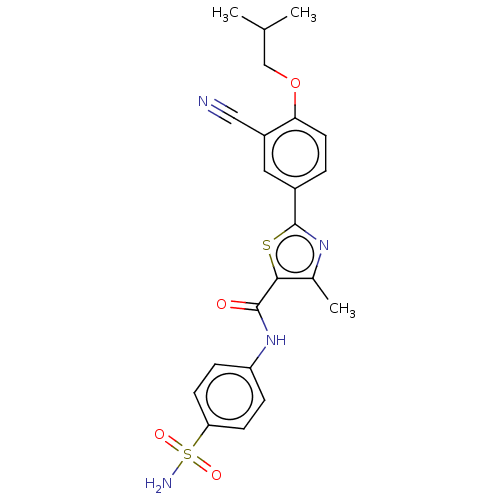
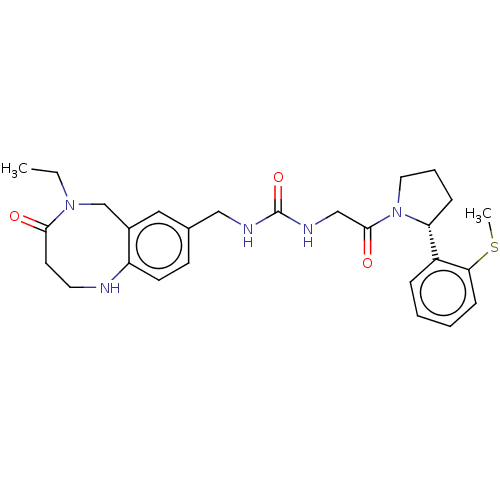
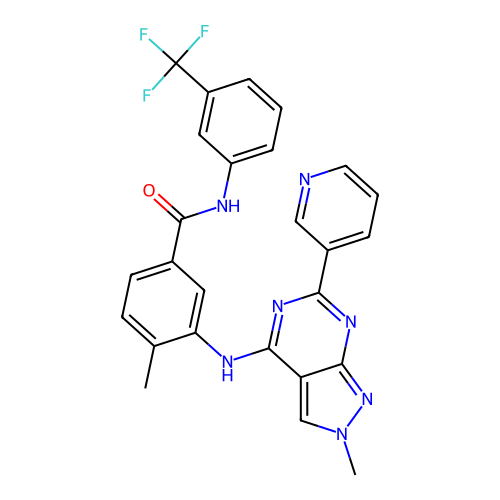
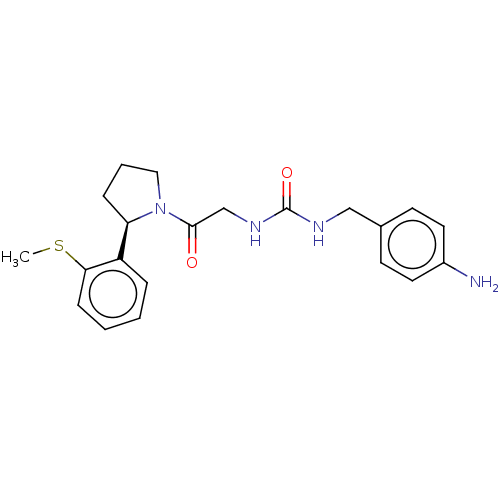
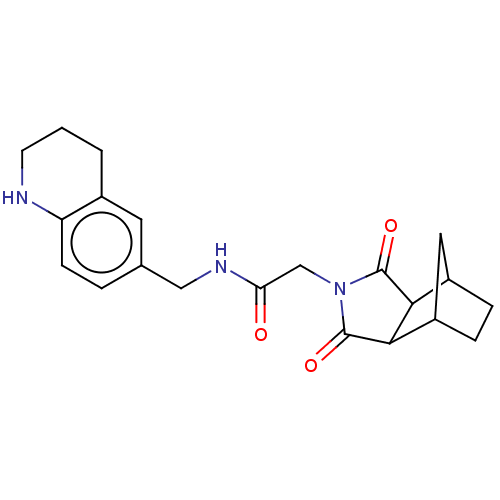
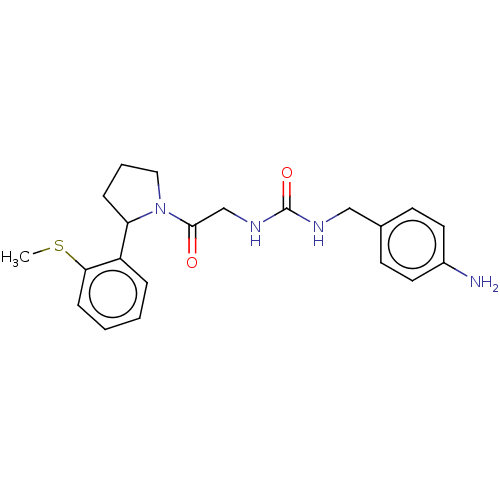
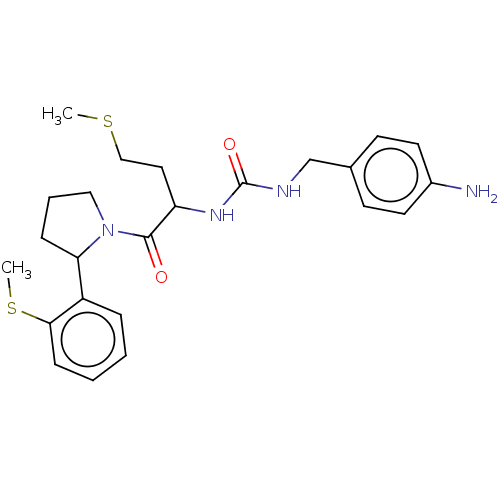
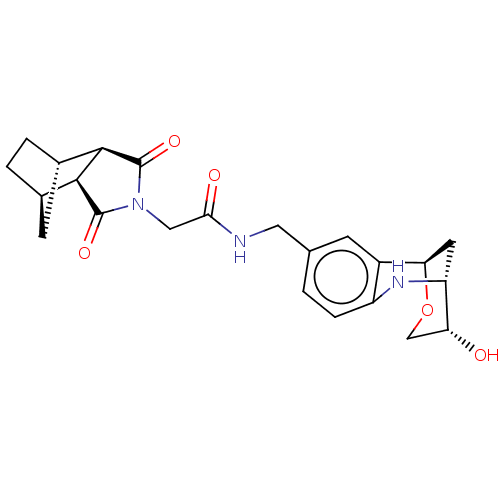
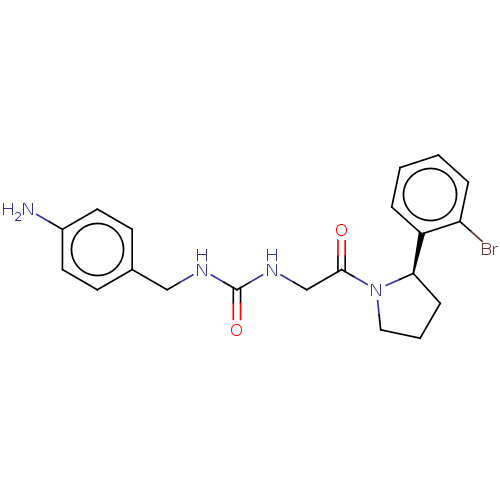
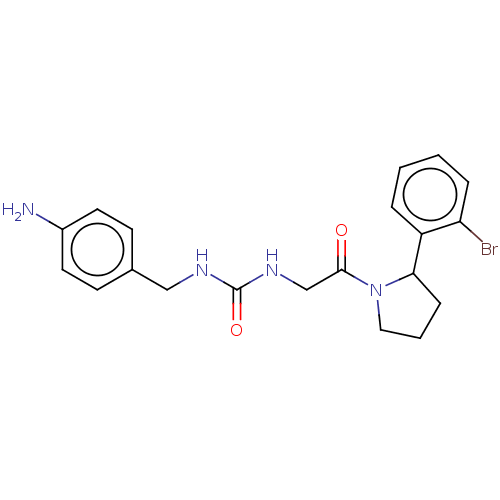
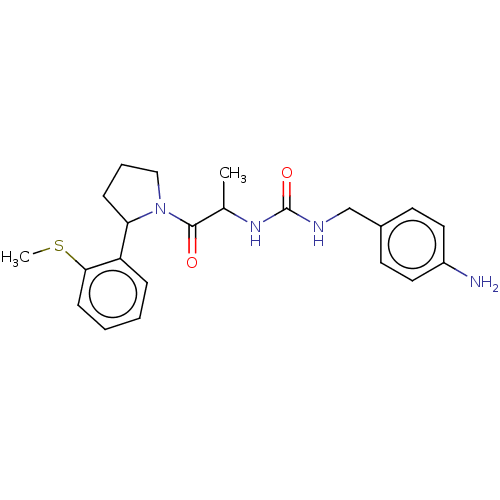
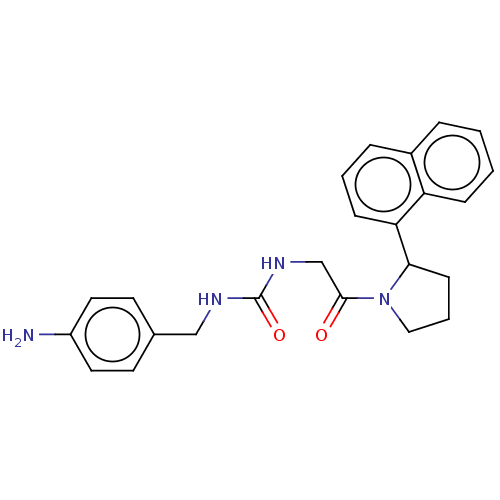
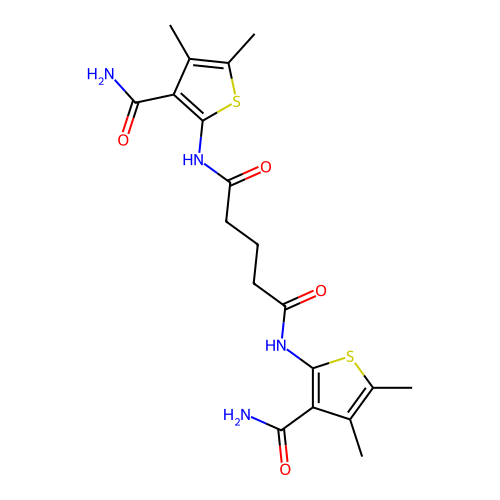
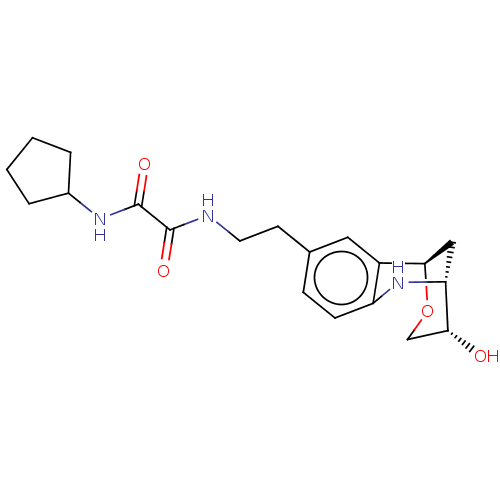
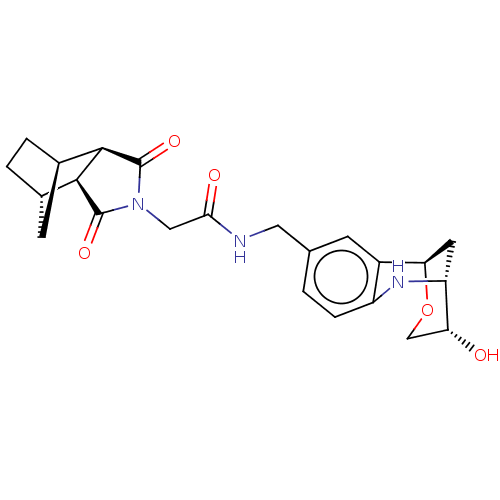
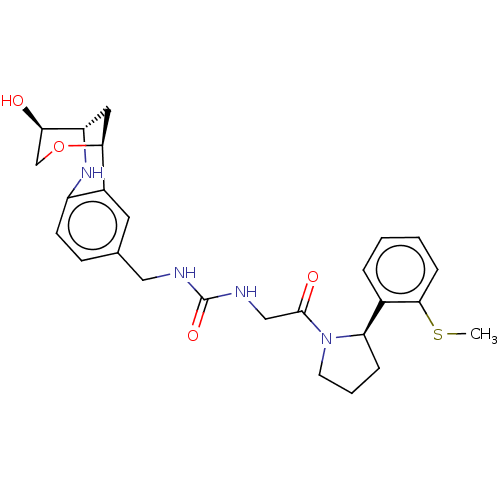
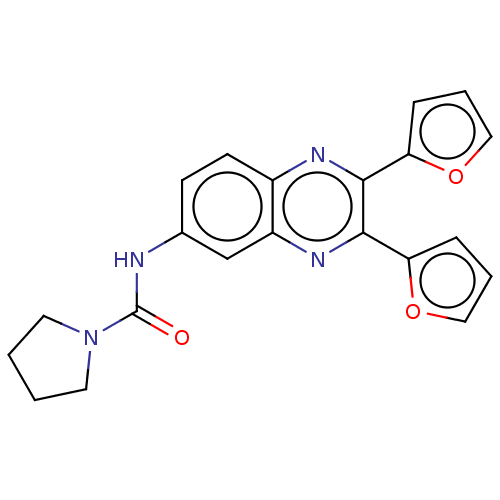
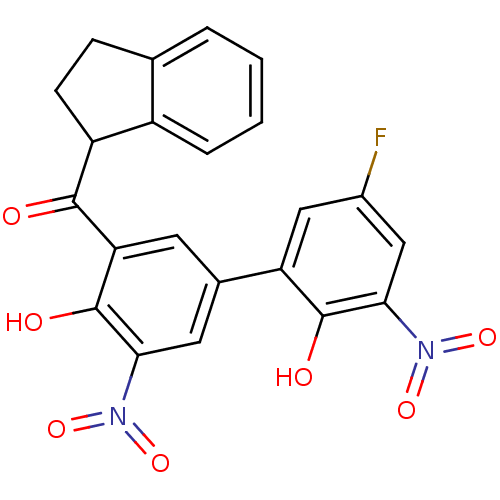
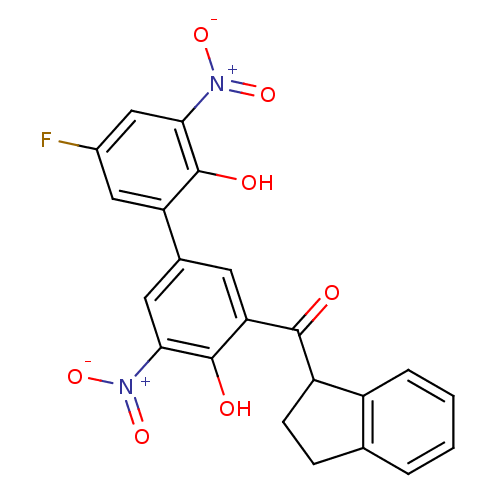
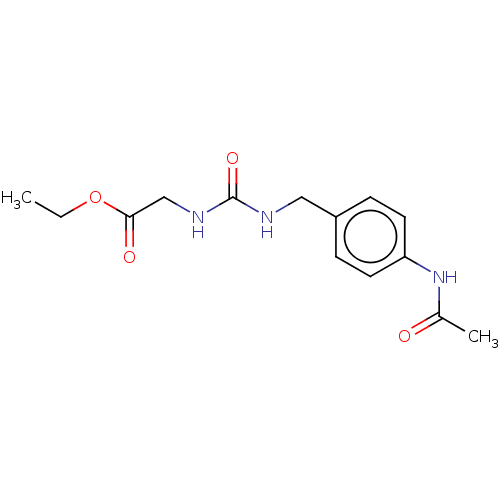
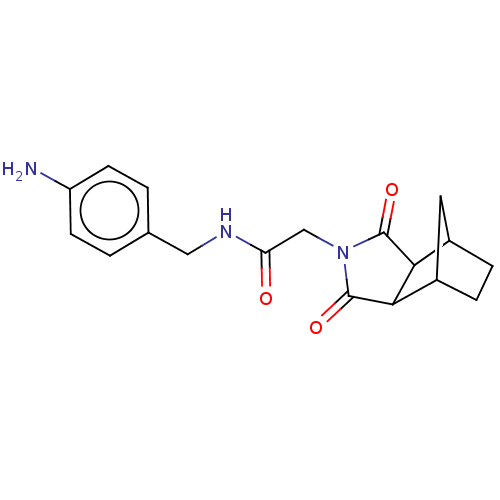
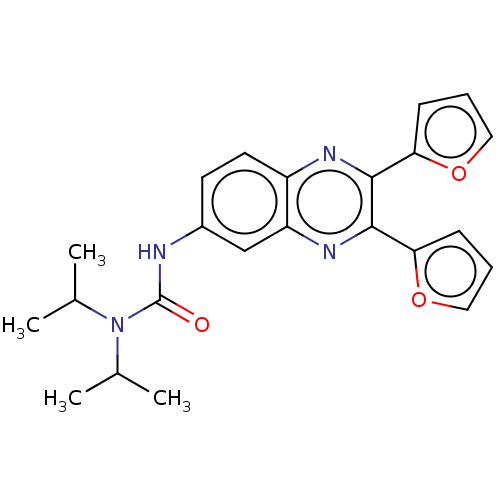
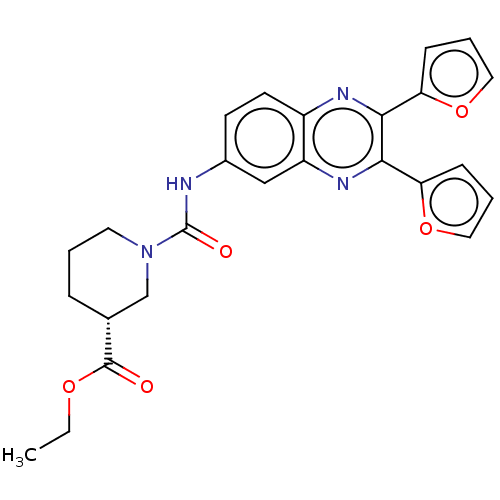
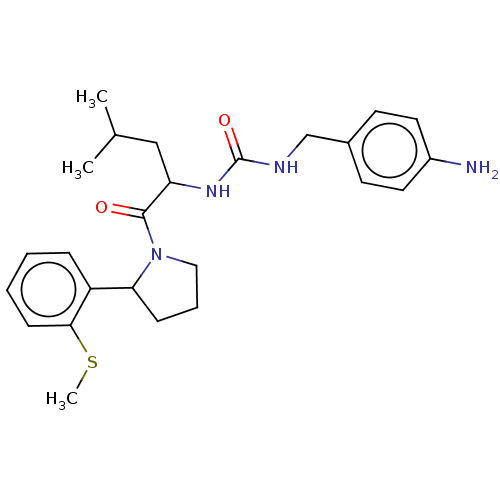
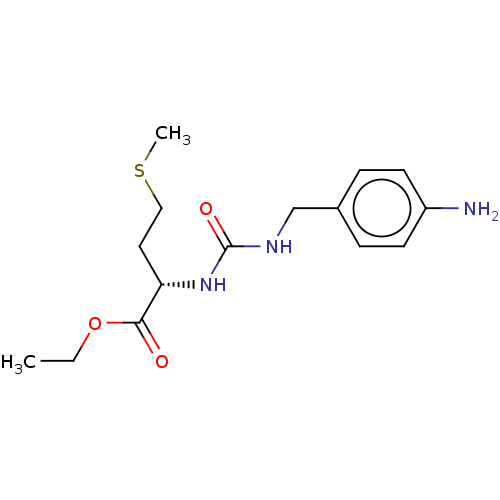
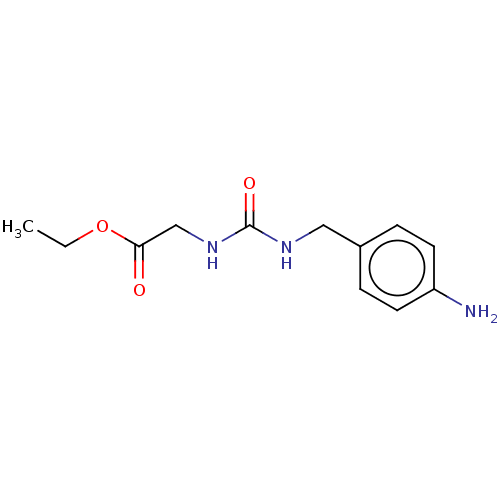
Affinity DataKi: 2.60nMAssay Description:Binding affinity to human recombinant cyclophilin D by surface plasmon resonance analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 3.60nMAssay Description:Inhibition of human recombinant cyclophilin D using Suc-AAPF-MCA as substrate preincubated for 1 hr followed by substrate addition measured per milli...More data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Inhibition of CypD (unknown origin) PPIase activity using N-Suc-AAPF-p-nitroanilide as substrate preincubated with enzyme for 10 mins followed by chy...More data for this Ligand-Target Pair
Affinity DataKi: 7nMAssay Description:Inhibition of fluorescein labeled cyclosporin binding to Cyp40 by fluorescence polarization competition assayMore data for this Ligand-Target Pair
Affinity DataKi: 8.20nMAssay Description:Inhibition of Cyclophilin D (unknown origin) activity in absence of detergentMore data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:Inhibition of Cyclophilin 40 PPIase activityMore data for this Ligand-Target Pair
Affinity DataKd: 13nMAssay Description:Binding affinity to Cyclophilin D (unknown origin) by isothermal titration calorimetryMore data for this Ligand-Target Pair
Affinity DataKd: 30nMAssay Description:Binding affinity to CypD (unknown origin) assessed as dissociation constant by SPR analysisMore data for this Ligand-Target Pair
Affinity DataKi: 34nMAssay Description:Inhibition of Cyclophilin 40 PPIase activityMore data for this Ligand-Target Pair
Affinity DataIC50: 60nMAssay Description:Displacement of fluorescein labelled cyclosporine A derivative from human recombinant CypD (30 to 207 residues) expressed in Escherichia coli cells i...More data for this Ligand-Target Pair
Affinity DataKi: 99nMAssay Description:Inhibition of Cyclophilin D (unknown origin) activity preincubated for 15 mins followed Suc-AAPF-pNA substrate addition by chymotrypsin coupled based...More data for this Ligand-Target Pair
Affinity DataKd: 101nMAssay Description:Inhibition of cyclosporin binding to Cyp40 by fluorescence polarization competition assayMore data for this Ligand-Target Pair
Affinity DataKd: 149nMAssay Description:Binding affinity to human recombinant cyclophilin D by surface plasmon resonance analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 190nMAssay Description:Displacement of fluorescein labelled cyclosporine A derivative from human recombinant CypD (30 to 207 residues) expressed in Escherichia coli cells i...More data for this Ligand-Target Pair
Affinity DataKd: 191nMAssay Description:Binding affinity to human PPID incubated for 45 mins by Kinobead based pull down assayMore data for this Ligand-Target Pair
Affinity DataKd: 230nMAssay Description:Binding affinity to CypD (unknown origin) assessed as dissociation constant by SPR analysisMore data for this Ligand-Target Pair
Affinity DataKd: 410nMAssay Description:Binding affinity to Cyclophilin D (unknown origin) by isothermal titration calorimetryMore data for this Ligand-Target Pair
Affinity DataIC50: 600nMAssay Description:Inhibition of CypD (unknown origin) PPIase activity using N-Suc-AAPF-p-nitroanilide as substrate preincubated with enzyme for 10 mins followed by chy...More data for this Ligand-Target Pair
Affinity DataIC50: 640nMpH: 7.8 T: 2°CAssay Description:Cyclophilin PPlase activity was measured at 20 C. by using the standard chymotrypsin coupled assay (Kofron J L, Kuzmic P, Kishore V, Colon-Bonilla E,...More data for this Ligand-Target Pair
Affinity DataIC50: 660nMpH: 7.8 T: 2°CAssay Description:Cyclophilin PPlase activity was measured at 20 C. by using the standard chymotrypsin coupled assay (Kofron J L, Kuzmic P, Kishore V, Colon-Bonilla E,...More data for this Ligand-Target Pair
Affinity DataIC50: 735nMAssay Description:Displacement of fluorescein labelled cyclosporine A derivative from human recombinant CypD (30 to 207 residues) expressed in Escherichia coli cells i...More data for this Ligand-Target Pair
Affinity DataKi: 950nMAssay Description:Inhibition of Cyclophilin D (unknown origin) activity preincubated for 15 mins followed Suc-AAPF-pNA substrate addition by chymotrypsin coupled based...More data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+3nMpH: 7.8 T: 2°CAssay Description:Cyclophilin PPlase activity was measured at 20 C. by using the standard chymotrypsin coupled assay (Kofron J L, Kuzmic P, Kishore V, Colon-Bonilla E,...More data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+3nMpH: 7.8 T: 2°CAssay Description:Cyclophilin PPlase activity was measured at 20 C. by using the standard chymotrypsin coupled assay (Kofron J L, Kuzmic P, Kishore V, Colon-Bonilla E,...More data for this Ligand-Target Pair
Affinity DataKd: 1.20E+3nMAssay Description:Binding affinity to Cyclophilin D (unknown origin) by isothermal titration calorimetryMore data for this Ligand-Target Pair
Affinity DataKi: 1.33E+3nMAssay Description:Binding affinity to human recombinant cyclophilin D by surface plasmon resonance analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.35E+3nMAssay Description:Displacement of fluorescein labelled cyclosporine A derivative from human recombinant CypD (30 to 207 residues) expressed in Escherichia coli cells i...More data for this Ligand-Target Pair
Affinity DataIC50: 1.40E+3nMpH: 7.8 T: 2°CAssay Description:Cyclophilin PPlase activity was measured at 20 C. by using the standard chymotrypsin coupled assay (Kofron J L, Kuzmic P, Kishore V, Colon-Bonilla E,...More data for this Ligand-Target Pair
Affinity DataIC50: 1.49E+3nMAssay Description:Inhibition of human recombinant cyclophilin D using Suc-AAPF-MCA as substrate preincubated for 1 hr followed by substrate addition measured per milli...More data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+3nMAssay Description:Inhibition of GST-tagged human CYP40 expressed in Escherichia coli BL21 (DE3) using NH2-EDASRMEEVD-COOH peptide as substrate preincubated for 15 mins...More data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+3nMAssay Description:Inhibition of CypD (unknown origin) PPIase activity using N-Suc-AAPF-p-nitroanilide as substrate preincubated with enzyme for 10 mins followed by chy...More data for this Ligand-Target Pair
Affinity DataIC50: 1.70E+3nMAssay Description:Displacement of fluorescein labelled cyclosporine A derivative from human recombinant CypD (30 to 207 residues) expressed in Escherichia coli cells i...More data for this Ligand-Target Pair
Affinity DataIC50: 1.70E+3nMAssay Description:Displacement of fluorescein labelled cyclosporine A derivative from human recombinant CypD (30 to 207 residues) expressed in Escherichia coli cells i...More data for this Ligand-Target Pair
Affinity DataKi: 2.00E+3nMAssay Description:Binding affinity to CypD (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMAssay Description:Displacement of fluorescein labelled cyclosporine A derivative from human recombinant CypD (30 to 207 residues) expressed in Escherichia coli cells i...More data for this Ligand-Target Pair
Affinity DataKi: 2.42E+3nM ΔG°: -29.9kJ/molepH: 7.8 T: 2°CAssay Description:PPIase activity assay were performed at 283 K in quartz cuvettes with a path length of 1 cm under vigorous stirring with a Hewlett-Packard 8453A UV-v...More data for this Ligand-Target Pair
Affinity DataIC50: 2.42E+3nMAssay Description:Inhibition of CypD (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 2.60E+3nMAssay Description:Inhibition of Cyclophilin D (unknown origin) activity preincubated for 15 mins followed Suc-AAPF-pNA substrate addition by chymotrypsin coupled based...More data for this Ligand-Target Pair
Affinity DataIC50: 2.80E+3nMAssay Description:Displacement of fluorescein labelled cyclosporine A derivative from human recombinant CypD (30 to 207 residues) expressed in Escherichia coli cells i...More data for this Ligand-Target Pair
Affinity DataKd: 2.80E+3nMAssay Description:Binding affinity to Cyclophilin D (unknown origin) by isothermal titration calorimetryMore data for this Ligand-Target Pair
Affinity DataKi: 3.00E+3nMAssay Description:Binding affinity to CypD (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 3.00E+3nMAssay Description:Binding affinity to CypD (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+3nMpH: 7.8 T: 2°CAssay Description:Cyclophilin PPlase activity was measured at 20 C. by using the standard chymotrypsin coupled assay (Kofron J L, Kuzmic P, Kishore V, Colon-Bonilla E,...More data for this Ligand-Target Pair
Affinity DataIC50: 3.50E+3nMAssay Description:Displacement of fluorescein labelled cyclosporine A derivative from human recombinant CypD (30 to 207 residues) expressed in Escherichia coli cells i...More data for this Ligand-Target Pair
Affinity DataIC50: 3.60E+3nMAssay Description:Displacement of fluorescein labelled cyclosporine A derivative from human recombinant CypD (30 to 207 residues) expressed in Escherichia coli cells i...More data for this Ligand-Target Pair
Affinity DataIC50: 4.10E+3nMAssay Description:Displacement of fluorescein labelled cyclosporine A derivative from human recombinant CypD (30 to 207 residues) expressed in Escherichia coli cells i...More data for this Ligand-Target Pair
Affinity DataKi: 4.90E+3nMAssay Description:Inhibition of Cyclophilin D (unknown origin) activity preincubated for 15 mins followed Suc-AAPF-pNA substrate addition by chymotrypsin coupled based...More data for this Ligand-Target Pair
Affinity DataKd: 5.12E+3nMAssay Description:Binding affinity to Cyclophilin D (unknown origin) by isothermal titration calorimetryMore data for this Ligand-Target Pair
Affinity DataKd: 5.40E+3nMAssay Description:Binding affinity to Cyclophilin D (unknown origin) by isothermal titration calorimetryMore data for this Ligand-Target Pair
Affinity DataKi: 5.90E+3nMAssay Description:Inhibition of Cyclophilin D (unknown origin) activity preincubated for 15 mins followed Suc-AAPF-pNA substrate addition by chymotrypsin coupled based...More data for this Ligand-Target Pair