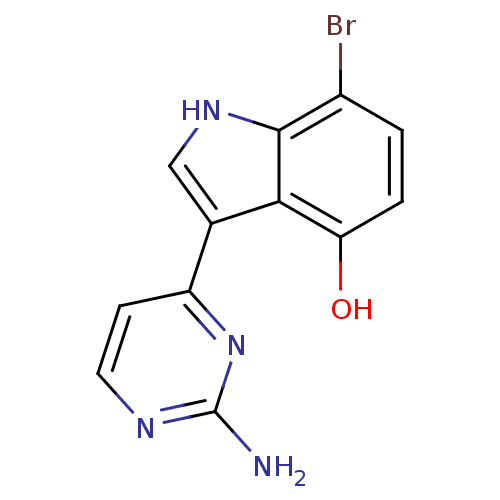
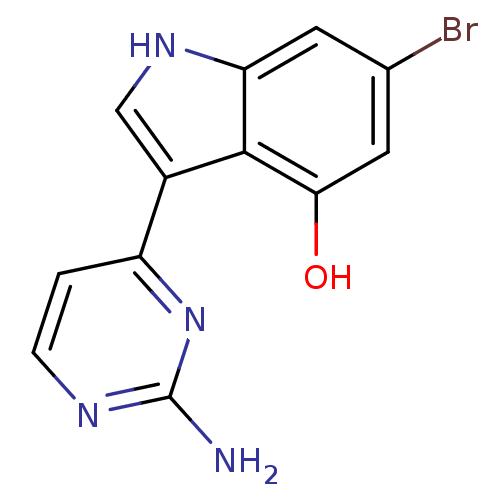
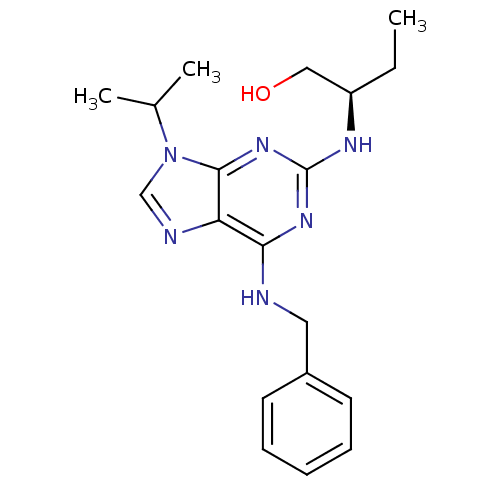
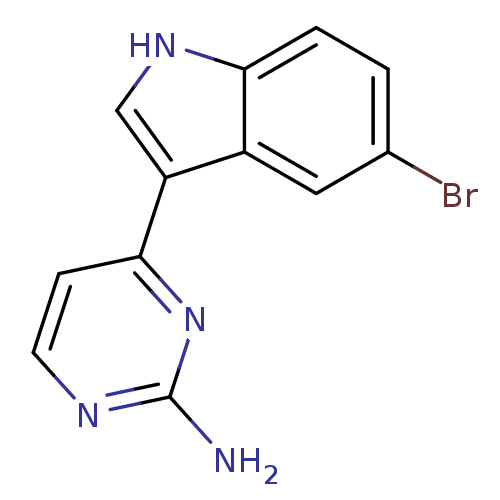
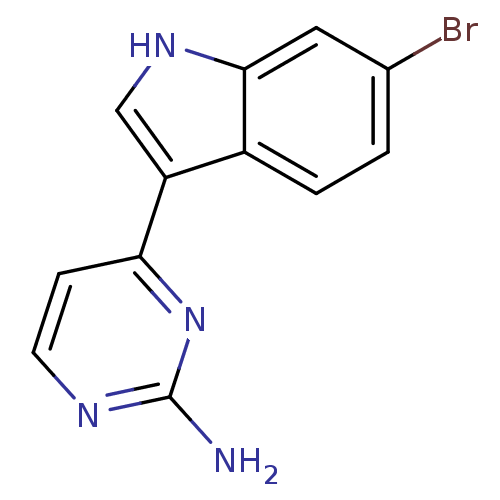
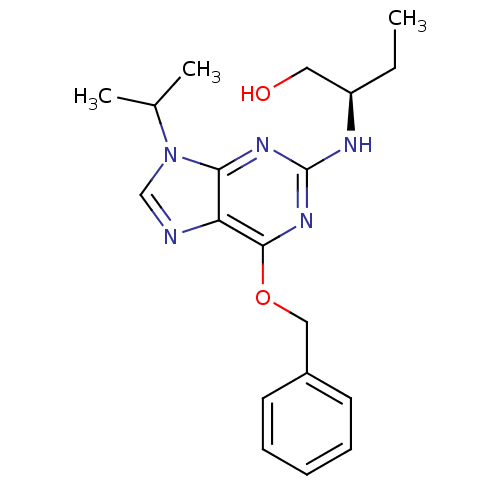
Affinity DataIC50: 400nMT: 2°CAssay Description:Kinase activities were assayed in buffers containing substrate, enzyme, and inhibitor at 30°C in the presence of 15 uM ATP/[gamma-32P] ATP. 32P ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMT: 2°CAssay Description:Kinase activities were assayed in buffers containing substrate, enzyme, and inhibitor at 30°C in the presence of 15 uM ATP/[gamma-32P] ATP. 32P ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.70E+4nMT: 2°CAssay Description:Kinase activities were assayed in buffers containing substrate, enzyme, and inhibitor at 30°C in the presence of 15 uM ATP/[gamma-32P] ATP. 32P ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+4nMT: 2°CAssay Description:Kinase activities were assayed in buffers containing substrate, enzyme, and inhibitor at 30°C in the presence of 15 uM ATP/[gamma-32P] ATP. 32P ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMT: 2°CAssay Description:Kinase activities were assayed in buffers containing substrate, enzyme, and inhibitor at 30°C in the presence of 15 uM ATP/[gamma-32P] ATP. 32P ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMpH: 7.2 T: 2°CAssay Description:Kinase activities were assayed in buffers containing substrate, enzyme, and inhibitor at 30 °C in the presence of 15 uM ATP/[gamma-32P] ATP. 32P...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+6nMpH: 7.2 T: 2°CAssay Description:Kinase activities were assayed in buffers containing substrate, enzyme, and inhibitor at 30 °C in the presence of 15 uM ATP/[gamma-32P] ATP. 32P...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+6nMpH: 7.2 T: 2°CAssay Description:Kinase activities were assayed in buffers containing substrate, enzyme, and inhibitor at 30 °C in the presence of 15 uM ATP/[gamma-32P] ATP. 32P...More data for this Ligand-Target Pair