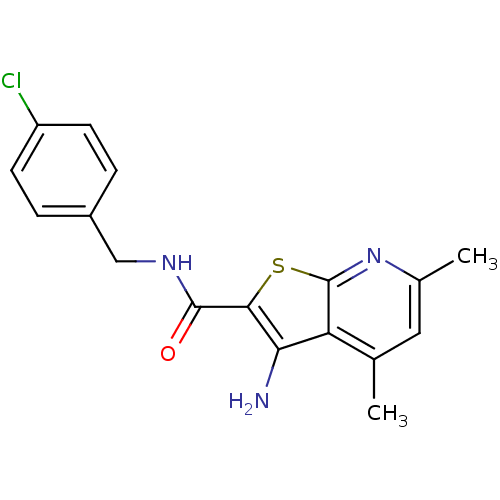
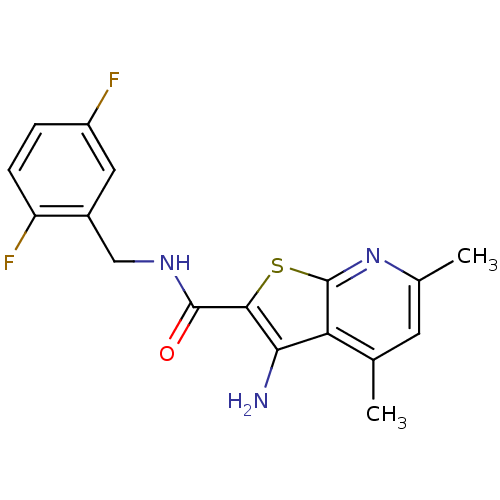
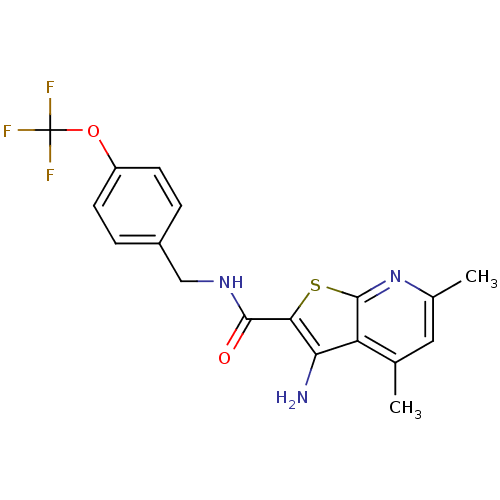
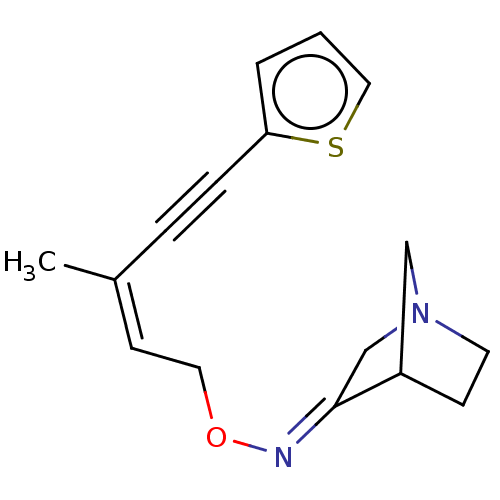
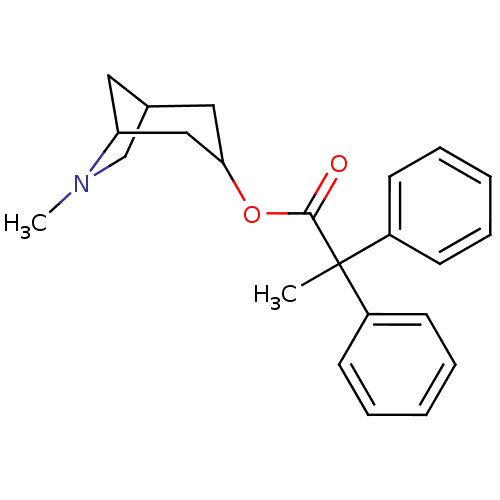
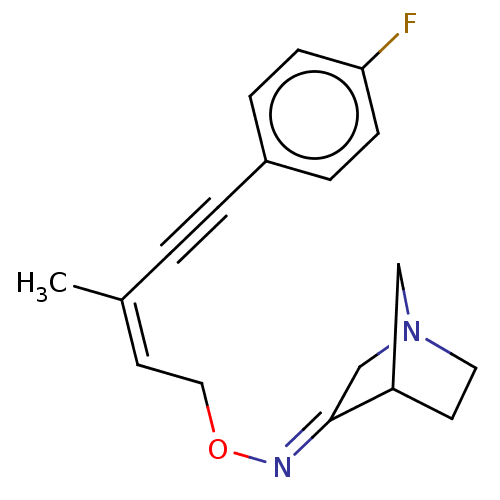
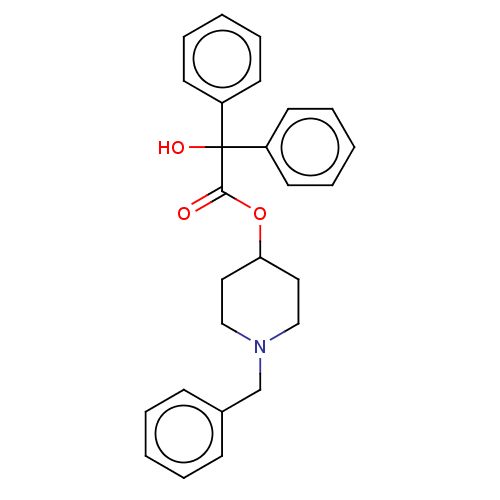
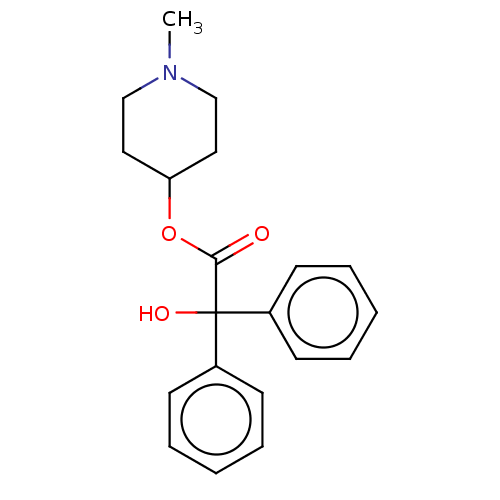
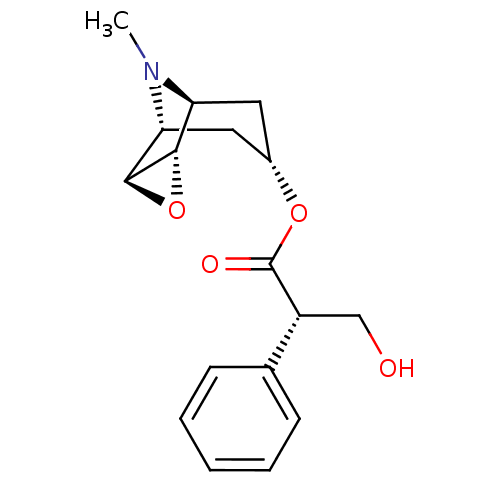
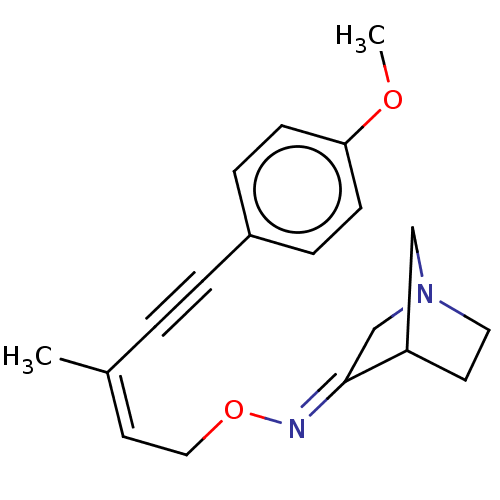
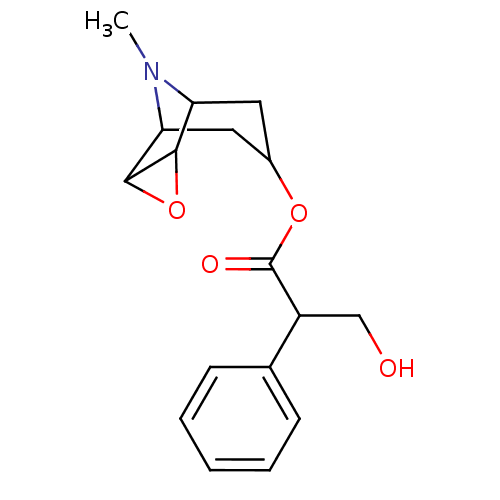
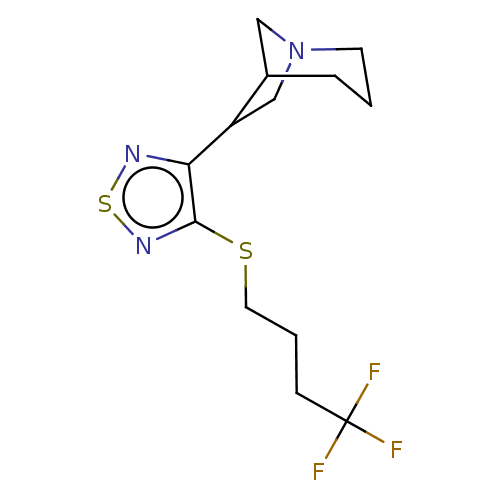
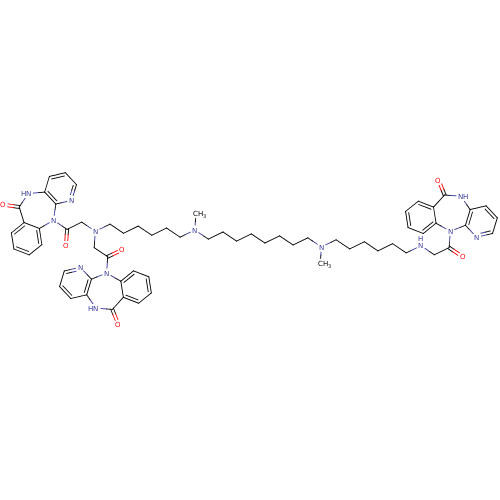
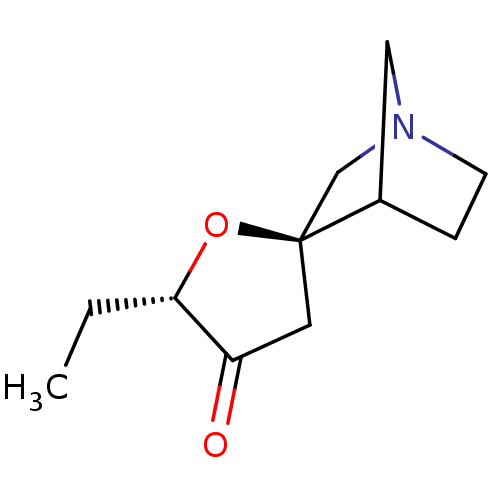
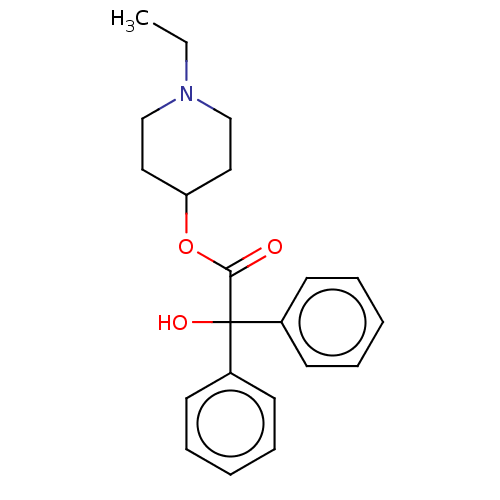
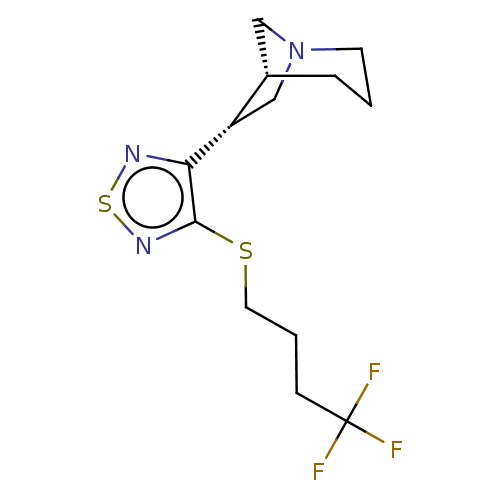
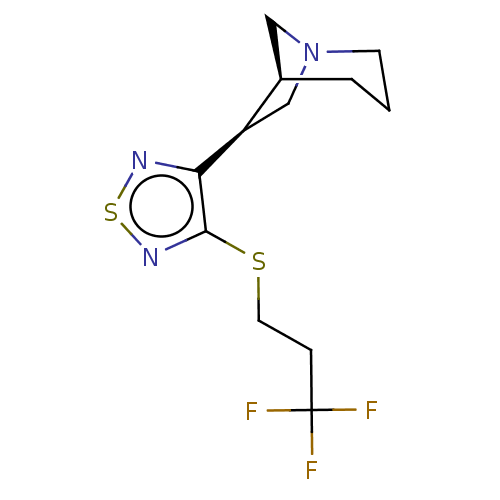
TargetMuscarinic acetylcholine receptor M4(Rat)
Vanderbilt Screening Center For Gpcrs, Ion Channels and Transporters
Curated by PubChem BioAssay
Vanderbilt Screening Center For Gpcrs, Ion Channels and Transporters
Curated by PubChem BioAssay
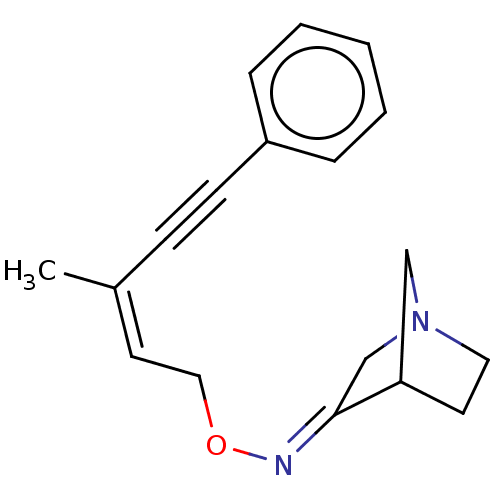
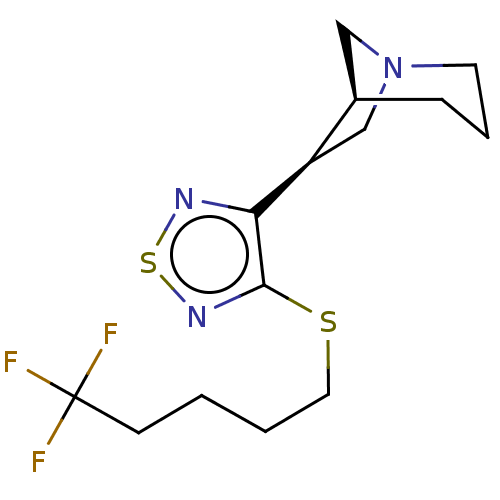
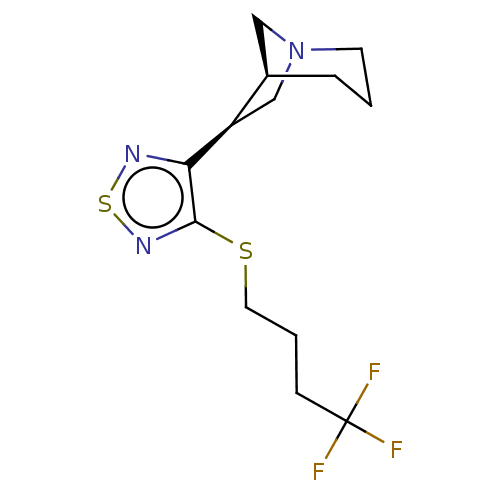
Affinity DataEC50: 0.000339nMAssay Description:Assay Provider: Colleen Niswender Assay Provider Affiliation: Vanderbilt University Grant Title: Discovery of a Highly Selective in vitro and in vivo...More data for this Ligand-Target Pair
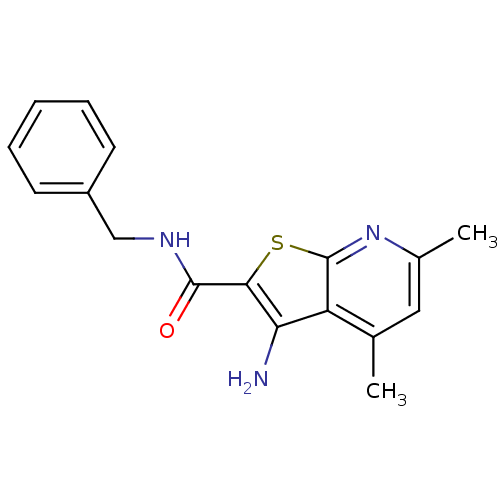
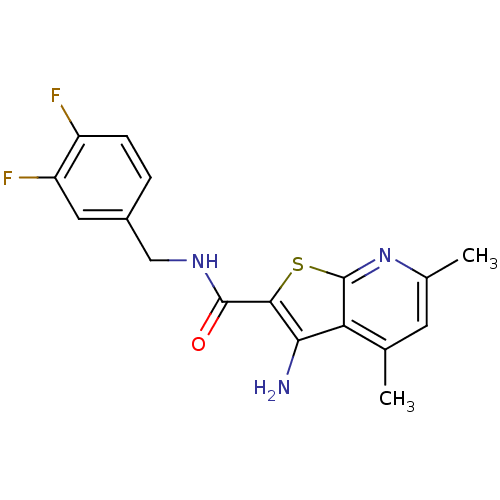
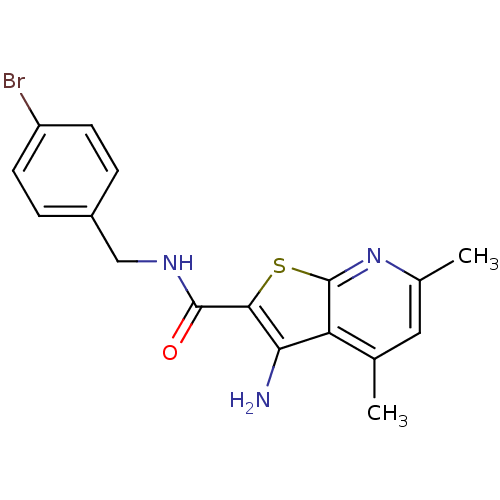
TargetMuscarinic acetylcholine receptor M4(Rat)
Vanderbilt Screening Center For Gpcrs, Ion Channels and Transporters
Curated by PubChem BioAssay
Vanderbilt Screening Center For Gpcrs, Ion Channels and Transporters
Curated by PubChem BioAssay
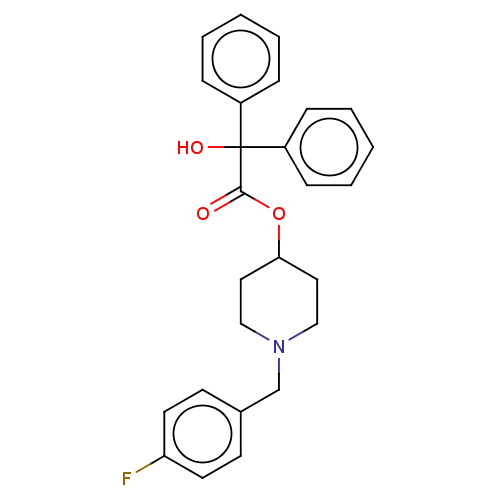
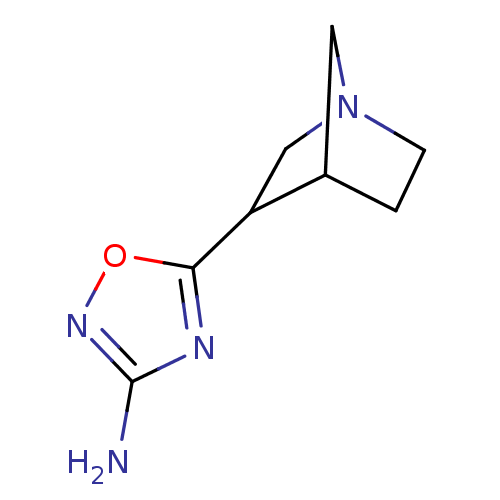
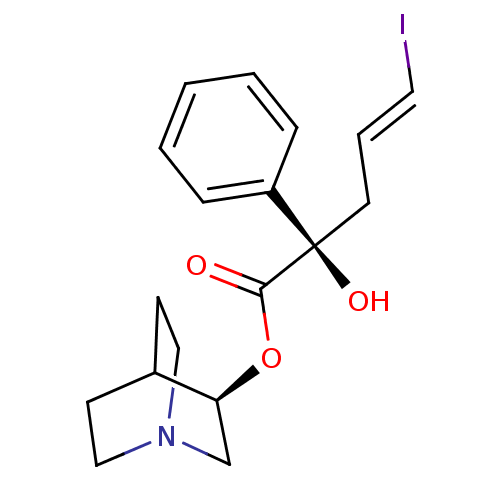
Affinity DataEC50: 0.000693nMAssay Description:Assay Provider: Colleen Niswender Assay Provider Affiliation: Vanderbilt University Grant Title: Discovery of a Highly Selective in vitro and in vivo...More data for this Ligand-Target Pair
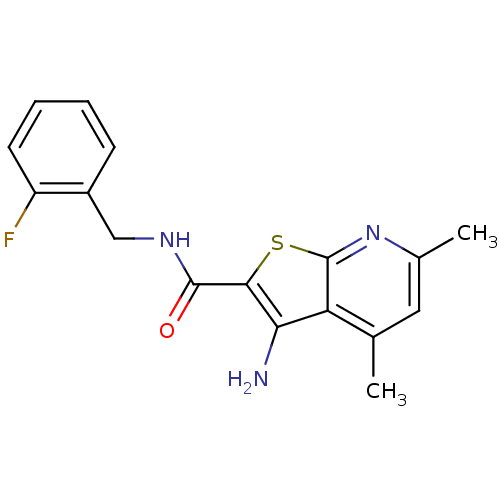
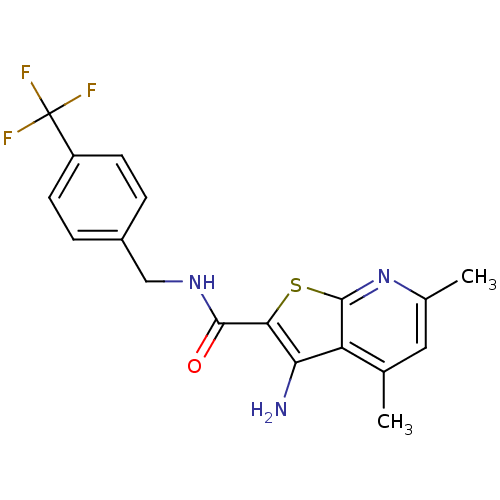
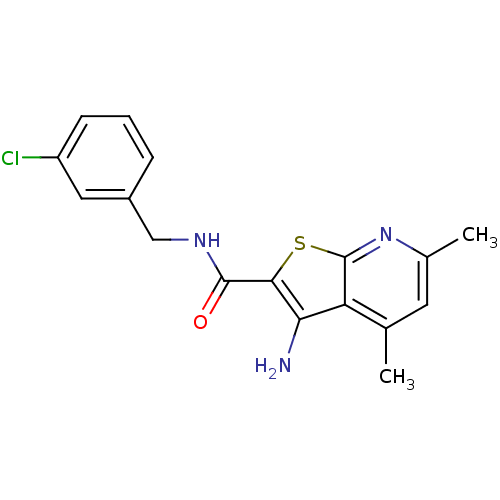
TargetMuscarinic acetylcholine receptor M4(Rat)
Vanderbilt Screening Center For Gpcrs, Ion Channels and Transporters
Curated by PubChem BioAssay
Vanderbilt Screening Center For Gpcrs, Ion Channels and Transporters
Curated by PubChem BioAssay
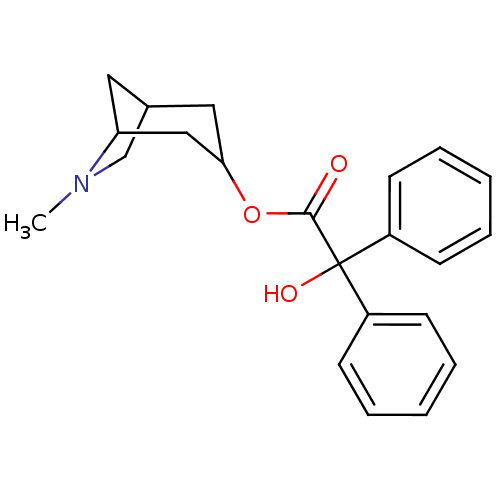
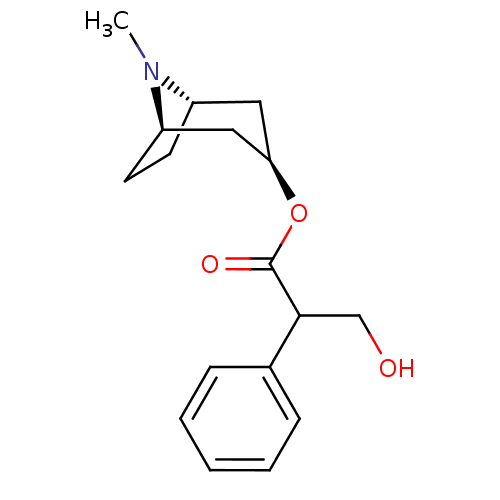
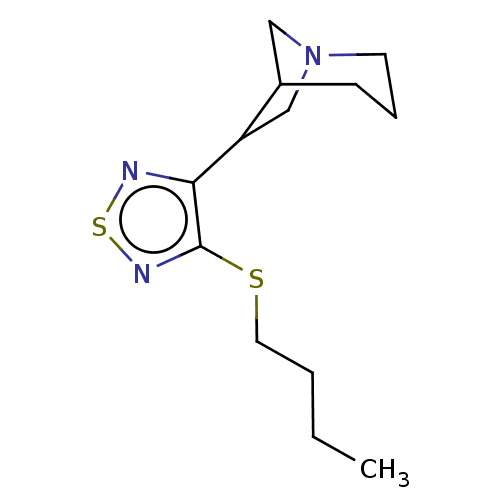
Affinity DataEC50: 0.000873nMAssay Description:Assay Provider: Colleen Niswender Assay Provider Affiliation: Vanderbilt University Grant Title: Discovery of a Highly Selective in vitro and in vivo...More data for this Ligand-Target Pair
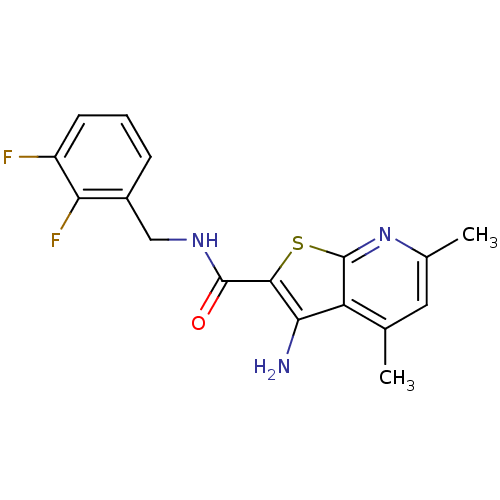
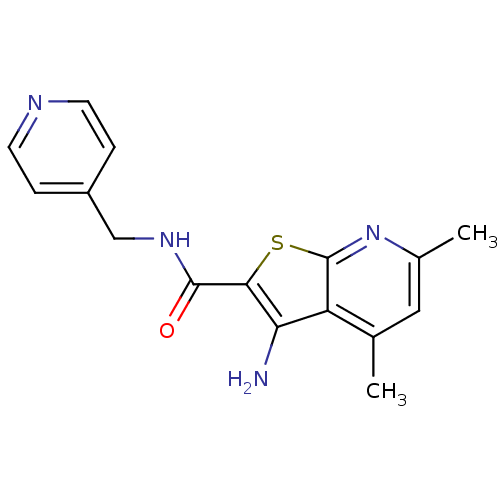
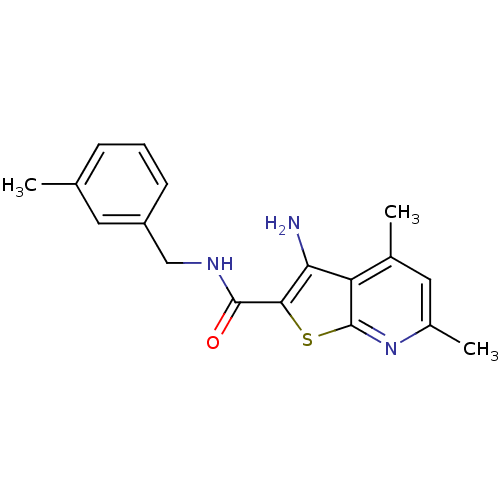
TargetMuscarinic acetylcholine receptor M4(Rat)
Vanderbilt Screening Center For Gpcrs, Ion Channels and Transporters
Curated by PubChem BioAssay
Vanderbilt Screening Center For Gpcrs, Ion Channels and Transporters
Curated by PubChem BioAssay
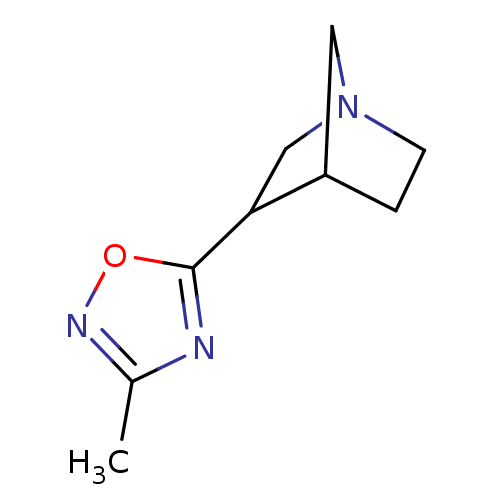
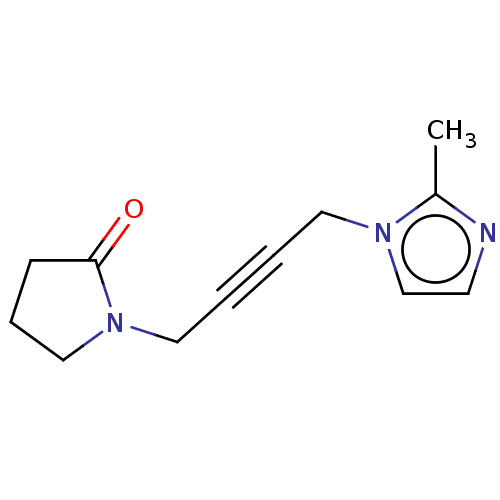
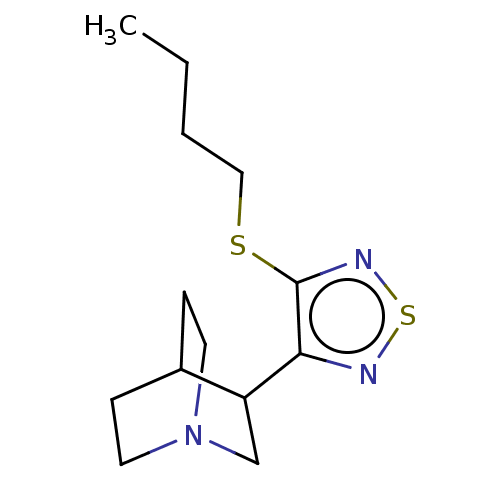
Affinity DataEC50: 0.000973nMAssay Description:Assay Provider: Colleen Niswender Assay Provider Affiliation: Vanderbilt University Grant Title: Discovery of a Highly Selective in vitro and in vivo...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M4(Rat)
Vanderbilt Screening Center For Gpcrs, Ion Channels and Transporters
Curated by PubChem BioAssay
Vanderbilt Screening Center For Gpcrs, Ion Channels and Transporters
Curated by PubChem BioAssay
Affinity DataEC50: 0.00100nMAssay Description:Assay Provider: Colleen Niswender Assay Provider Affiliation: Vanderbilt University Grant Title: Discovery of a Highly Selective in vitro and in vivo...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M4(Rat)
Vanderbilt Screening Center For Gpcrs, Ion Channels and Transporters
Curated by PubChem BioAssay
Vanderbilt Screening Center For Gpcrs, Ion Channels and Transporters
Curated by PubChem BioAssay
Affinity DataEC50: 0.00127nMAssay Description:Assay Provider: Colleen Niswender Assay Provider Affiliation: Vanderbilt University Grant Title: Discovery of a Highly Selective in vitro and in vivo...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M4(Rat)
Vanderbilt Screening Center For Gpcrs, Ion Channels and Transporters
Curated by PubChem BioAssay
Vanderbilt Screening Center For Gpcrs, Ion Channels and Transporters
Curated by PubChem BioAssay
Affinity DataEC50: 0.00234nMAssay Description:Assay Provider: Colleen Niswender Assay Provider Affiliation: Vanderbilt University Grant Title: Discovery of a Highly Selective in vitro and in vivo...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M4(Rat)
Vanderbilt Screening Center For Gpcrs, Ion Channels and Transporters
Curated by PubChem BioAssay
Vanderbilt Screening Center For Gpcrs, Ion Channels and Transporters
Curated by PubChem BioAssay
Affinity DataEC50: 0.00242nMAssay Description:Assay Provider: Colleen Niswender Assay Provider Affiliation: Vanderbilt University Grant Title: Discovery of a Highly Selective in vitro and in vivo...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M4(Rat)
Vanderbilt Screening Center For Gpcrs, Ion Channels and Transporters
Curated by PubChem BioAssay
Vanderbilt Screening Center For Gpcrs, Ion Channels and Transporters
Curated by PubChem BioAssay
Affinity DataEC50: 0.00242nMAssay Description:Assay Provider: Colleen Niswender Assay Provider Affiliation: Vanderbilt University Grant Title: Discovery of a Highly Selective in vitro and in vivo...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M4(Rat)
Vanderbilt Screening Center For Gpcrs, Ion Channels and Transporters
Curated by PubChem BioAssay
Vanderbilt Screening Center For Gpcrs, Ion Channels and Transporters
Curated by PubChem BioAssay
Affinity DataEC50: 0.00282nMAssay Description:Assay Provider: Colleen Niswender Assay Provider Affiliation: Vanderbilt University Grant Title: Discovery of a Highly Selective in vitro and in vivo...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M4(Rat)
Vanderbilt Screening Center For Gpcrs, Ion Channels and Transporters
Curated by PubChem BioAssay
Vanderbilt Screening Center For Gpcrs, Ion Channels and Transporters
Curated by PubChem BioAssay
Affinity DataEC50: 0.00506nMAssay Description:Assay Provider: Colleen Niswender Assay Provider Affiliation: Vanderbilt University Grant Title: Discovery of a Highly Selective in vitro and in vivo...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M4(Rat)
Vanderbilt Screening Center For Gpcrs, Ion Channels and Transporters
Curated by PubChem BioAssay
Vanderbilt Screening Center For Gpcrs, Ion Channels and Transporters
Curated by PubChem BioAssay
Affinity DataEC50: 0.00516nMAssay Description:Assay Provider: Colleen Niswender Assay Provider Affiliation: Vanderbilt University Grant Title: Discovery of a Highly Selective in vitro and in vivo...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M4(Rat)
Vanderbilt Screening Center For Gpcrs, Ion Channels and Transporters
Curated by PubChem BioAssay
Vanderbilt Screening Center For Gpcrs, Ion Channels and Transporters
Curated by PubChem BioAssay
Affinity DataEC50: 0.00559nMAssay Description:Assay Provider: Colleen Niswender Assay Provider Affiliation: Vanderbilt University Grant Title: Discovery of a Highly Selective in vitro and in vivo...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M4(Rat)
Vanderbilt Screening Center For Gpcrs, Ion Channels and Transporters
Curated by PubChem BioAssay
Vanderbilt Screening Center For Gpcrs, Ion Channels and Transporters
Curated by PubChem BioAssay
Affinity DataEC50: 0.00903nMAssay Description:Assay Provider: Colleen Niswender Assay Provider Affiliation: Vanderbilt University Grant Title: Discovery of a Highly Selective in vitro and in vivo...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M4(Rat)
Vanderbilt Screening Center For Gpcrs, Ion Channels and Transporters
Curated by PubChem BioAssay
Vanderbilt Screening Center For Gpcrs, Ion Channels and Transporters
Curated by PubChem BioAssay
Affinity DataEC50: 0.0102nMAssay Description:Assay Provider: Colleen Niswender Assay Provider Affiliation: Vanderbilt University Grant Title: Discovery of a Highly Selective in vitro and in vivo...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0200nMAssay Description:Binding affinity against muscarinic receptor using radioligand [3H]cis-methyldioxolane (CMD) binding assay in membrane preparations of rat neocortex.More data for this Ligand-Target Pair
Affinity DataKi: 0.0300nMAssay Description:Inhibition of [3H]QNB binding against muscarinic acetylcholine receptor in rat brain.More data for this Ligand-Target Pair
Affinity DataIC50: 0.0300nMAssay Description:Binding affinity against muscarinic receptor using radioligand [3H]cis-methyldioxolane (CMD) binding assay in membrane preparations of rat neocortex.More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:In vitro for binding affinity for muscarinic acetylcholine receptors in homogenized rat brain in the presence of [3H]scopolamine.More data for this Ligand-Target Pair
Affinity DataKi: 0.0700nMAssay Description:In vitro for binding affinity for muscarinic acetylcholine receptors in homogenized rat brain in the presence of [3H]scopolamine.More data for this Ligand-Target Pair
Affinity DataKi: 0.0800nMAssay Description:Inhibition of [3H]pirenzepine binding to muscarinic receptor in rat cortical homogenatesMore data for this Ligand-Target Pair
Affinity DataIC50: 0.100nMAssay Description:Binding affinity against muscarinic receptor using radioligand [3H]cis-methyldioxolane (CMD) binding assay in membrane preparations of rat neocortex.More data for this Ligand-Target Pair
Affinity DataKi: 0.118nMAssay Description:Inhibition of [3H]pirenzepine binding against muscarinic acetylcholine receptor in rat brain.More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M4(Rat)
Vanderbilt Screening Center For Gpcrs, Ion Channels and Transporters
Curated by PubChem BioAssay
Vanderbilt Screening Center For Gpcrs, Ion Channels and Transporters
Curated by PubChem BioAssay
Affinity DataIC50: 0.150nMAssay Description:Binding affinity against muscarinic receptor using radioligand [3H]cis-methyldioxolane (CMD) binding assay in membrane preparations of rat neocortex.More data for this Ligand-Target Pair
Affinity DataKi: 0.160nMAssay Description:In vitro for binding affinity for muscarinic acetylcholine receptors in homogenized rat brain in the presence of [3H]scopolamine.More data for this Ligand-Target Pair
Affinity DataKi: 0.166nMAssay Description:Inhibition of [3H]pirenzepine binding against muscarinic acetylcholine receptor in rat brain.More data for this Ligand-Target Pair
Affinity DataKi: 0.196nMAssay Description:Inhibition of [3H]QNB binding against muscarinic acetylcholine receptor in rat heart.More data for this Ligand-Target Pair
Affinity DataKi: 0.197nMAssay Description:Inhibition of [3H]pirenzepine binding against muscarinic acetylcholine receptor in rat brain.More data for this Ligand-Target Pair
Affinity DataKi: 0.200nMAssay Description:Binding affinity against Muscarinic acetylcholine receptor using [3H]-OXO-M as the radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 0.224nMAssay Description:Inhibition of [3H]QNB binding against muscarinic acetylcholine receptor in rat brain.More data for this Ligand-Target Pair
Affinity DataKi: 0.240nMAssay Description:Binding affinity against Muscarinic acetylcholine receptors using [3H]oxotremorine-M as radioligand in rat cortexMore data for this Ligand-Target Pair
Affinity DataKi: 0.254nMAssay Description:Inhibition of [3H]QNB binding against muscarinic acetylcholine receptor in rat brain.More data for this Ligand-Target Pair
Affinity DataKi: 0.297nMAssay Description:Inhibition of [3H]QNB binding against muscarinic acetylcholine receptor in rat heart.More data for this Ligand-Target Pair
Affinity DataIC50: 0.300nMAssay Description:Compound was evaluated for inhibition of [3H]cis-methyldioxolane binding to label agonist sites (RCMD) in rat neocortexMore data for this Ligand-Target Pair
Affinity DataKi: 0.300nMAssay Description:The compound was tested for inhibition of [3H]NMS binding against muscarinic acetylcholine receptor in rat brainMore data for this Ligand-Target Pair
Affinity DataKi: 0.300nMAssay Description:Binding affinity to muscarinic acetylcholine receptor was determined in presence of [3H]OXO-M radioligand (in vitro)More data for this Ligand-Target Pair
Affinity DataKi: 0.302nMAssay Description:Inhibition of [3H]QNB binding against muscarinic acetylcholine receptor in rat heart.More data for this Ligand-Target Pair
Affinity DataKi: 0.330nMAssay Description:Binding affinity against Muscarinic acetylcholine receptors using [3H]oxotremorine-M as radioligand in rat cortexMore data for this Ligand-Target Pair
Affinity DataKi: 0.337nMAssay Description:Inhibition of [3H]QNB binding against muscarinic acetylcholine receptor in rat brain.More data for this Ligand-Target Pair
Affinity DataKi: 0.337nMAssay Description:Inhibition of [3H]QNB binding to muscarinic acetylcholine receptor of rat heart membrane preparation.More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M4(Rat)
Vanderbilt Screening Center For Gpcrs, Ion Channels and Transporters
Curated by PubChem BioAssay
Vanderbilt Screening Center For Gpcrs, Ion Channels and Transporters
Curated by PubChem BioAssay
Affinity DataKd: 0.356nMAssay Description:In vitro binding affinity towards Muscarinic acetylcholine receptor M4 using [3H]QNB as radioligand from rat heart tissueMore data for this Ligand-Target Pair
Affinity DataKi: 0.360nMAssay Description:Binding affinity against Muscarinic acetylcholine receptors using [3H]oxotremorine-M as radioligand in rat cortexMore data for this Ligand-Target Pair
Affinity DataKi: 0.400nMAssay Description:Binding affinity against Muscarinic acetylcholine receptors using [3H]oxotremorine-M as radioligand in rat cortexMore data for this Ligand-Target Pair
Affinity DataKi: 0.400nMAssay Description:Binding affinity against Muscarinic acetylcholine receptor using [3H]oxotremorine-M as radioligand in rat cortexMore data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M4(Rat)
Vanderbilt Screening Center For Gpcrs, Ion Channels and Transporters
Curated by PubChem BioAssay
Vanderbilt Screening Center For Gpcrs, Ion Channels and Transporters
Curated by PubChem BioAssay
Affinity DataKi: 0.400nMAssay Description:Binding affinity against Muscarinic acetylcholine receptor using [3H]-OXO-M as the radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 0.410nMAssay Description:In vitro for binding affinity for muscarinic acetylcholine receptors in homogenized rat brain in the presence of [3H]scopolamine.More data for this Ligand-Target Pair
Affinity DataKi: 0.420nMAssay Description:Binding affinity against Muscarinic acetylcholine receptors using [3H]oxotremorine-M as radioligand in rat cortexMore data for this Ligand-Target Pair
Affinity DataKi: 0.440nMAssay Description:Binding affinity against Muscarinic acetylcholine receptors using [3H]oxotremorine-M as radioligand in rat cortexMore data for this Ligand-Target Pair