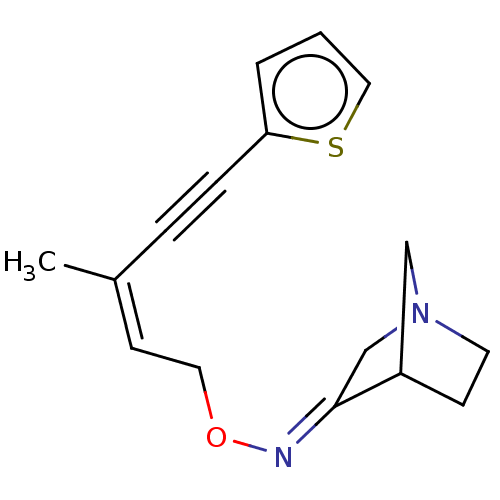
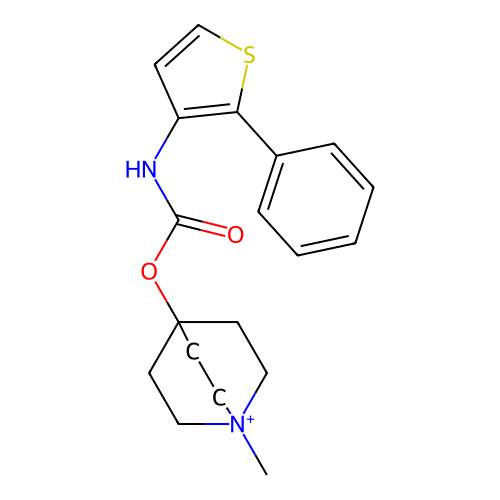
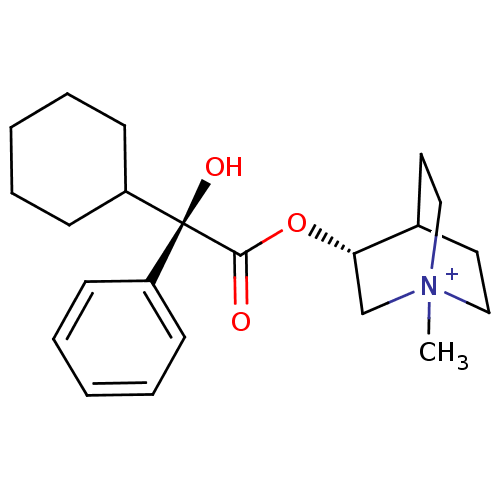
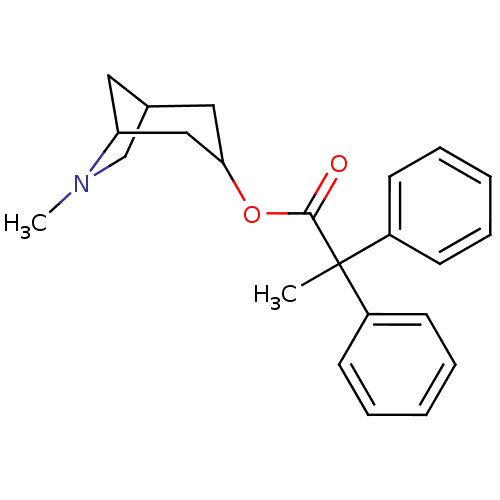
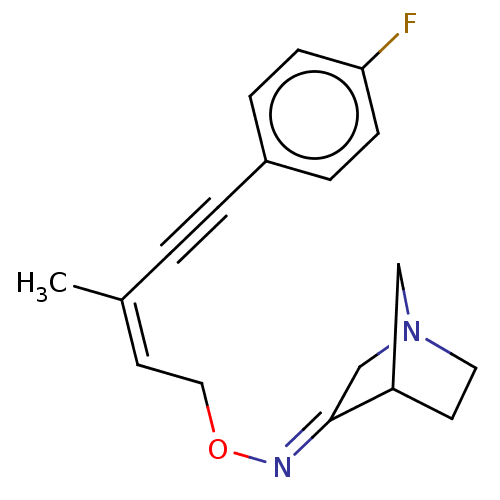
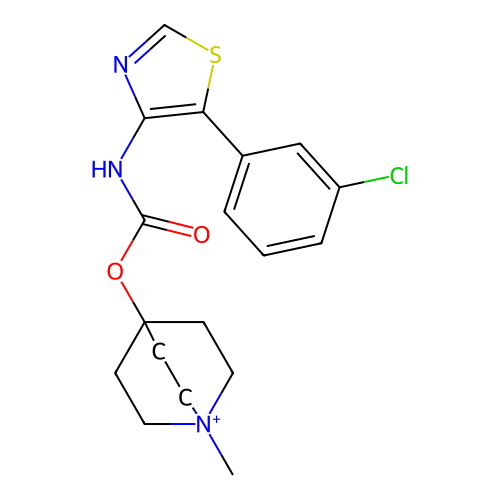
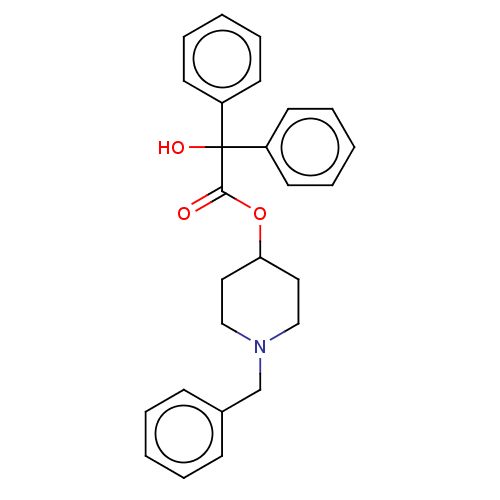
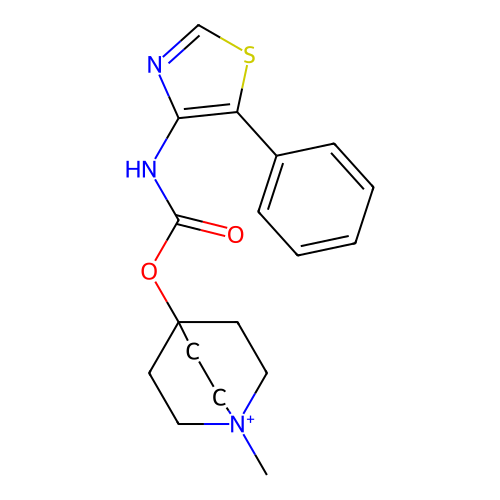
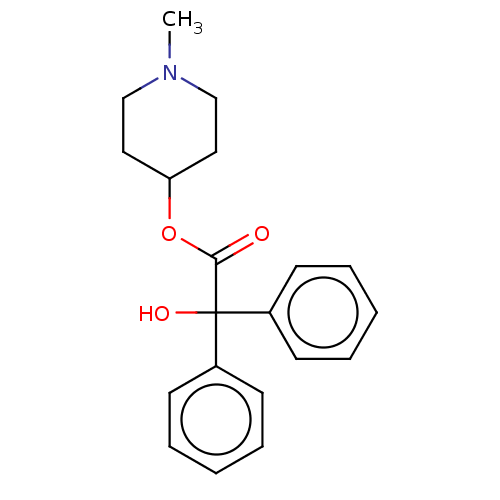
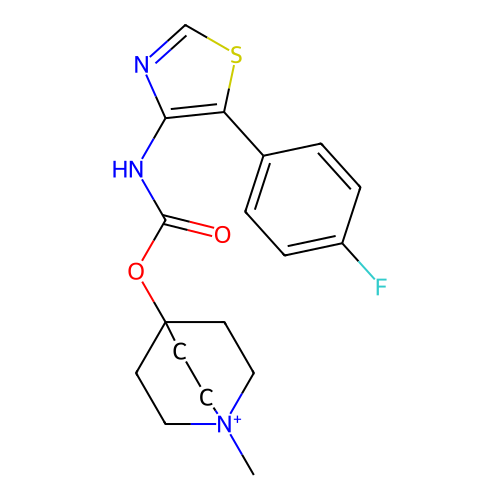
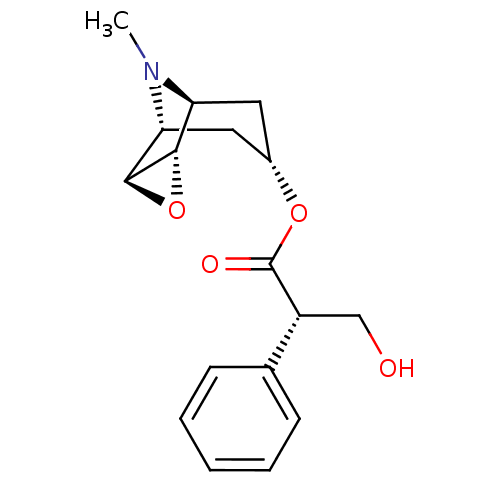
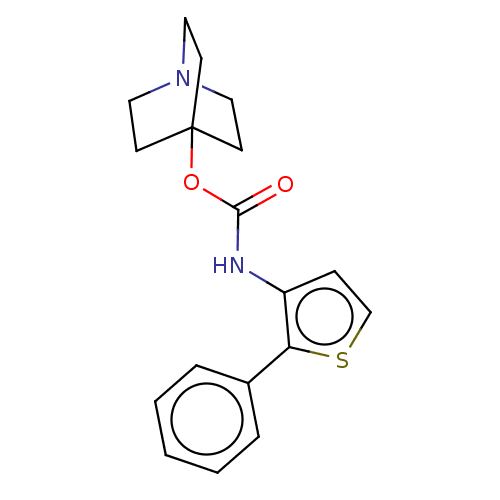
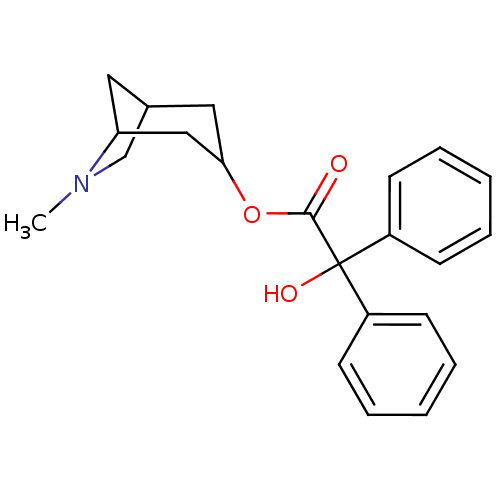
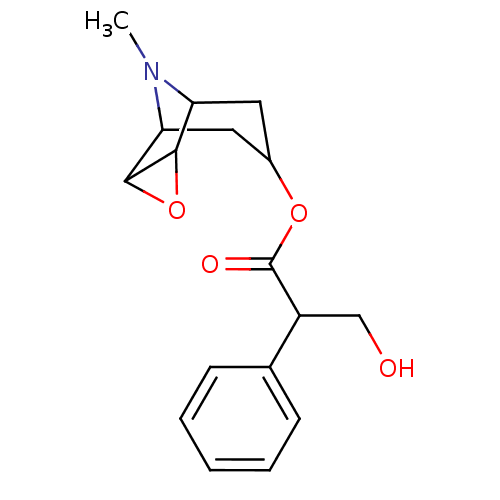
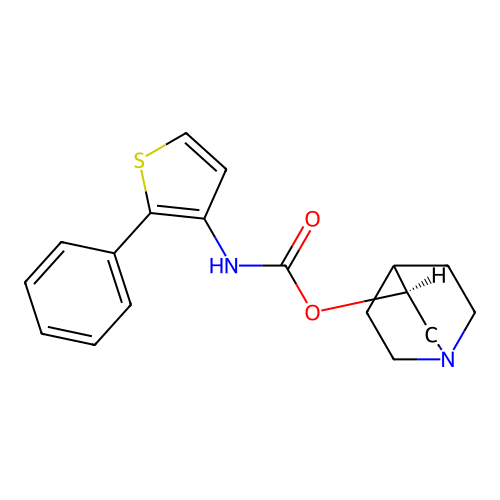
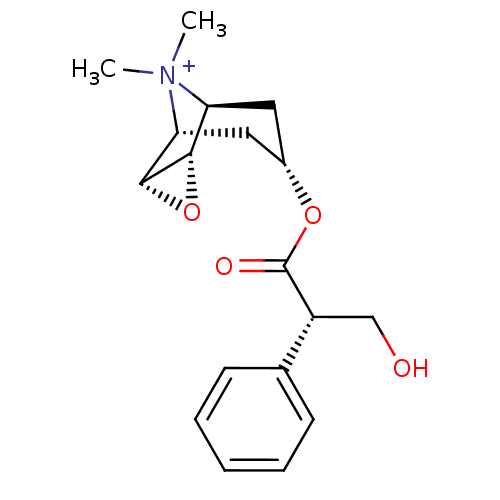
Affinity DataKi: 0.0100nMAssay Description:Binding affinity towards rat Muscarinic acetylcholine receptor M1 was determinedMore data for this Ligand-Target Pair
Affinity DataIC50: 0.0200nMAssay Description:Binding affinity against muscarinic receptor using radioligand [3H]cis-methyldioxolane (CMD) binding assay in membrane preparations of rat neocortex.More data for this Ligand-Target Pair
Affinity DataKi: 0.0200nMAssay Description:Displacement of [3H]-NMS from Sprague-Dawley rat muscarinic M1 receptor in cortex membraneMore data for this Ligand-Target Pair
Affinity DataKi: 0.0200nMAssay Description:Binding affinity towards rat Muscarinic acetylcholine receptor M1 was determinedMore data for this Ligand-Target Pair
Affinity DataKi: 0.0300nMAssay Description:Inhibition of [3H]QNB binding against muscarinic acetylcholine receptor in rat brain.More data for this Ligand-Target Pair
Affinity DataIC50: 0.0300nMAssay Description:Binding affinity against muscarinic receptor using radioligand [3H]cis-methyldioxolane (CMD) binding assay in membrane preparations of rat neocortex.More data for this Ligand-Target Pair
Affinity DataKi: 0.0383nMAssay Description:The compound was tested for binding activity against muscarinic acetylcholine receptor M1, using [3H]QNB as the radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 0.0410nMAssay Description:Displacement of [3H]-NMS from Sprague-Dawley rat muscarinic M1 receptor in cortex membraneMore data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:In vitro for binding affinity for muscarinic acetylcholine receptors in homogenized rat brain in the presence of [3H]scopolamine.More data for this Ligand-Target Pair
Affinity DataKi: 0.0550nMAssay Description:Displacement of [3H]-NMS from Sprague-Dawley rat muscarinic M1 receptor in cortex membraneMore data for this Ligand-Target Pair
Affinity DataKi: 0.0700nMAssay Description:In vitro for binding affinity for muscarinic acetylcholine receptors in homogenized rat brain in the presence of [3H]scopolamine.More data for this Ligand-Target Pair
Affinity DataKi: 0.0730nMAssay Description:Displacement of [3H]-NMS from Sprague-Dawley rat muscarinic M1 receptor in cortex membraneMore data for this Ligand-Target Pair
Affinity DataKi: 0.0800nMAssay Description:Inhibition of [3H]pirenzepine binding to muscarinic receptor in rat cortical homogenatesMore data for this Ligand-Target Pair
Affinity DataKi: 0.0950nMAssay Description:Displacement of [3H]-NMS from Sprague-Dawley rat muscarinic M1 receptor in cortex membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 0.100nMAssay Description:Binding affinity against muscarinic receptor using radioligand [3H]cis-methyldioxolane (CMD) binding assay in membrane preparations of rat neocortex.More data for this Ligand-Target Pair
Affinity DataKi: 0.118nMAssay Description:Inhibition of [3H]pirenzepine binding against muscarinic acetylcholine receptor in rat brain.More data for this Ligand-Target Pair
Affinity DataKi: 0.120nMAssay Description:Displacement of [3H]-NMS from Sprague-Dawley rat muscarinic M1 receptor in cortex membraneMore data for this Ligand-Target Pair
Affinity DataKi: 0.130nMAssay Description:Binding affinity towards rat Muscarinic acetylcholine receptor M1 was determinedMore data for this Ligand-Target Pair
Affinity DataKi: 0.150nMAssay Description:Ability to displace [3H]pirenzepine (PZ) from M1 receptor in rat cortex homogenateMore data for this Ligand-Target Pair
Affinity DataIC50: 0.150nMAssay Description:Binding affinity against muscarinic receptor using radioligand [3H]cis-methyldioxolane (CMD) binding assay in membrane preparations of rat neocortex.More data for this Ligand-Target Pair
Affinity DataKi: 0.150nMAssay Description:Displacement of [3H]pirenzepine from muscarinic M1 receptor of rat cortex homogenates.More data for this Ligand-Target Pair
Affinity DataKi: 0.150nMAssay Description:Displacement of [3H]pirenzepine from muscarinic acetylcholine receptor M1 in rat cortex homogenatesMore data for this Ligand-Target Pair
Affinity DataIC50: 0.150nMAssay Description:Inhibitory constant by the inhibition of [3H]N-methylscopolamine (NMS) binding to m1 receptor of transfected A9L cellsMore data for this Ligand-Target Pair
Affinity DataKi: 0.160nMAssay Description:In vitro for binding affinity for muscarinic acetylcholine receptors in homogenized rat brain in the presence of [3H]scopolamine.More data for this Ligand-Target Pair
Affinity DataKi: 0.160nMAssay Description:Displacement of [3H]-NMS from Sprague-Dawley rat muscarinic M1 receptor in cortex membraneMore data for this Ligand-Target Pair
Affinity DataKi: 0.166nMAssay Description:Inhibition of [3H]pirenzepine binding against muscarinic acetylcholine receptor in rat brain.More data for this Ligand-Target Pair
Affinity DataKi: 0.190nMAssay Description:Displacement of [3H]-NMS from Sprague-Dawley rat muscarinic M1 receptor in cortex membraneMore data for this Ligand-Target Pair
Affinity DataKi: 0.196nMAssay Description:Inhibition of [3H]QNB binding against muscarinic acetylcholine receptor in rat heart.More data for this Ligand-Target Pair
Affinity DataKi: 0.197nMAssay Description:Inhibition of [3H]pirenzepine binding against muscarinic acetylcholine receptor in rat brain.More data for this Ligand-Target Pair
Affinity DataKi: 0.200nMAssay Description:Binding affinity against Muscarinic acetylcholine receptor using [3H]-OXO-M as the radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 0.220nMAssay Description:The compound was tested for binding activity against muscarinic acetylcholine receptor M1, using [3H]QNB as the radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 0.224nMAssay Description:Inhibition of [3H]QNB binding against muscarinic acetylcholine receptor in rat brain.More data for this Ligand-Target Pair
Affinity DataKd: 0.230nMAssay Description:In vitro binding affinity towards Muscarinic acetylcholine receptor M1 using [3H]QNB as radioligand from rat heart tissueMore data for this Ligand-Target Pair
Affinity DataKi: 0.240nMAssay Description:Binding affinity against Muscarinic acetylcholine receptors using [3H]oxotremorine-M as radioligand in rat cortexMore data for this Ligand-Target Pair
Affinity DataKi: 0.254nMAssay Description:Inhibition of [3H]QNB binding against muscarinic acetylcholine receptor in rat brain.More data for this Ligand-Target Pair
Affinity DataKi: 0.260nMAssay Description:Displacement of [3H]pirenzepine from muscarinic acetylcholine receptor M1 in rat cortex homogenatesMore data for this Ligand-Target Pair
Affinity DataKi: 0.260nMAssay Description:Ability to displace [3H]pirenzepine (PZ) from M1 receptor in rat cortex homogenateMore data for this Ligand-Target Pair
Affinity DataKi: 0.290nMAssay Description:Displacement of [3H]-NMS from Sprague-Dawley rat muscarinic M1 receptor in cortex membraneMore data for this Ligand-Target Pair
Affinity DataKi: 0.297nMAssay Description:Inhibition of [3H]QNB binding against muscarinic acetylcholine receptor in rat heart.More data for this Ligand-Target Pair
Affinity DataIC50: 0.300nMAssay Description:Compound was evaluated for inhibition of [3H]cis-methyldioxolane binding to label agonist sites (RCMD) in rat neocortexMore data for this Ligand-Target Pair
Affinity DataKi: 0.300nMAssay Description:The compound was tested for inhibition of [3H]NMS binding against muscarinic acetylcholine receptor in rat brainMore data for this Ligand-Target Pair
Affinity DataKi: 0.300nMAssay Description:Binding affinity to muscarinic acetylcholine receptor was determined in presence of [3H]OXO-M radioligand (in vitro)More data for this Ligand-Target Pair
Affinity DataKi: 0.302nMAssay Description:Inhibition of [3H]QNB binding against muscarinic acetylcholine receptor in rat heart.More data for this Ligand-Target Pair
Affinity DataKi: 0.310nMAssay Description:Displacement of [3H]-NMS from Sprague-Dawley rat muscarinic M1 receptor in cortex membraneMore data for this Ligand-Target Pair