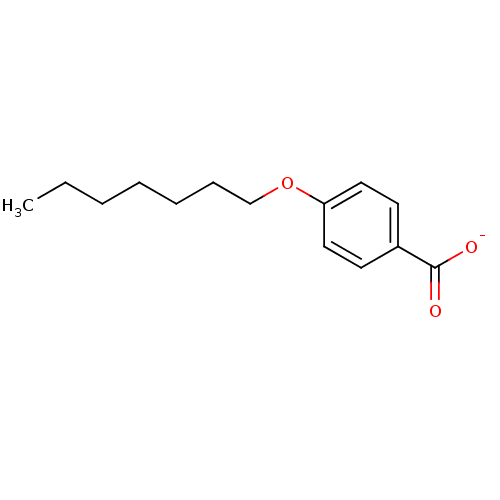
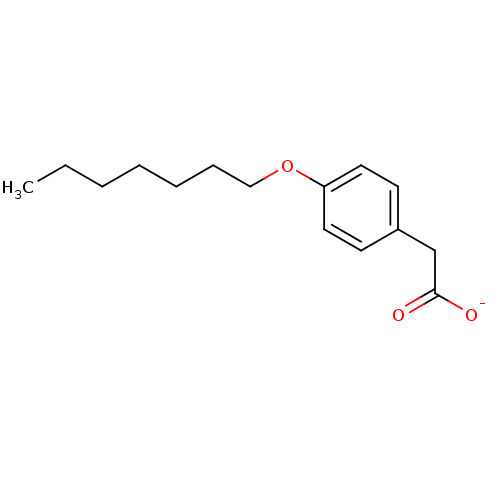
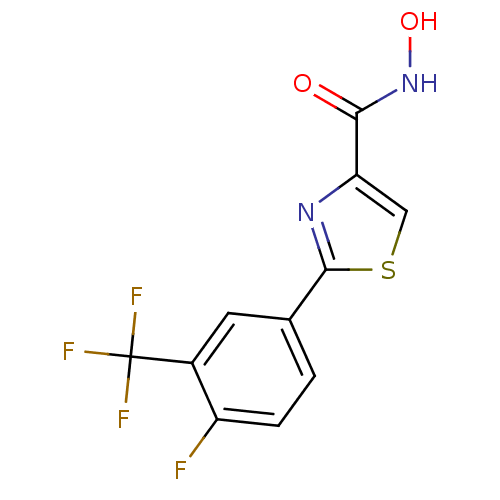
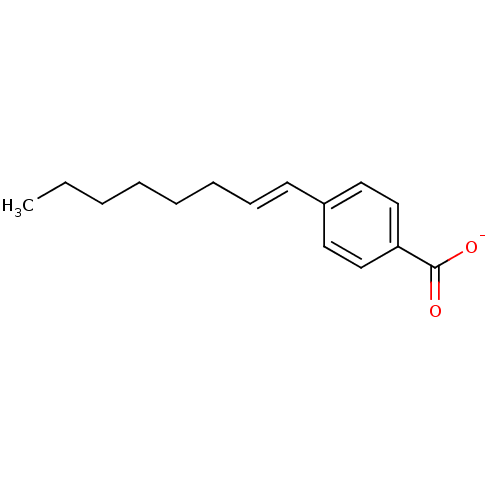
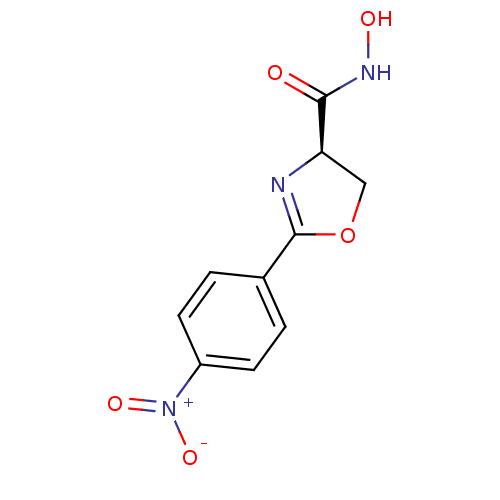
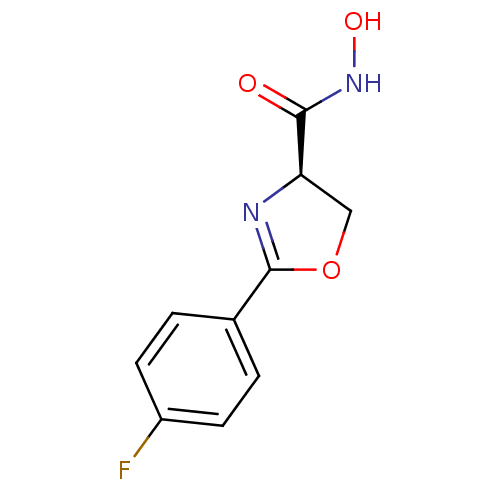
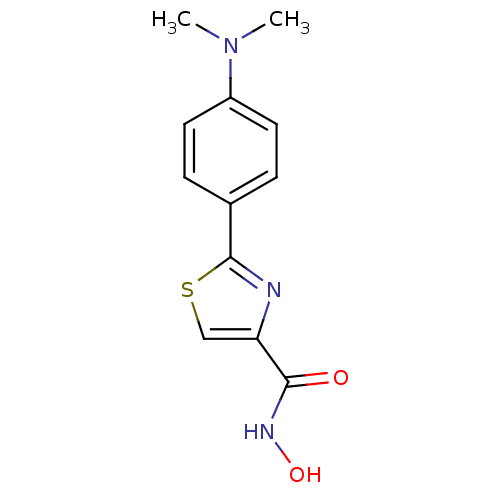
Affinity DataKi: 1nM ΔG°: -51.4kJ/molepH: 7.0 T: 2°CAssay Description:OctetRedMore data for this Ligand-Target Pair
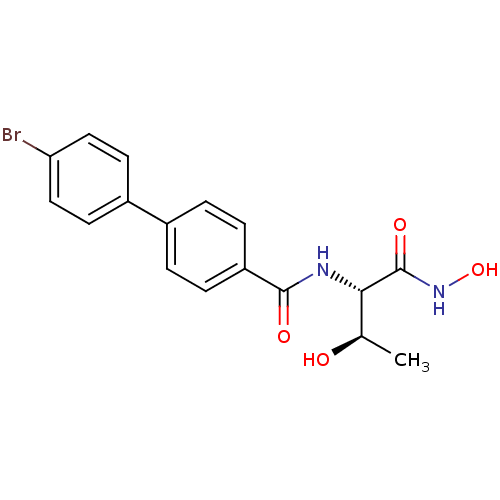
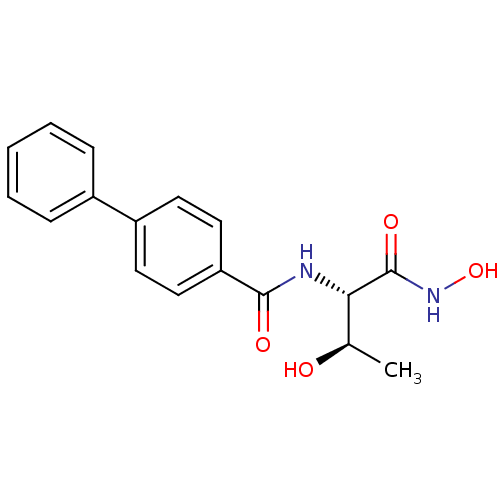
Affinity DataKi: 1nM ΔG°: -51.4kJ/molepH: 7.0 T: 2°CAssay Description:OctetRedMore data for this Ligand-Target Pair
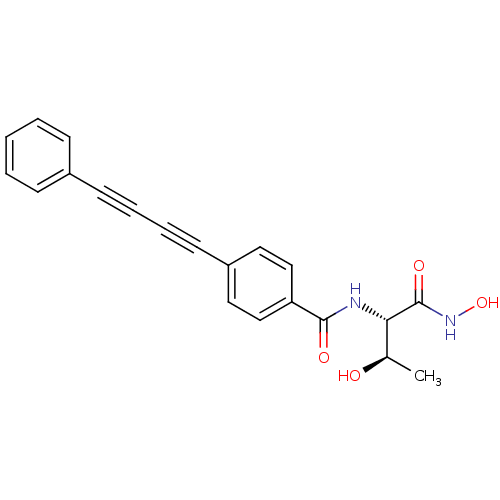
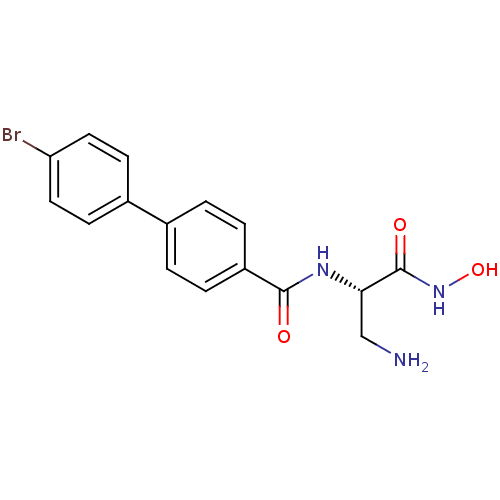
Affinity DataKi: 1nM ΔG°: -51.4kJ/molepH: 7.0 T: 2°CAssay Description:OctetRedMore data for this Ligand-Target Pair
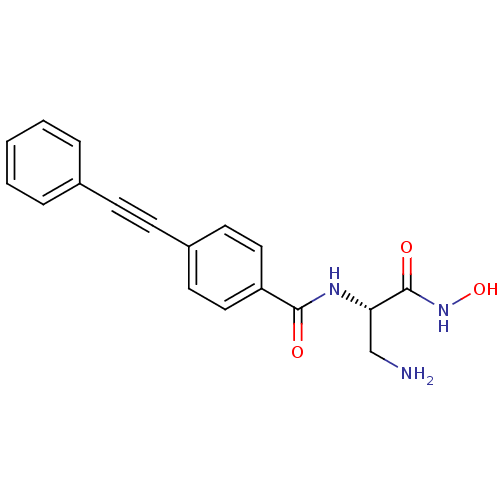
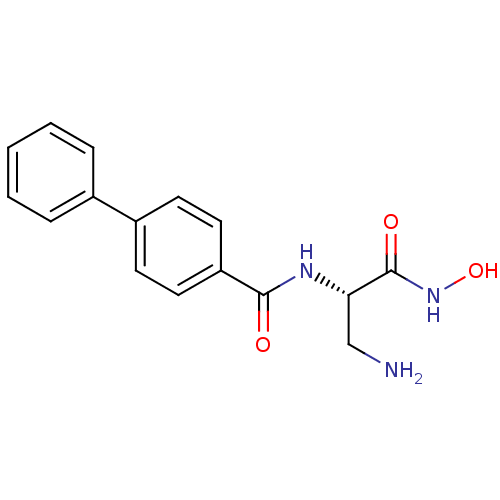
Affinity DataKd: 8nMpH: 7.0 T: 2°CAssay Description:ThermofluorMore data for this Ligand-Target Pair
Affinity DataKi: 10nM ΔG°: -45.7kJ/molepH: 7.0 T: 2°CAssay Description:OctetRedMore data for this Ligand-Target Pair
Affinity DataKi: 10nM ΔG°: -45.7kJ/molepH: 7.0 T: 2°CAssay Description:OctetRedMore data for this Ligand-Target Pair
Affinity DataKd: 20nMpH: 7.0 T: 2°CAssay Description:ThermofluorMore data for this Ligand-Target Pair
Affinity DataKd: 33nMpH: 7.0 T: 2°CAssay Description:ThermofluorMore data for this Ligand-Target Pair
Affinity DataKd: 59nMpH: 7.0 T: 2°CAssay Description:ThermofluorMore data for this Ligand-Target Pair
Affinity DataKd: 75nMpH: 7.0 T: 2°CAssay Description:ThermofluorMore data for this Ligand-Target Pair
Affinity DataKd: 108nMpH: 7.0 T: 2°CAssay Description:ThermofluorMore data for this Ligand-Target Pair
Affinity DataKd: 190nMpH: 7.0 T: 2°CAssay Description:ThermofluorMore data for this Ligand-Target Pair
Affinity DataIC50: 192nMAssay Description:Inhibition of Aquifex aeolicus LpxC transformed in Escherichia coli BL21(DE3) pLysS cellsMore data for this Ligand-Target Pair
Affinity DataKd: 393nMpH: 7.0 T: 2°CAssay Description:ThermofluorMore data for this Ligand-Target Pair
Affinity DataKd: 900nMpH: 7.0 T: 2°CAssay Description:Ligand binding to A. aeolicus LpxC was assayed by isothermal titration using isothermal microcalorimeter from Microcal, Inc. (Northampton, MA). The e...More data for this Ligand-Target Pair
Affinity DataKd: 1.03E+3nMpH: 7.0 T: 2°CAssay Description:ThermofluorMore data for this Ligand-Target Pair
Affinity DataKd: 1.82E+3nMpH: 7.0 T: 2°CAssay Description:OctetRedMore data for this Ligand-Target Pair
Affinity DataKd: 2.00E+3nMpH: 7.0 T: 2°CAssay Description:Ligand binding to A. aeolicus LpxC was assayed by isothermal titration using isothermal microcalorimeter from Microcal, Inc. (Northampton, MA). The e...More data for this Ligand-Target Pair
Affinity DataKd: 2.30E+3nMpH: 7.0 T: 2°CAssay Description:Ligand binding to A. aeolicus LpxC was assayed by isothermal titration using isothermal microcalorimeter from Microcal, Inc. (Northampton, MA). The e...More data for this Ligand-Target Pair
Affinity DataKd: 2.50E+3nMpH: 7.0 T: 2°CAssay Description:ThermofluorMore data for this Ligand-Target Pair
Affinity DataKd: 2.60E+3nMpH: 7.0 T: 2°CAssay Description:Ligand binding to A. aeolicus LpxC was assayed by isothermal titration using isothermal microcalorimeter from Microcal, Inc. (Northampton, MA). The e...More data for this Ligand-Target Pair
Affinity DataKd: 2.91E+3nMpH: 7.0 T: 2°CAssay Description:ThermofluorMore data for this Ligand-Target Pair
Affinity DataKd: 3.80E+3nMpH: 7.0 T: 2°CAssay Description:ThermofluorMore data for this Ligand-Target Pair
Affinity DataKd: 6.20E+3nMpH: 7.0 T: 2°CAssay Description:Ligand binding to A. aeolicus LpxC was assayed by isothermal titration using isothermal microcalorimeter from Microcal, Inc. (Northampton, MA). The e...More data for this Ligand-Target Pair
Affinity DataKd: 6.40E+3nMpH: 7.0 T: 2°CAssay Description:Ligand binding to A. aeolicus LpxC was assayed by isothermal titration using isothermal microcalorimeter from Microcal, Inc. (Northampton, MA). The e...More data for this Ligand-Target Pair
Affinity DataKd: 7.00E+3nMpH: 7.0 T: 2°CAssay Description:Ligand binding to A. aeolicus LpxC was assayed by isothermal titration using isothermal microcalorimeter from Microcal, Inc. (Northampton, MA). The e...More data for this Ligand-Target Pair
Affinity DataKd: 7.25E+3nMpH: 7.0 T: 2°CAssay Description:ThermofluorMore data for this Ligand-Target Pair
Affinity DataKd: 8.60E+3nMpH: 7.0 T: 2°CAssay Description:Ligand binding to A. aeolicus LpxC was assayed by isothermal titration using isothermal microcalorimeter from Microcal, Inc. (Northampton, MA). The e...More data for this Ligand-Target Pair
Affinity DataKd: 9.00E+3nMpH: 7.0 T: 2°CAssay Description:Ligand binding to A. aeolicus LpxC was assayed by isothermal titration using isothermal microcalorimeter from Microcal, Inc. (Northampton, MA). The e...More data for this Ligand-Target Pair
Affinity DataKd: 1.11E+4nMpH: 7.0 T: 2°CAssay Description:ThermofluorMore data for this Ligand-Target Pair
Affinity DataKd: 1.56E+4nMpH: 7.0 T: 2°CAssay Description:ThermofluorMore data for this Ligand-Target Pair
Affinity DataKd: 2.15E+4nMpH: 7.0 T: 2°CAssay Description:ThermofluorMore data for this Ligand-Target Pair
Affinity DataKd: 2.82E+4nMpH: 7.0 T: 2°CAssay Description:ThermofluorMore data for this Ligand-Target Pair