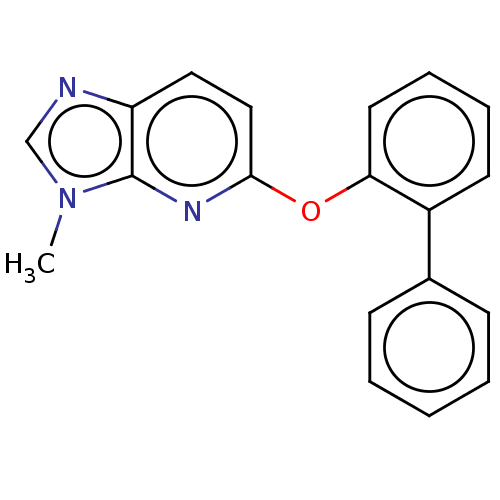
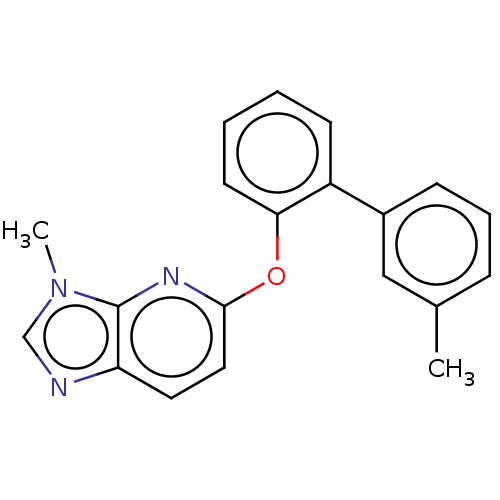
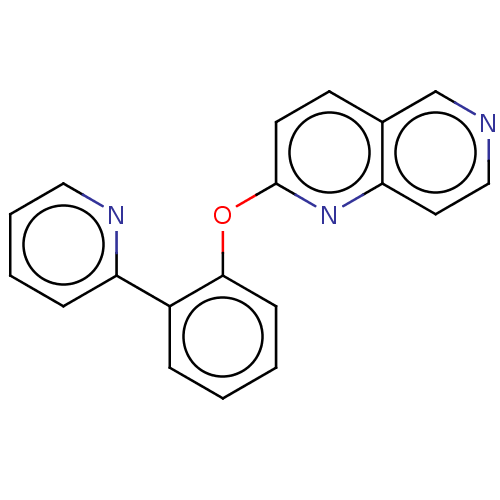
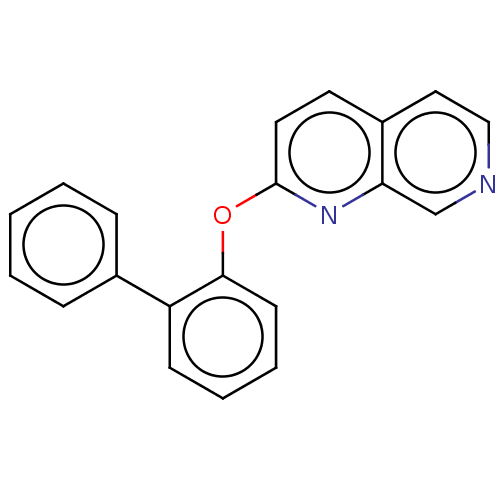
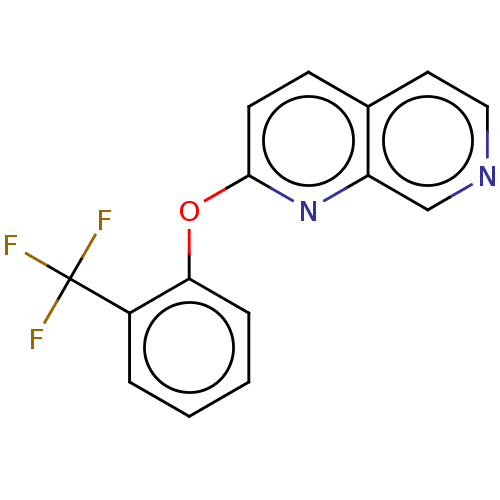
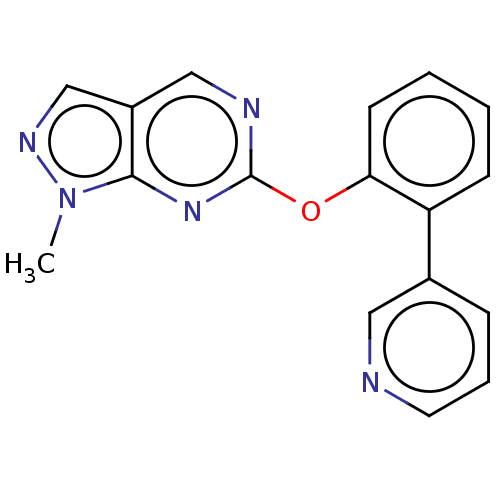
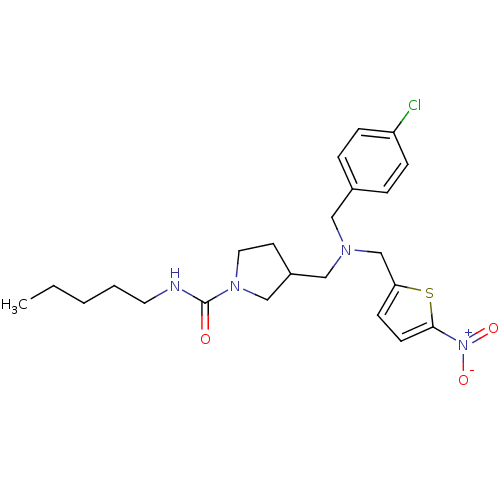
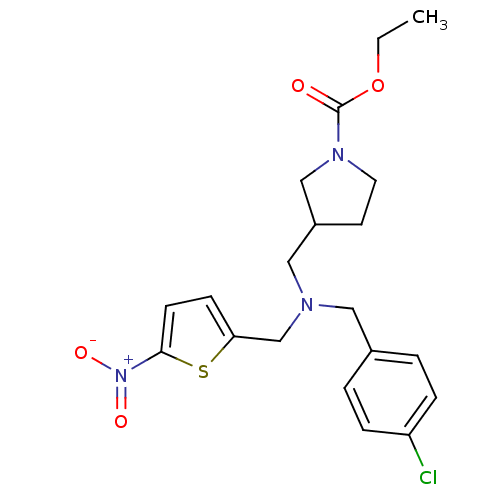
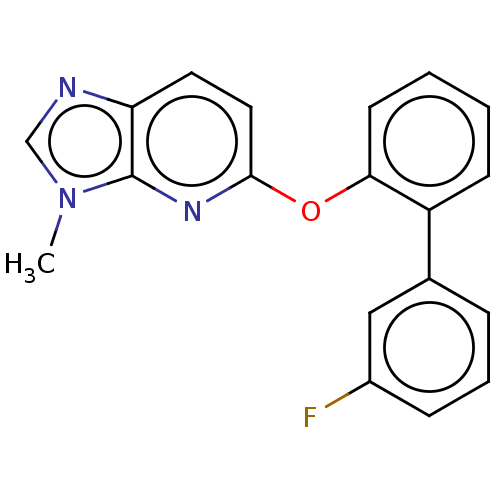
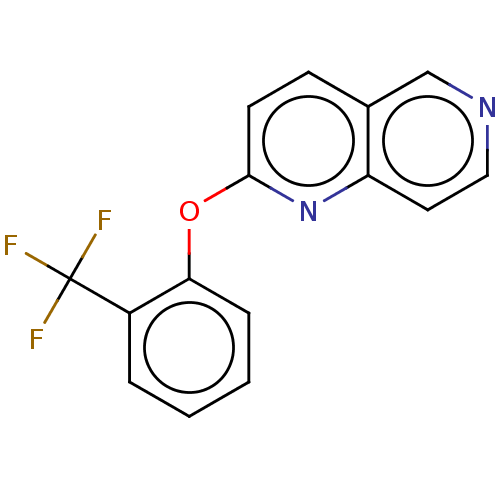
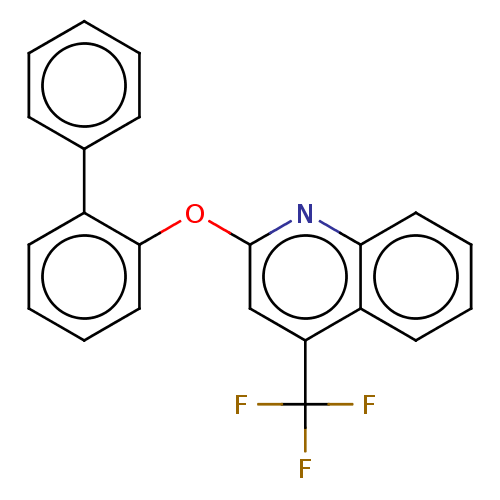
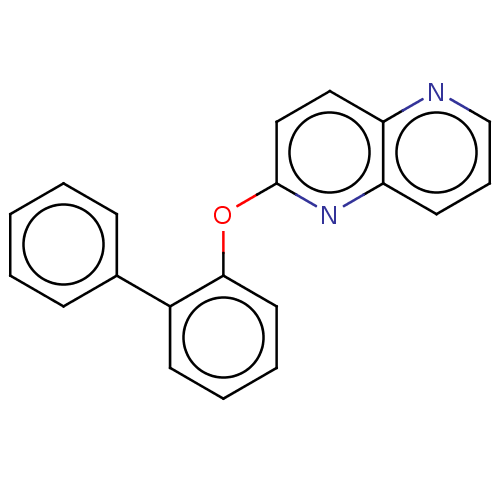
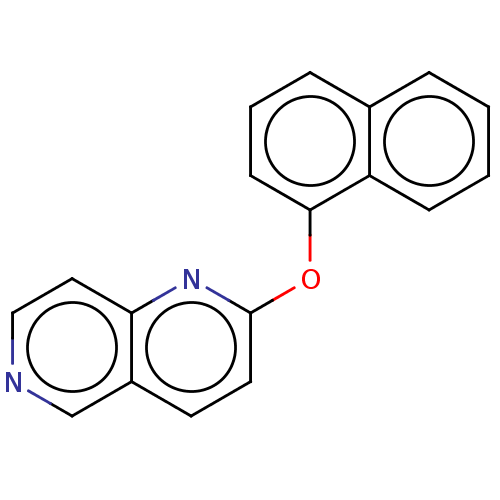
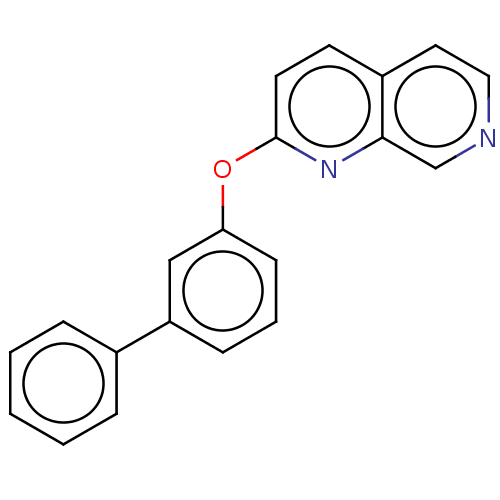
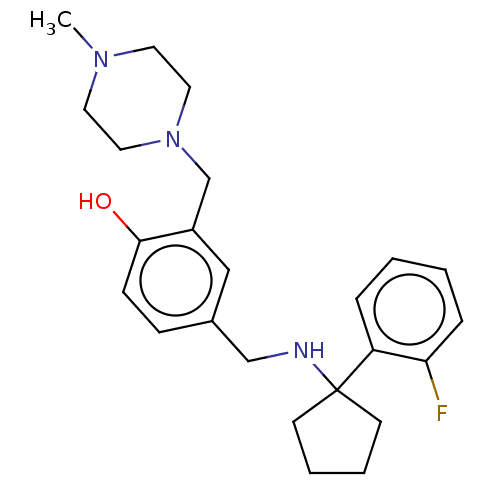
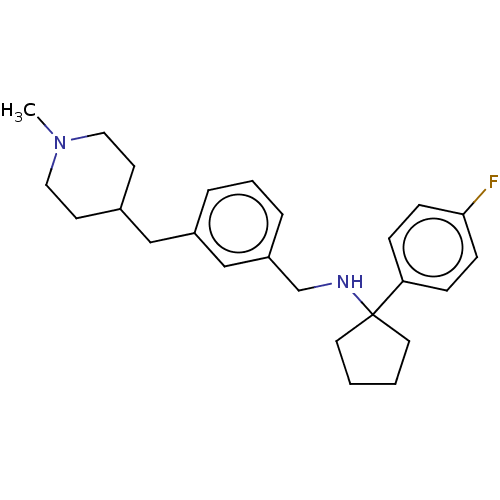Affinity DataIC50: 250nMAssay Description:The capacity for compounds to function as ligands for either REV-ERBα or REV-ERBβ was assessed using a fluorescent resonance energy transfe...More data for this Ligand-Target Pair
Affinity DataIC50: 250nMAssay Description:The capacity for compounds to function as ligands for either REV-ERBα or REV-ERBβ was assessed using a fluorescent resonance energy transfe...More data for this Ligand-Target Pair
Affinity DataIC50: 250nMAssay Description:The capacity for compounds to function as ligands for either REV-ERBα or REV-ERBβ was assessed using a fluorescent resonance energy transfe...More data for this Ligand-Target Pair
Affinity DataIC50: 250nMAssay Description:The capacity for compounds to function as ligands for either REV-ERBα or REV-ERBβ was assessed using a fluorescent resonance energy transfe...More data for this Ligand-Target Pair
Affinity DataIC50: 250nMAssay Description:The capacity for compounds to function as ligands for either REV-ERBα or REV-ERBβ was assessed using a fluorescent resonance energy transfe...More data for this Ligand-Target Pair
Affinity DataIC50: 250nMAssay Description:The capacity for compounds to function as ligands for either REV-ERBα or REV-ERBβ was assessed using a fluorescent resonance energy transfe...More data for this Ligand-Target Pair
Affinity DataIC50: 560nMAssay Description:Increased REV-ERB-beta LBD dependent repressor activity in HEK293 cell reporter assayMore data for this Ligand-Target Pair
Affinity DataIC50: 800nMAssay Description:Increased REV-ERB-beta LBD dependent repressor activity in HEK293 cell reporter assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:The capacity for compounds to function as ligands for either REV-ERBα or REV-ERBβ was assessed using a fluorescent resonance energy transfe...More data for this Ligand-Target Pair
Affinity DataIC50: 1.34E+3nMAssay Description:Inhibition of REV-ERBbeta (unknown origin) expressed in HEK293 cells incubated for 24 hrs assessed as inhibition of receptor-mediated transcriptional...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:The capacity for compounds to function as ligands for either REV-ERBα or REV-ERBβ was assessed using a fluorescent resonance energy transfe...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:The capacity for compounds to function as ligands for either REV-ERBα or REV-ERBβ was assessed using a fluorescent resonance energy transfe...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:The capacity for compounds to function as ligands for either REV-ERBα or REV-ERBβ was assessed using a fluorescent resonance energy transfe...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:The capacity for compounds to function as ligands for either REV-ERBα or REV-ERBβ was assessed using a fluorescent resonance energy transfe...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:The capacity for compounds to function as ligands for either REV-ERBα or REV-ERBβ was assessed using a fluorescent resonance energy transfe...More data for this Ligand-Target Pair
Affinity DataIC50: 1.75E+4nMAssay Description:Inhibition of REV-ERBbeta (unknown origin) expressed in HEK293 cells incubated for 24 hrs assessed as inhibition of receptor-mediated transcriptional...More data for this Ligand-Target Pair