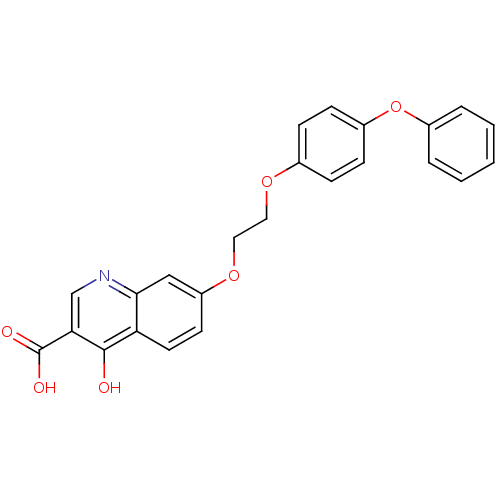
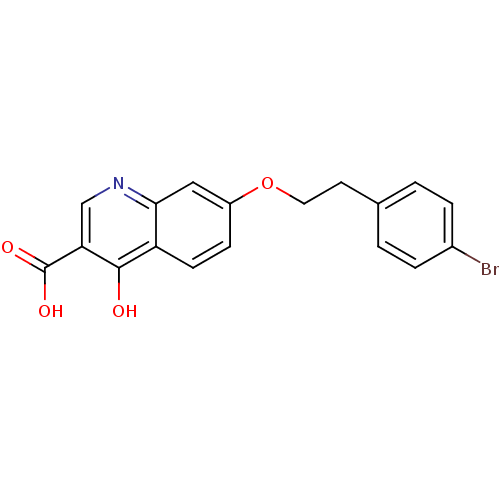
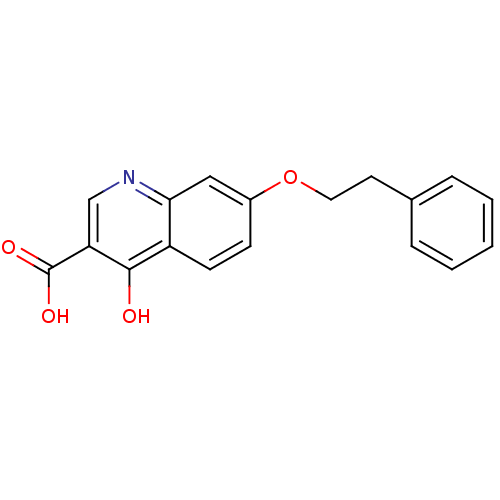
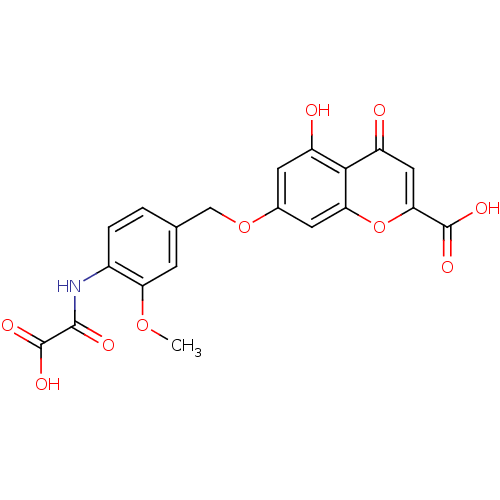
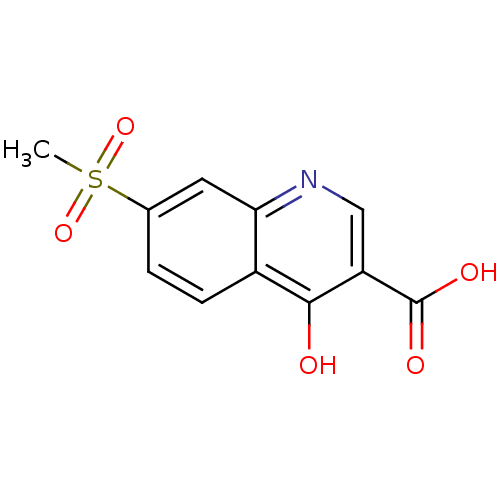
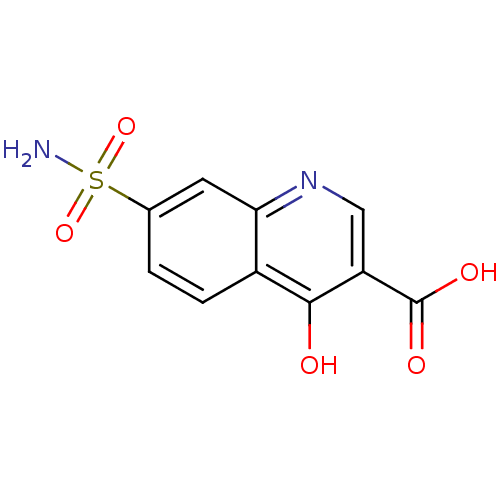
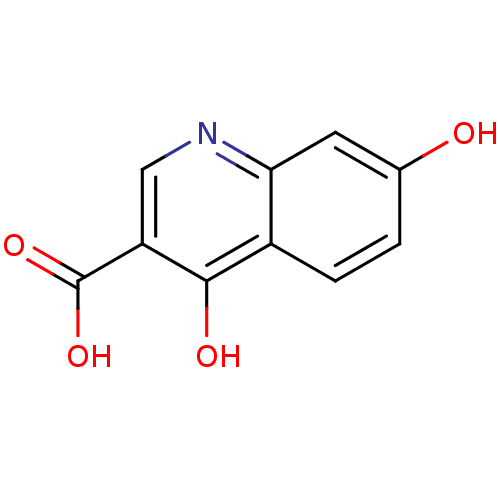
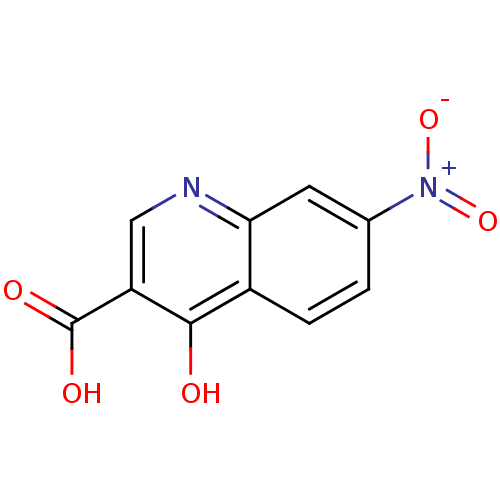
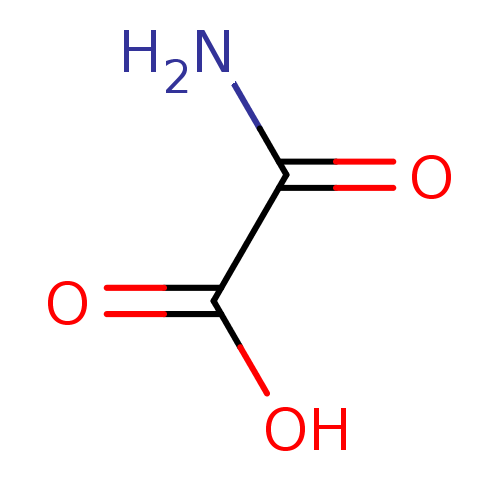
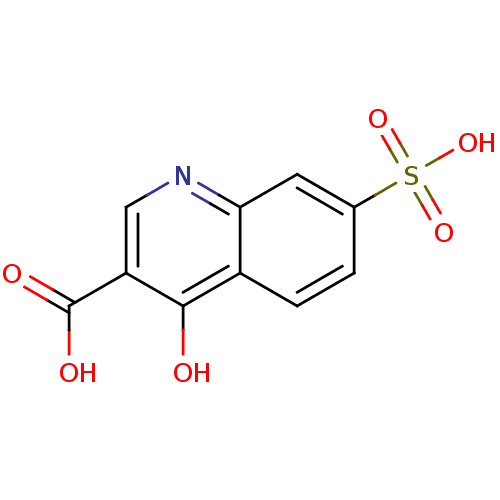
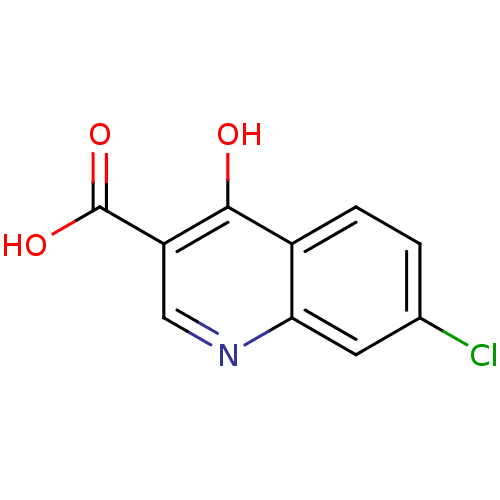
Affinity DataIC50: 4.88E+3nMAssay Description:LDH was assayed spectrophotometrically for reduction of pyruvate using NADH by recording the changed in absorbance at 340 nm. The reaction was initia...More data for this Ligand-Target Pair
Affinity DataIC50: 5.80E+3nMAssay Description:LDH was assayed spectrophotometrically for reduction of pyruvate using NADH by recording the changed in absorbance at 340 nm. The reaction was initia...More data for this Ligand-Target Pair
Affinity DataIC50: 6.31E+3nMAssay Description:LDH was assayed spectrophotometrically for reduction of pyruvate using NADH by recording the changed in absorbance at 340 nm. The reaction was initia...More data for this Ligand-Target Pair
Affinity DataIC50: 1.09E+4nMAssay Description:LDH was assayed spectrophotometrically for reduction of pyruvate using NADH by recording the changed in absorbance at 340 nm. The reaction was initia...More data for this Ligand-Target Pair
Affinity DataIC50: 1.36E+4nMAssay Description:LDH was assayed spectrophotometrically for reduction of pyruvate using NADH by recording the changed in absorbance at 340 nm. The reaction was initia...More data for this Ligand-Target Pair
Affinity DataIC50: 3.09E+4nMAssay Description:Ability to inhibit cytoplasmic malate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataKi: 3.63E+4nMAssay Description:In vitro inhibitory activity against pig heart cytoplasmic malate dehydrogenase (MDH)More data for this Ligand-Target Pair
Affinity DataIC50: 6.17E+4nMAssay Description:Ability to inhibit cytoplasmic malate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 7.24E+4nMAssay Description:Ability to inhibit cytoplasmic malate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 8.13E+4nMAssay Description:Ability to inhibit cytoplasmic malate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 9.12E+4nMAssay Description:Ability to inhibit cytoplasmic malate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 1.55E+5nMAssay Description:Ability to inhibit cytoplasmic malate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 2.32E+5nMAssay Description:LDH was assayed spectrophotometrically for reduction of pyruvate using NADH by recording the changed in absorbance at 340 nm. The reaction was initia...More data for this Ligand-Target Pair
Affinity DataIC50: 3.72E+5nMAssay Description:Ability to inhibit cytoplasmic malate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 4.37E+5nMAssay Description:Ability to inhibit cytoplasmic malate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 6.03E+5nMAssay Description:Ability to inhibit cytoplasmic malate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 6.92E+5nMAssay Description:Ability to inhibit cytoplasmic malate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 9.12E+5nMAssay Description:Ability to inhibit cytoplasmic malate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 9.55E+5nMAssay Description:Ability to inhibit cytoplasmic malate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+6nMAssay Description:Ability to inhibit cytoplasmic malate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 1.29E+6nMAssay Description:Ability to inhibit cytoplasmic malate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+6nMAssay Description:LDH was assayed spectrophotometrically for reduction of pyruvate using NADH by recording the changed in absorbance at 340 nm. The reaction was initia...More data for this Ligand-Target Pair
Affinity DataIC50: 3.09E+6nMAssay Description:Ability to inhibit cytoplasmic malate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 4.68E+6nMAssay Description:Ability to inhibit cytoplasmic malate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+7nMAssay Description:Ability to inhibit cytoplasmic malate dehydrogenaseMore data for this Ligand-Target Pair