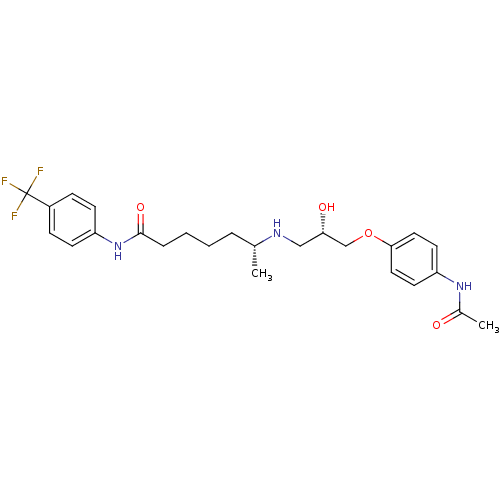
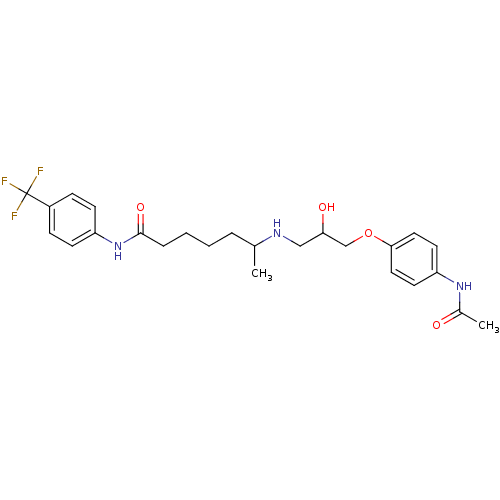
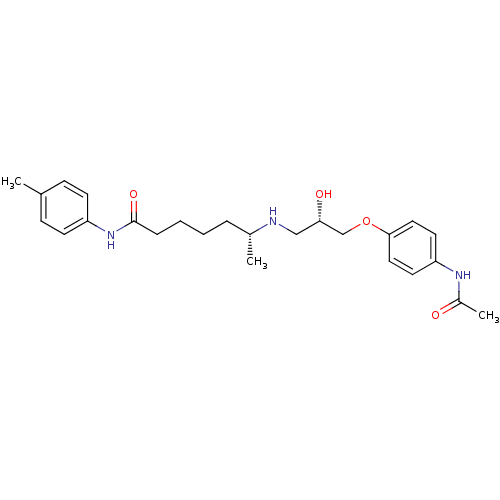
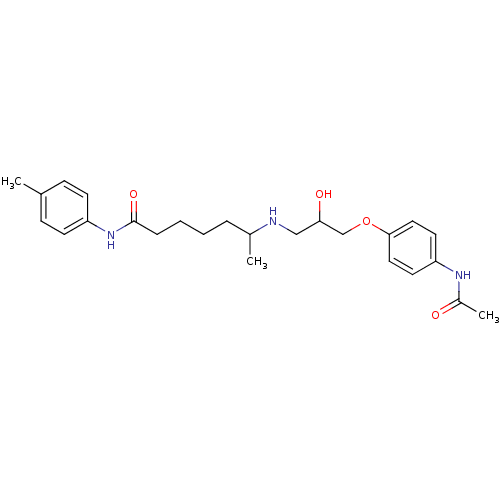
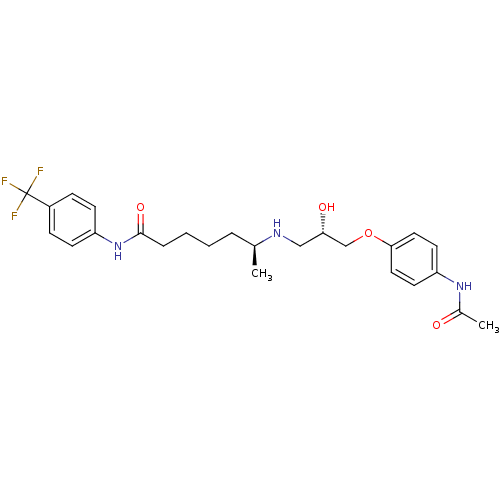
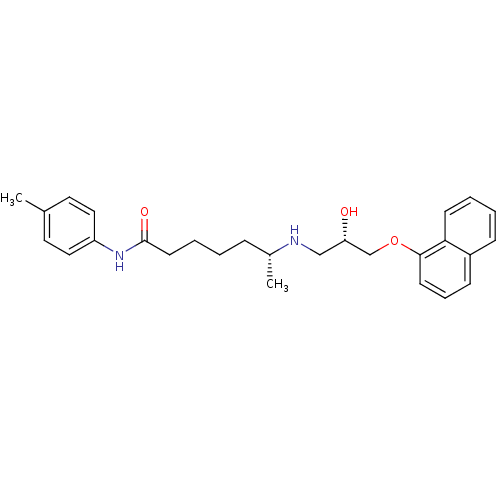
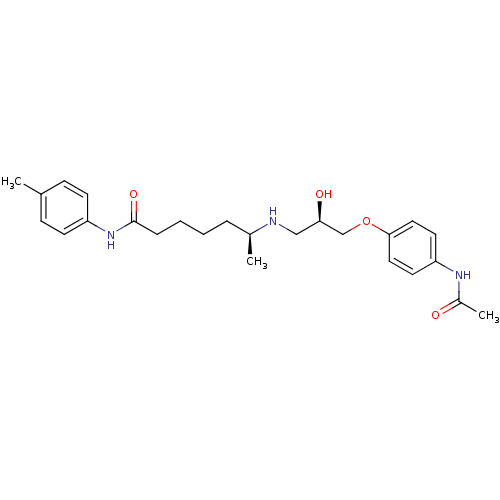
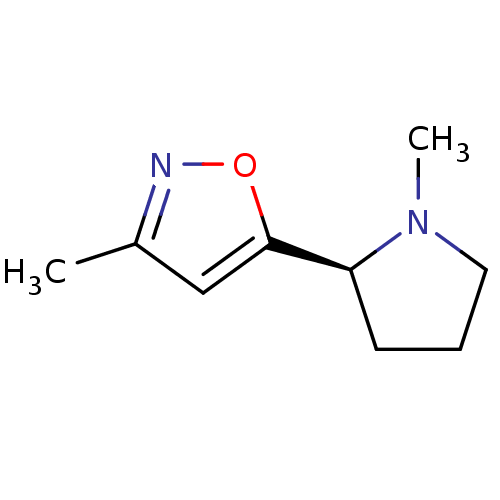
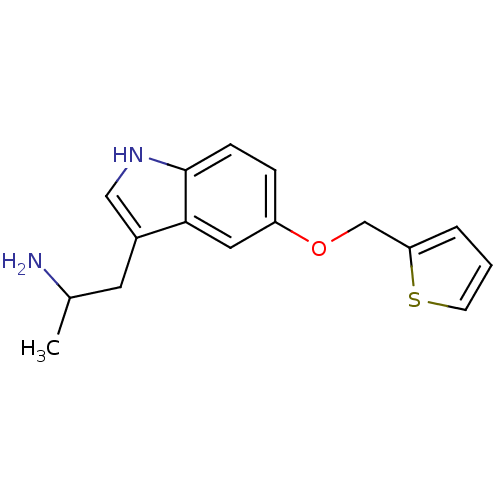
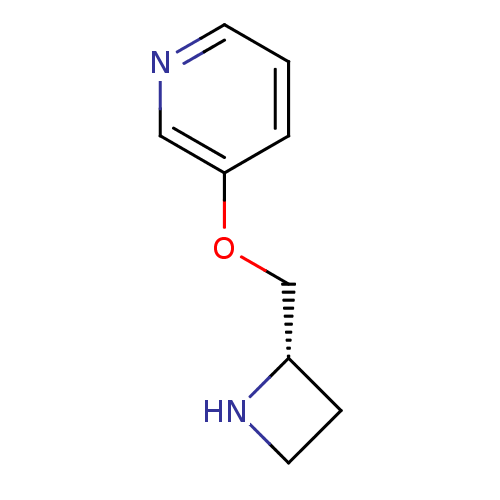
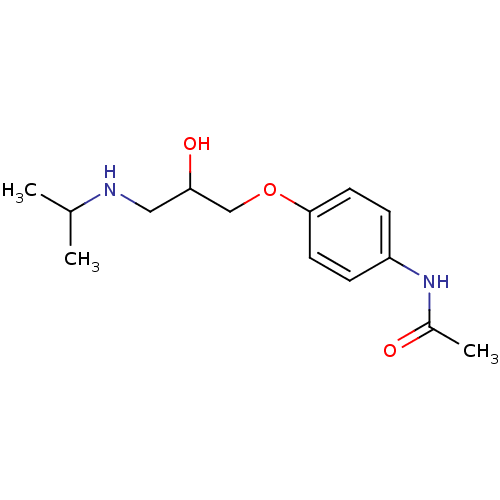
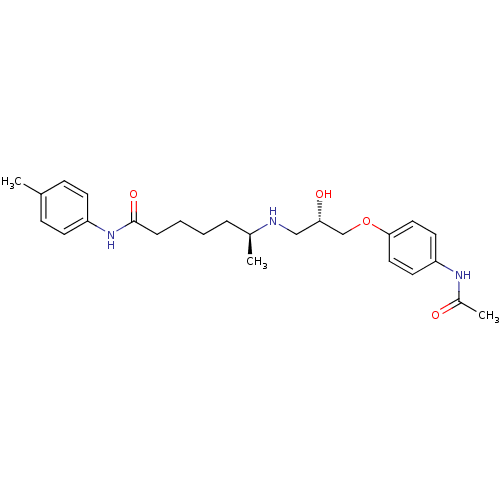
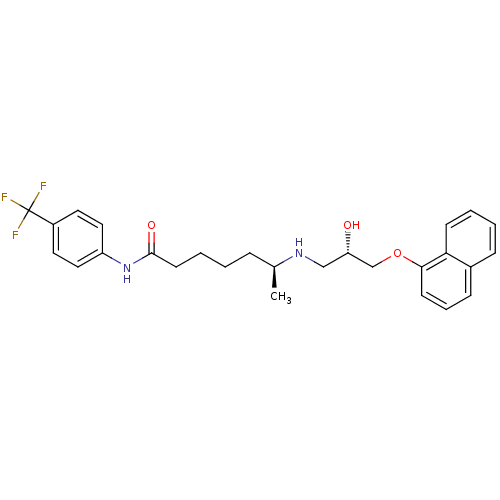
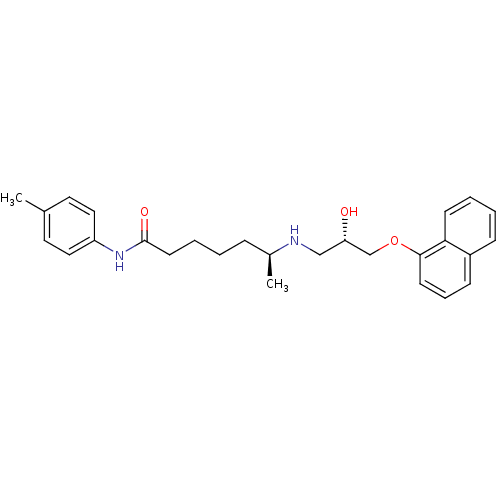
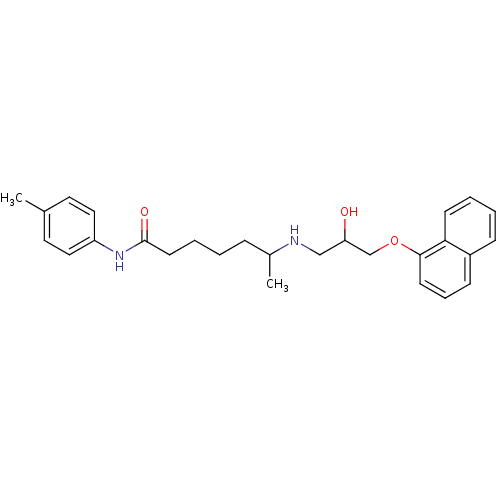
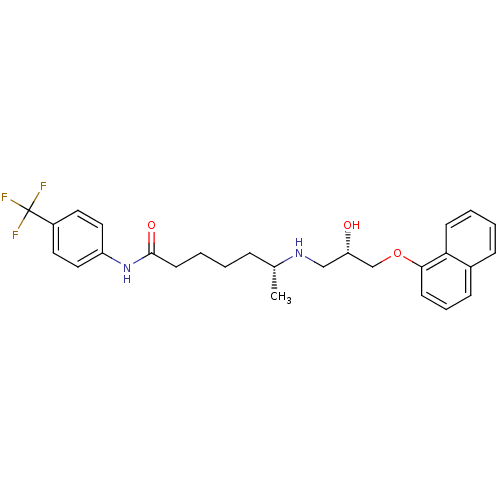
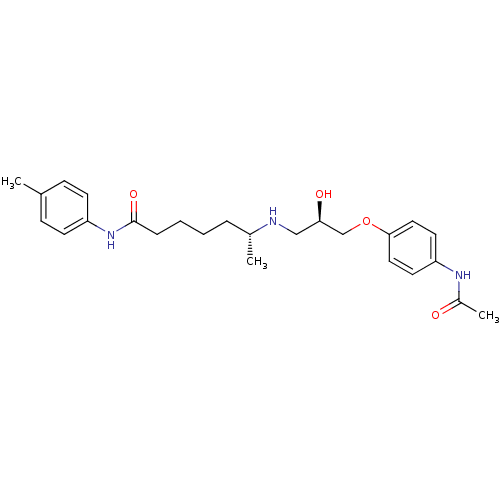
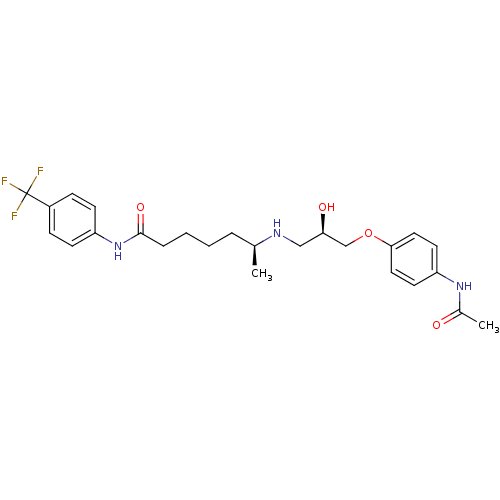
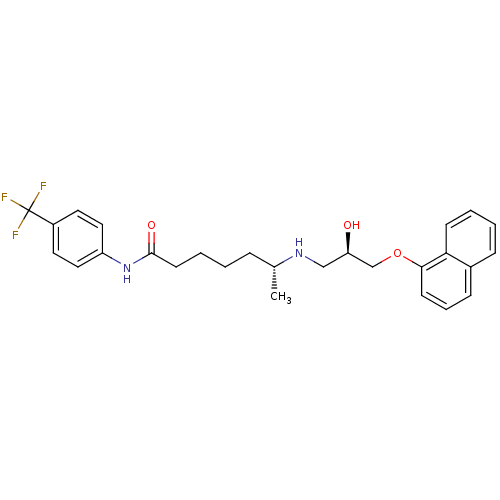
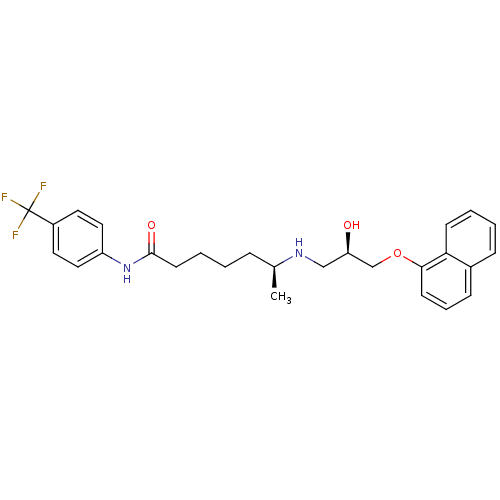
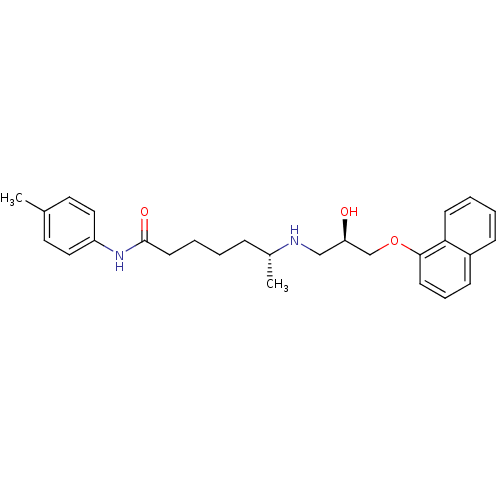
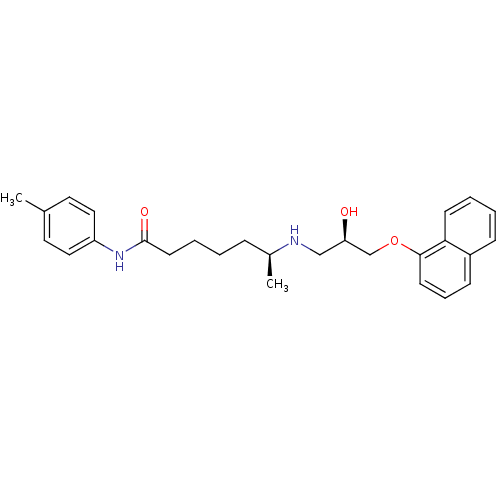
Affinity DataKi: 146nMAssay Description:Ki value was determined by accumulation of c-AMP in S-49 mouse lymphoma cells (Beta2).More data for this Ligand-Target Pair
Affinity DataKi: 390nMAssay Description:Ki value was determined by accumulation of c-AMP in S-49 mouse lymphoma cells (Beta2).More data for this Ligand-Target Pair
Affinity DataKi: 810nMAssay Description:Ki value was determined by accumulation of c-AMP in S-49 mouse lymphoma cells (Beta2).More data for this Ligand-Target Pair
Affinity DataKi: 1.18E+3nMAssay Description:Ki value was determined by accumulation of c-AMP in S-49 mouse lymphoma cells (Beta2).More data for this Ligand-Target Pair
Affinity DataKi: 2.17E+3nMAssay Description:Ki value was determined by accumulation of c-AMP in S-49 mouse lymphoma cells (Beta2).More data for this Ligand-Target Pair
Affinity DataKi: 2.85E+3nMAssay Description:Ki value was determined by accumulation of c-AMP in S-49 mouse lymphoma cells (Beta2).More data for this Ligand-Target Pair
Affinity DataKi: 9.07E+3nMAssay Description:Ki value was determined by accumulation of c-AMP in S-49 mouse lymphoma cells (Beta2).More data for this Ligand-Target Pair
Affinity DataKi: 9.24E+3nMAssay Description:Ki value was determined by accumulation of c-AMP in S-49 mouse lymphoma cells (Beta2).More data for this Ligand-Target Pair
Affinity DataKi: 1.35E+4nMAssay Description:Ki value was determined by accumulation of c-AMP in S-49 mouse lymphoma cells (Beta2).More data for this Ligand-Target Pair
Affinity DataKi: 1.56E+4nMAssay Description:Ki value was determined by accumulation of c-AMP in S-49 mouse lymphoma cells (Beta2).More data for this Ligand-Target Pair
Affinity DataKi: 3.90E+4nMAssay Description:Ki value was determined by accumulation of c-AMP in S-49 mouse lymphoma cells (Beta2).More data for this Ligand-Target Pair
Affinity DataKi: 4.10E+4nMAssay Description:Ki value was determined by accumulation of c-AMP in S-49 mouse lymphoma cells (Beta2).More data for this Ligand-Target Pair
Affinity DataKi: 4.33E+4nMAssay Description:Ki value was determined by accumulation of c-AMP in S-49 mouse lymphoma cells (Beta2).More data for this Ligand-Target Pair
Affinity DataKi: 4.59E+4nMAssay Description:Ki value was determined by accumulation of c-AMP in S-49 mouse lymphoma cells (Beta2).More data for this Ligand-Target Pair
Affinity DataKi: 5.36E+4nMAssay Description:Ki value was determined by accumulation of c-AMP in S-49 mouse lymphoma cells (Beta2).More data for this Ligand-Target Pair
Affinity DataKi: 6.20E+4nMAssay Description:Ki value was determined by accumulation of c-AMP in S-49 mouse lymphoma cells (Beta2).More data for this Ligand-Target Pair
Affinity DataKi: 7.80E+4nMAssay Description:Ki value was determined by accumulation of c-AMP in S-49 mouse lymphoma cells (Beta2).More data for this Ligand-Target Pair
Affinity DataKi: 1.30E+5nMAssay Description:Ki value was determined by accumulation of c-AMP in S-49 mouse lymphoma cells (Beta2).More data for this Ligand-Target Pair
Affinity DataKi: 1.79E+5nMAssay Description:Ki value was determined by accumulation of c-AMP in S-49 mouse lymphoma cells (Beta2).More data for this Ligand-Target Pair
Affinity DataKi: 1.95E+5nMAssay Description:Ki value was determined by accumulation of c-AMP in S-49 mouse lymphoma cells (Beta2).More data for this Ligand-Target Pair
Affinity DataKi: 1.95E+5nMAssay Description:Ki value was determined by accumulation of c-AMP in S-49 mouse lymphoma cells (Beta2).More data for this Ligand-Target Pair