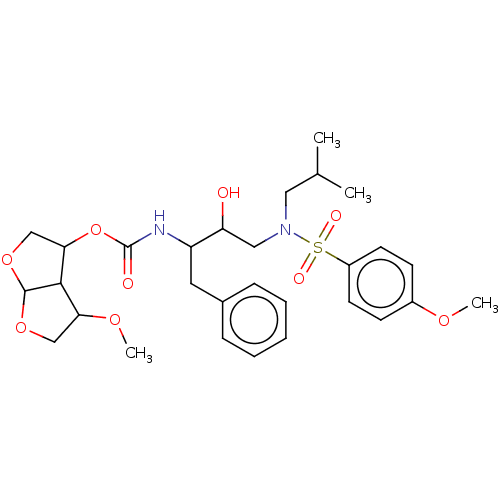
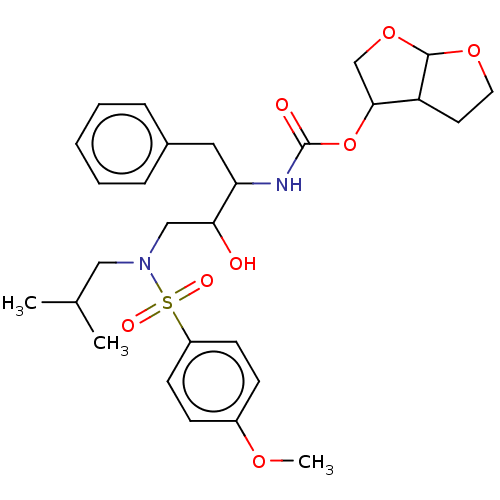
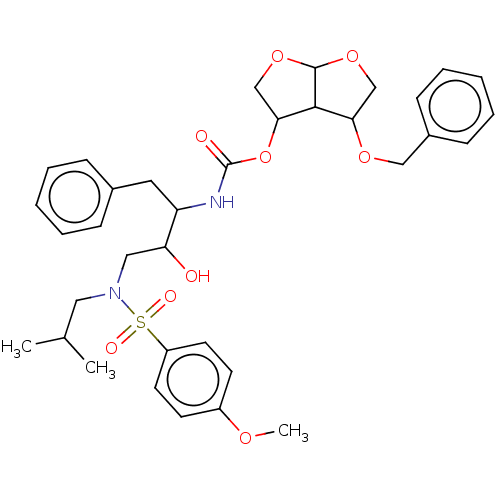
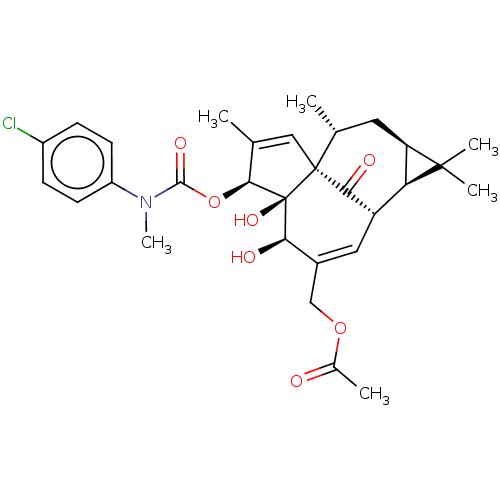
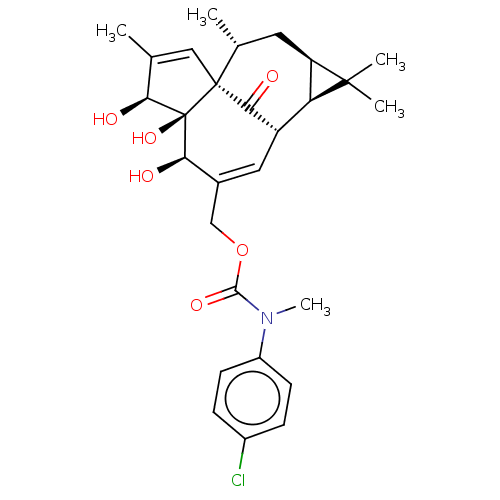
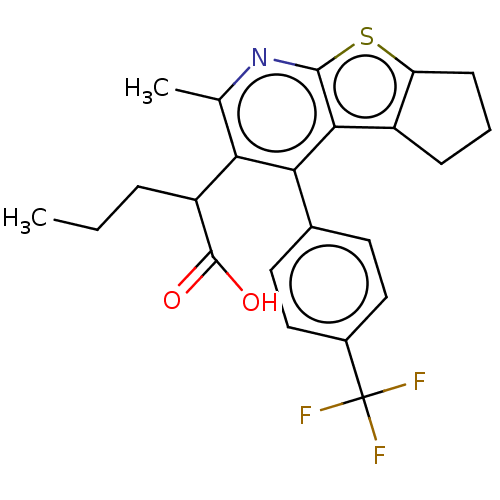
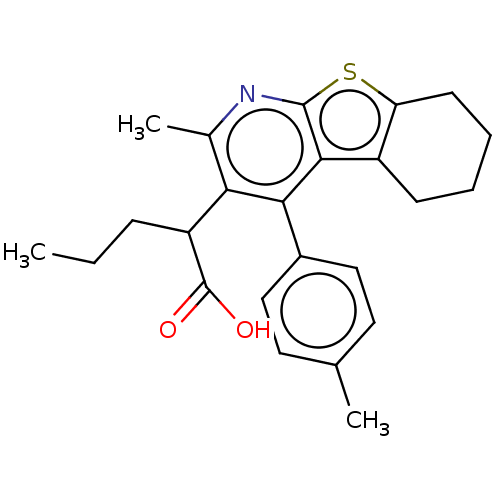
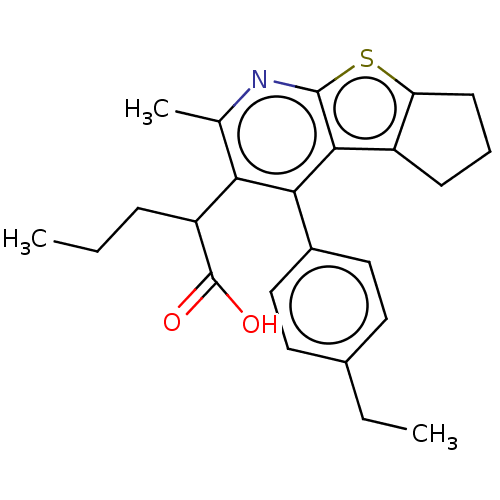
Affinity DataKi: 0.00290nMAssay Description:The enzyme inhibitory activity (Ki) was determined according to an assay protocol reported by Toth and Marshall (Toth, M. V.; Marshall, G. R. Int. J....More data for this Ligand-Target Pair
Affinity DataKi: 0.0140nMAssay Description:The enzyme inhibitory activity (Ki) was determined according to an assay protocol reported by Toth and Marshall (Toth, M. V.; Marshall, G. R. Int. J....More data for this Ligand-Target Pair
Affinity DataKi: 0.0350nMAssay Description:The enzyme inhibitory activity (Ki) was determined according to an assay protocol reported by Toth and Marshall (Toth, M. V.; Marshall, G. R. Int. J....More data for this Ligand-Target Pair
Affinity DataKi: 0.0730nMAssay Description:The enzyme inhibitory activity (Ki) was determined according to an assay protocol reported by Toth and Marshall (Toth, M. V.; Marshall, G. R. Int. J....More data for this Ligand-Target Pair
Affinity DataKi: 0.300nMAssay Description:The enzyme inhibitory activity (Ki) was determined according to an assay protocol reported by Toth and Marshall (Toth, M. V.; Marshall, G. R. Int. J....More data for this Ligand-Target Pair
Affinity DataEC50: 7nMAssay Description:For the Jurkat HIV Latency assay, compounds are dissolved and titrated in DMSO and diluted 100-fold in assay medium (RPMI-1640 containing 10% fetal b...More data for this Ligand-Target Pair
Affinity DataEC50: 22nMAssay Description:For the Jurkat HIV Latency assay, compounds are dissolved and titrated in DMSO and diluted 100-fold in assay medium (RPMI-1640 containing 10% fetal b...More data for this Ligand-Target Pair
Affinity DataIC50: 50nMpH: 7.4 T: 2°CAssay Description:The AlphaScreen assay was performed according to the manufacturer's protocol (Perkin Elmer, Benelux). Reactions were performed in 25 ul final volume ...More data for this Ligand-Target Pair
Affinity DataIC50: 55.4nMpH: 7.4 T: 2°CAssay Description:The AlphaScreen assay was performed according to the manufacturer's protocol (Perkin Elmer, Benelux). Reactions were performed in 25 ul final volume ...More data for this Ligand-Target Pair
Affinity DataIC50: 200nMAssay Description:Inhibition of HIV integrase.More data for this Ligand-Target Pair
Affinity DataIC50: 250nMAssay Description:Inhibition of HIV integrase.More data for this Ligand-Target Pair
Affinity DataEC50: 278nMAssay Description:For the Jurkat HIV Latency assay, compounds are dissolved and titrated in DMSO and diluted 100-fold in assay medium (RPMI-1640 containing 10% fetal b...More data for this Ligand-Target Pair
Affinity DataIC50: 360nMAssay Description:Inhibition of HIV integrase.More data for this Ligand-Target Pair
Affinity DataIC50: 370nMpH: 7.4 T: 2°CAssay Description:The AlphaScreen assay was performed according to the manufacturer's protocol (Perkin Elmer, Benelux). Reactions were performed in 25 ul final volume ...More data for this Ligand-Target Pair
Affinity DataIC50: 430nMpH: 7.4 T: 2°CAssay Description:The AlphaScreen assay was performed according to the manufacturer's protocol (Perkin Elmer, Benelux). Reactions were performed in 25 ul final volume ...More data for this Ligand-Target Pair
Affinity DataEC50: 616nMAssay Description:For the Jurkat HIV Latency assay, compounds are dissolved and titrated in DMSO and diluted 100-fold in assay medium (RPMI-1640 containing 10% fetal b...More data for this Ligand-Target Pair
Affinity DataIC50: 620nMpH: 7.4 T: 2°CAssay Description:The AlphaScreen assay was performed according to the manufacturer's protocol (Perkin Elmer, Benelux). Reactions were performed in 25 ul final volume ...More data for this Ligand-Target Pair
Affinity DataIC50: 750nMpH: 7.4 T: 2°CAssay Description:The AlphaScreen assay was performed according to the manufacturer's protocol (Perkin Elmer, Benelux). Reactions were performed in 25 ul final volume ...More data for this Ligand-Target Pair
Affinity DataIC50: 780nMpH: 7.4 T: 2°CAssay Description:The AlphaScreen assay was performed according to the manufacturer's protocol (Perkin Elmer, Benelux). Reactions were performed in 25 ul final volume ...More data for this Ligand-Target Pair
Affinity DataIC50: 950nMAssay Description:Inhibition of HIV integrase.More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMpH: 7.4 T: 2°CAssay Description:The AlphaScreen assay was performed according to the manufacturer's protocol (Perkin Elmer, Benelux). Reactions were performed in 25 ul final volume ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMpH: 7.4 T: 2°CAssay Description:The AlphaScreen assay was performed according to the manufacturer's protocol (Perkin Elmer, Benelux). Reactions were performed in 25 ul final volume ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMpH: 7.4 T: 2°CAssay Description:The AlphaScreen assay was performed according to the manufacturer's protocol (Perkin Elmer, Benelux). Reactions were performed in 25 ul final volume ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of HIV integrase.More data for this Ligand-Target Pair
Affinity DataEC50: 1.05E+3nMAssay Description:For the Jurkat HIV Latency assay, compounds are dissolved and titrated in DMSO and diluted 100-fold in assay medium (RPMI-1640 containing 10% fetal b...More data for this Ligand-Target Pair
Affinity DataEC50: 1.11E+3nMAssay Description:For the Jurkat HIV Latency assay, compounds are dissolved and titrated in DMSO and diluted 100-fold in assay medium (RPMI-1640 containing 10% fetal b...More data for this Ligand-Target Pair
Affinity DataIC50: 1.11E+3nMpH: 7.4 T: 2°CAssay Description:The AlphaScreen assay was performed according to the manufacturer's protocol (Perkin Elmer, Benelux). Reactions were performed in 25 ul final volume ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+3nMAssay Description:Inhibition of HIV integrase.More data for this Ligand-Target Pair
Affinity DataIC50: 1.21E+3nMpH: 7.4 T: 2°CAssay Description:The AlphaScreen assay was performed according to the manufacturer's protocol (Perkin Elmer, Benelux). Reactions were performed in 25 ul final volume ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.37E+3nMpH: 7.4 T: 2°CAssay Description:The AlphaScreen assay was performed according to the manufacturer's protocol (Perkin Elmer, Benelux). Reactions were performed in 25 ul final volume ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.57E+3nMpH: 7.4 T: 2°CAssay Description:The AlphaScreen assay was performed according to the manufacturer's protocol (Perkin Elmer, Benelux). Reactions were performed in 25 ul final volume ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+3nMAssay Description:Inhibition of HIV integrase.More data for this Ligand-Target Pair
Affinity DataIC50: 2.19E+3nMpH: 7.4 T: 2°CAssay Description:The AlphaScreen assay was performed according to the manufacturer's protocol (Perkin Elmer, Benelux). Reactions were performed in 25 ul final volume ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+3nMpH: 5.5Assay Description:Binding constants for inhibition of HIV protease were determined as previously described.More data for this Ligand-Target Pair
Affinity DataIC50: 2.66E+3nMpH: 7.4 T: 2°CAssay Description:The AlphaScreen assay was performed according to the manufacturer's protocol (Perkin Elmer, Benelux). Reactions were performed in 25 ul final volume ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.72E+3nMpH: 7.4 T: 2°CAssay Description:The AlphaScreen assay was performed according to the manufacturer's protocol (Perkin Elmer, Benelux). Reactions were performed in 25 ul final volume ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.85E+3nMpH: 7.4 T: 2°CAssay Description:The AlphaScreen assay was performed according to the manufacturer's protocol (Perkin Elmer, Benelux). Reactions were performed in 25 ul final volume ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+3nMpH: 7.4 T: 2°CAssay Description:The AlphaScreen assay was performed according to the manufacturer's protocol (Perkin Elmer, Benelux). Reactions were performed in 25 ul final volume ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.20E+3nMpH: 7.4 T: 2°CAssay Description:The AlphaScreen assay was performed according to the manufacturer's protocol (Perkin Elmer, Benelux). Reactions were performed in 25 ul final volume ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.30E+3nMpH: 5.5Assay Description:Binding constants for inhibition of HIV protease were determined as previously described.More data for this Ligand-Target Pair
Affinity DataIC50: 3.44E+3nMpH: 7.4 T: 2°CAssay Description:The AlphaScreen assay was performed according to the manufacturer's protocol (Perkin Elmer, Benelux). Reactions were performed in 25 ul final volume ...More data for this Ligand-Target Pair
Affinity DataEC50: 3.50E+3nMAssay Description:For the Jurkat HIV Latency assay, compounds are dissolved and titrated in DMSO and diluted 100-fold in assay medium (RPMI-1640 containing 10% fetal b...More data for this Ligand-Target Pair
Affinity DataEC50: 3.68E+3nMAssay Description:For the Jurkat HIV Latency assay, compounds are dissolved and titrated in DMSO and diluted 100-fold in assay medium (RPMI-1640 containing 10% fetal b...More data for this Ligand-Target Pair
Affinity DataIC50: 4.31E+3nMpH: 7.4 T: 2°CAssay Description:The AlphaScreen assay was performed according to the manufacturer's protocol (Perkin Elmer, Benelux). Reactions were performed in 25 ul final volume ...More data for this Ligand-Target Pair
Affinity DataEC50: 4.33E+3nMAssay Description:For the Jurkat HIV Latency assay, compounds are dissolved and titrated in DMSO and diluted 100-fold in assay medium (RPMI-1640 containing 10% fetal b...More data for this Ligand-Target Pair
Affinity DataIC50: 4.34E+3nMpH: 7.4 T: 2°CAssay Description:The AlphaScreen assay was performed according to the manufacturer's protocol (Perkin Elmer, Benelux). Reactions were performed in 25 ul final volume ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.48E+3nMpH: 7.4 T: 2°CAssay Description:The AlphaScreen assay was performed according to the manufacturer's protocol (Perkin Elmer, Benelux). Reactions were performed in 25 ul final volume ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.50E+3nMpH: 7.4 T: 2°CAssay Description:The AlphaScreen assay was performed according to the manufacturer's protocol (Perkin Elmer, Benelux). Reactions were performed in 25 ul final volume ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.54E+3nMpH: 7.4 T: 2°CAssay Description:The AlphaScreen assay was performed according to the manufacturer's protocol (Perkin Elmer, Benelux). Reactions were performed in 25 ul final volume ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.76E+3nMpH: 7.4 T: 2°CAssay Description:The AlphaScreen assay was performed according to the manufacturer's protocol (Perkin Elmer, Benelux). Reactions were performed in 25 ul final volume ...More data for this Ligand-Target Pair