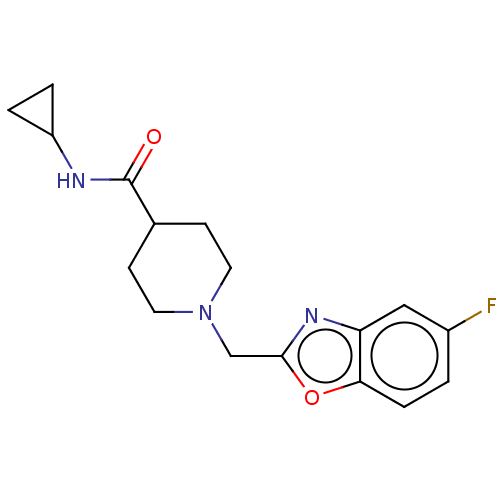
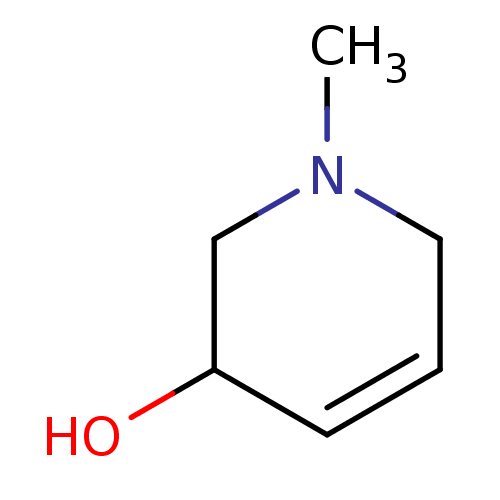
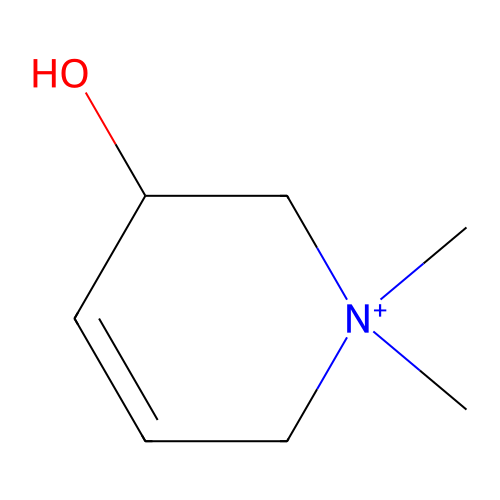
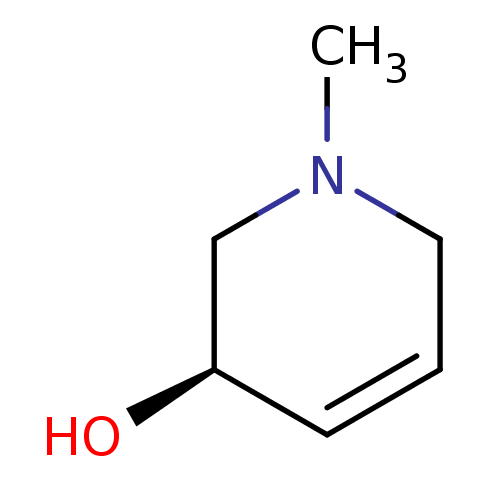
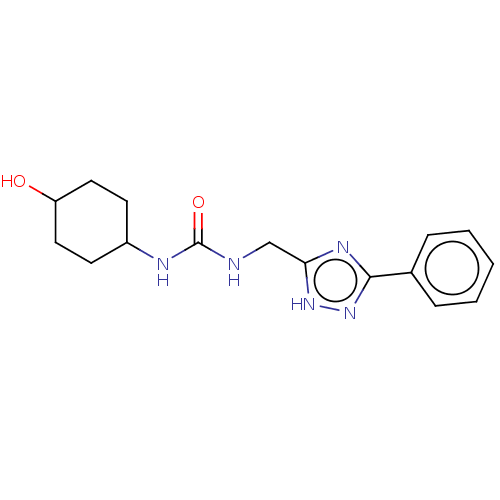
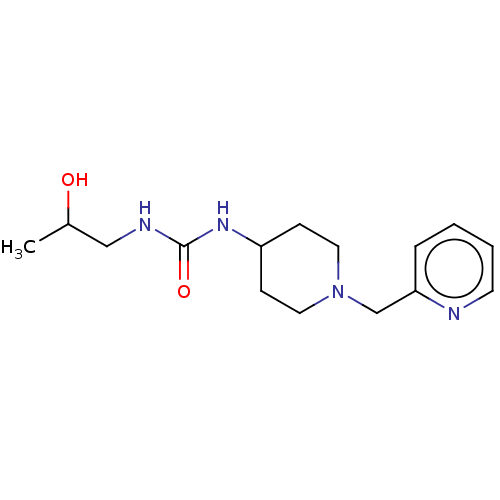
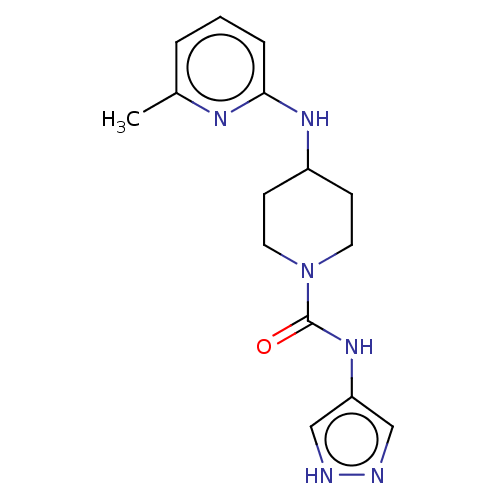
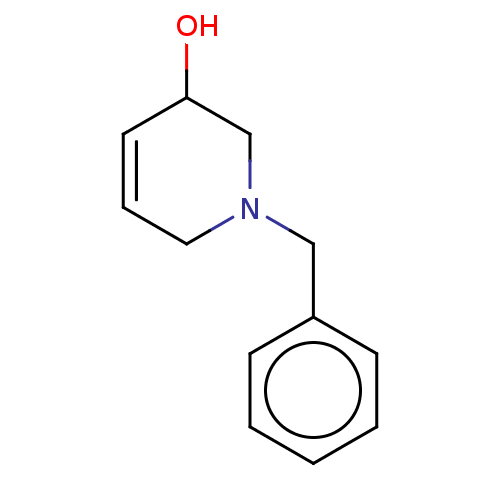
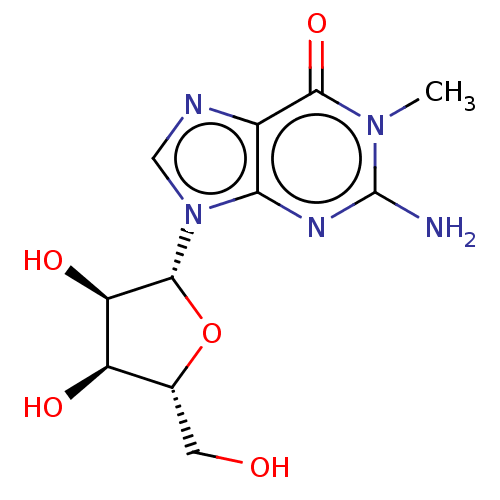
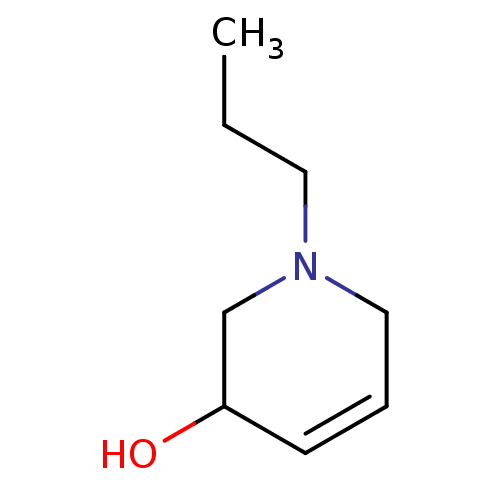
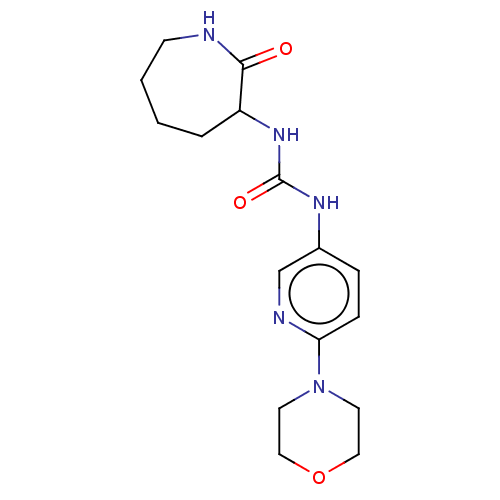
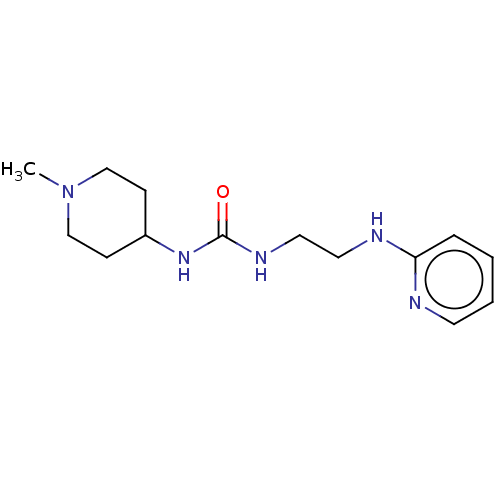
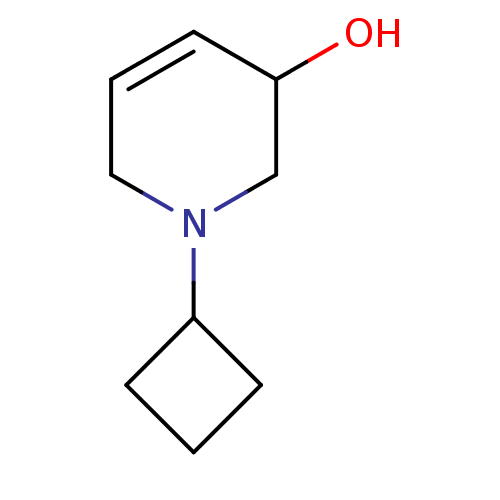
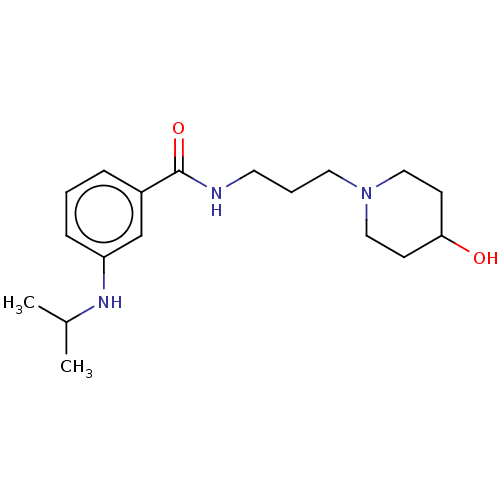
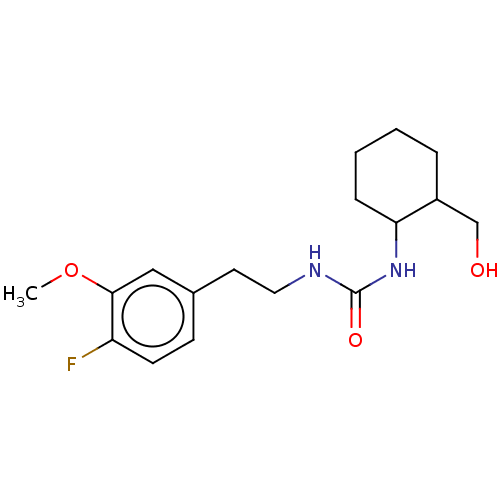
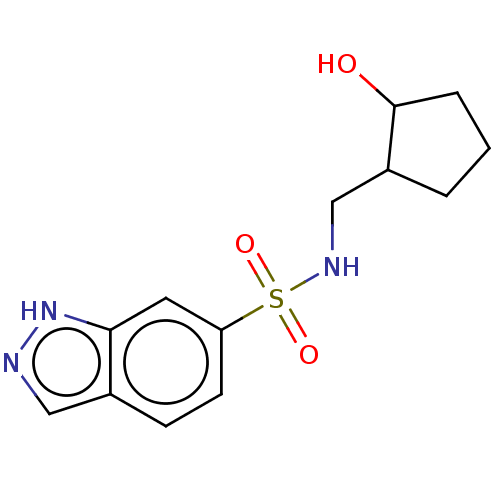
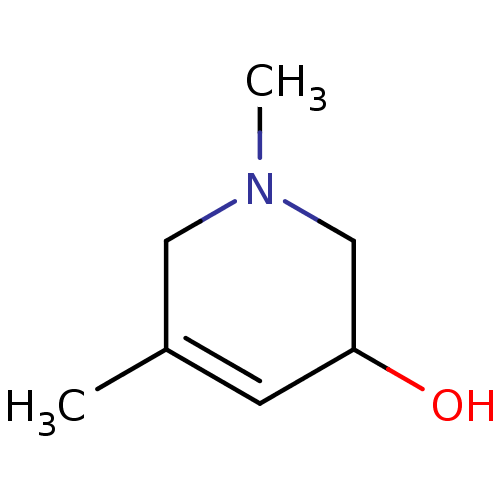
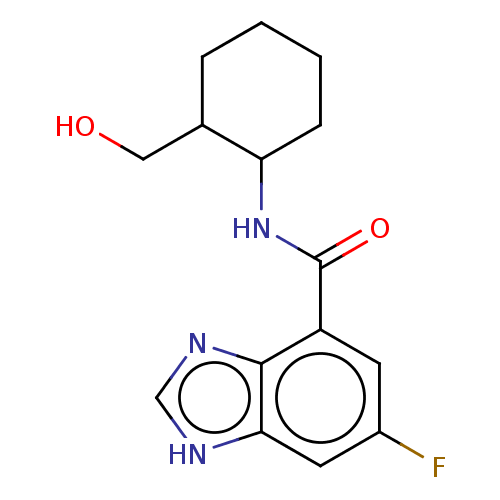
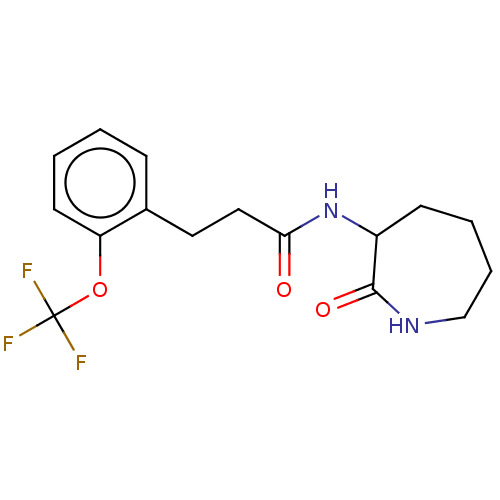
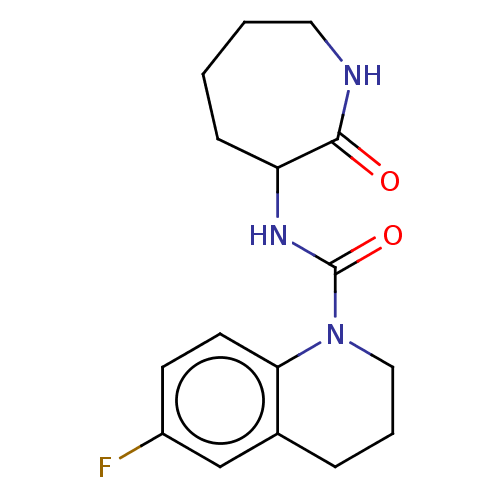
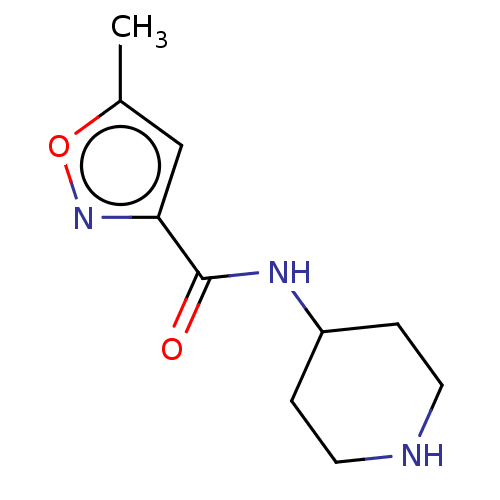
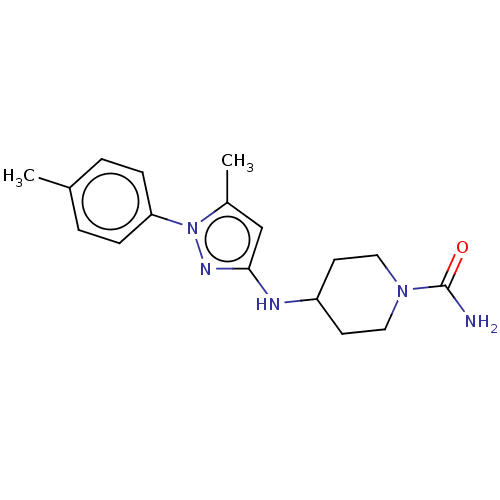
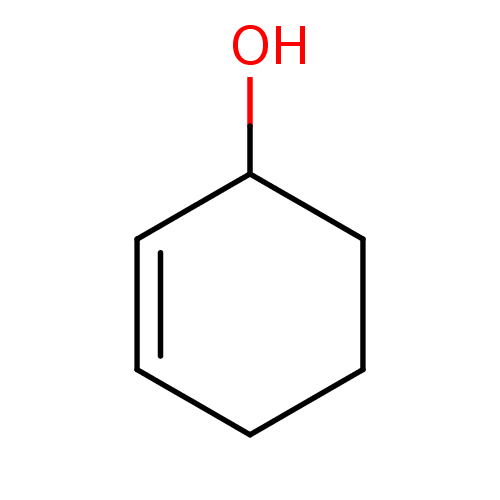
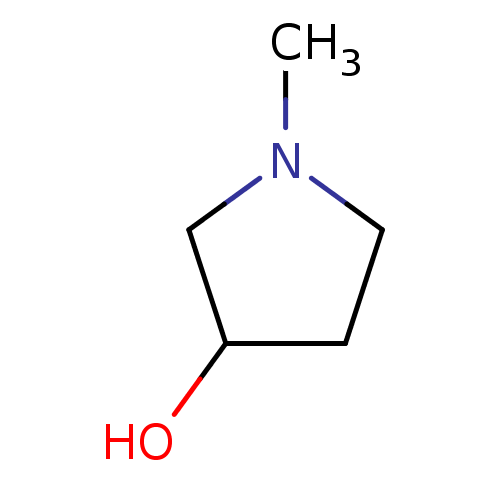
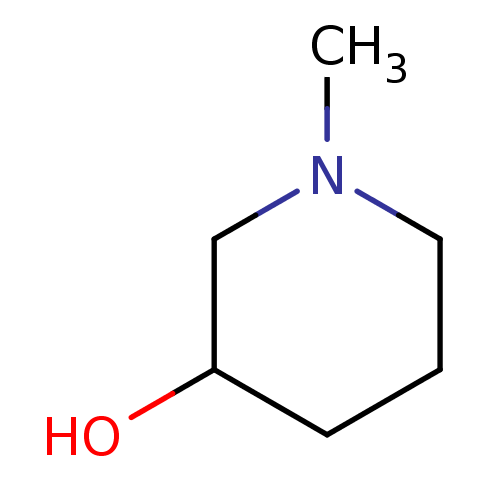
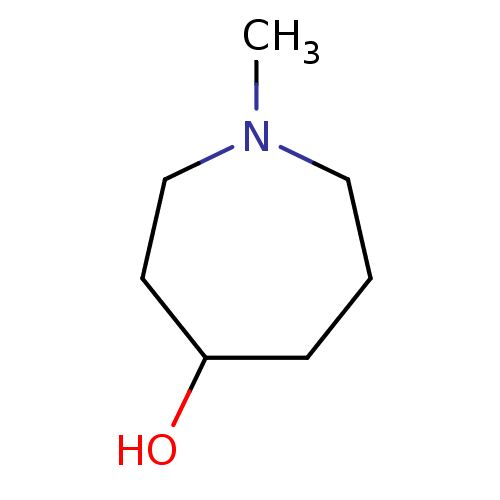
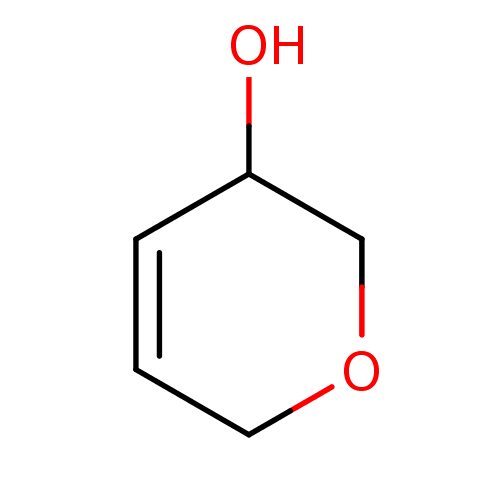
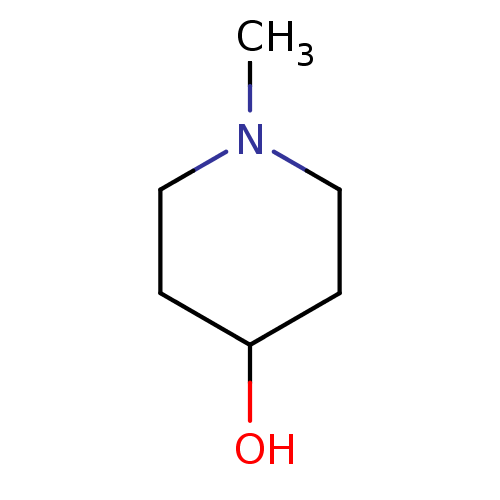
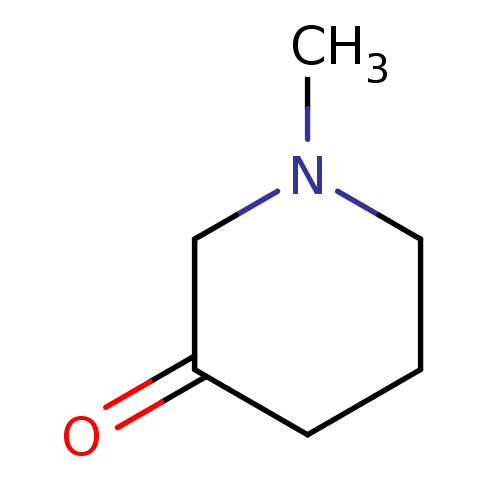
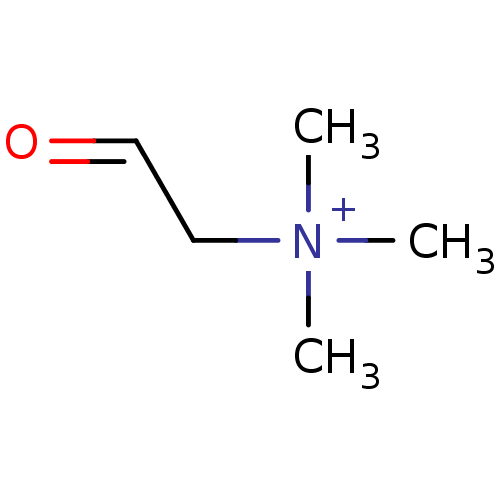TargetCholine trimethylamine-lyase(Desulfovibrio alaskensis (strain ATCC BAA 1058 / D...)
Harvard University
Curated by ChEMBL
Harvard University
Curated by ChEMBL
Affinity DataIC50: 2.40E+3nMAssay Description:Inhibition of Desulfovibrio desulfuricans (strain G20) CutC assessed as reduction in TMA by spectrophotometryMore data for this Ligand-Target Pair
TargetCholine trimethylamine-lyase(Desulfovibrio alaskensis (strain ATCC BAA 1058 / D...)
Harvard University
Curated by ChEMBL
Harvard University
Curated by ChEMBL
Affinity DataIC50: 2.40E+3nMAssay Description:Inhibition of N-terminal His6-tagged Desulfovibrio desulfuricans G20 CutC encoding 52 residues assessed as reduction in acetaldehyde using choline as...More data for this Ligand-Target Pair
TargetCholine trimethylamine-lyase(Desulfovibrio alaskensis (strain ATCC BAA 1058 / D...)
Harvard University
Curated by ChEMBL
Harvard University
Curated by ChEMBL
Affinity DataIC50: 2.90E+3nMAssay Description:Inhibition of Desulfovibrio desulfuricans (strain G20) CutC assessed as reduction in TMA by spectrophotometryMore data for this Ligand-Target Pair
TargetCholine trimethylamine-lyase(Desulfovibrio alaskensis (strain ATCC BAA 1058 / D...)
Harvard University
Curated by ChEMBL
Harvard University
Curated by ChEMBL
Affinity DataIC50: 3.90E+3nMAssay Description:Inhibition of N-terminal His6-tagged Desulfovibrio desulfuricans G20 CutC encoding 52 residues assessed as reduction in acetaldehyde using choline as...More data for this Ligand-Target Pair
TargetCholine trimethylamine-lyase(Desulfovibrio alaskensis (strain ATCC BAA 1058 / D...)
Harvard University
Curated by ChEMBL
Harvard University
Curated by ChEMBL
Affinity DataIC50: 6.00E+3nMAssay Description:Inhibition of Desulfovibrio desulfuricans (strain G20) CutC assessed as reduction in TMA by spectrophotometryMore data for this Ligand-Target Pair
TargetCholine trimethylamine-lyase(Desulfovibrio alaskensis (strain ATCC BAA 1058 / D...)
Harvard University
Curated by ChEMBL
Harvard University
Curated by ChEMBL
Affinity DataIC50: 7.00E+3nMAssay Description:Inhibition of Desulfovibrio desulfuricans (strain G20) CutC assessed as reduction in TMA by spectrophotometryMore data for this Ligand-Target Pair
TargetCholine trimethylamine-lyase(Desulfovibrio alaskensis (strain ATCC BAA 1058 / D...)
Harvard University
Curated by ChEMBL
Harvard University
Curated by ChEMBL
Affinity DataIC50: 2.20E+4nMAssay Description:Inhibition of Desulfovibrio desulfuricans (strain G20) CutC assessed as reduction in TMA by spectrophotometryMore data for this Ligand-Target Pair
TargetCholine trimethylamine-lyase(Desulfovibrio alaskensis (strain ATCC BAA 1058 / D...)
Harvard University
Curated by ChEMBL
Harvard University
Curated by ChEMBL
Affinity DataIC50: 2.60E+4nMAssay Description:Inhibition of N-terminal His6-tagged Desulfovibrio desulfuricans G20 CutC encoding 52 residues assessed as reduction in acetaldehyde using choline as...More data for this Ligand-Target Pair
TargetCholine trimethylamine-lyase(Desulfovibrio alaskensis (strain ATCC BAA 1058 / D...)
Harvard University
Curated by ChEMBL
Harvard University
Curated by ChEMBL
Affinity DataIC50: 4.90E+4nMAssay Description:Inhibition of N-terminal His6-tagged Desulfovibrio desulfuricans G20 CutC encoding 52 residues assessed as reduction in acetaldehyde using choline as...More data for this Ligand-Target Pair
TargetCholine trimethylamine-lyase(Desulfovibrio alaskensis (strain ATCC BAA 1058 / D...)
Harvard University
Curated by ChEMBL
Harvard University
Curated by ChEMBL
Affinity DataIC50: 7.30E+4nMAssay Description:Inhibition of N-terminal His6-tagged Desulfovibrio desulfuricans G20 CutC encoding 52 residues assessed as reduction in acetaldehyde using choline as...More data for this Ligand-Target Pair
TargetCholine trimethylamine-lyase(Desulfovibrio alaskensis (strain ATCC BAA 1058 / D...)
Harvard University
Curated by ChEMBL
Harvard University
Curated by ChEMBL
Affinity DataIC50: 8.50E+4nMAssay Description:Inhibition of N-terminal His6-tagged Desulfovibrio desulfuricans G20 CutC encoding 52 residues assessed as reduction in acetaldehyde using choline as...More data for this Ligand-Target Pair
TargetCholine trimethylamine-lyase(Desulfovibrio alaskensis (strain ATCC BAA 1058 / D...)
Harvard University
Curated by ChEMBL
Harvard University
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of Desulfovibrio desulfuricans (strain G20) CutC assessed as reduction in TMA by spectrophotometryMore data for this Ligand-Target Pair
TargetCholine trimethylamine-lyase(Desulfovibrio alaskensis (strain ATCC BAA 1058 / D...)
Harvard University
Curated by ChEMBL
Harvard University
Curated by ChEMBL
Affinity DataIC50: 1.10E+5nMAssay Description:Inhibition of N-terminal His6-tagged Desulfovibrio desulfuricans G20 CutC encoding 52 residues assessed as reduction in acetaldehyde using choline as...More data for this Ligand-Target Pair
TargetCholine trimethylamine-lyase(Desulfovibrio alaskensis (strain ATCC BAA 1058 / D...)
Harvard University
Curated by ChEMBL
Harvard University
Curated by ChEMBL
Affinity DataIC50: 1.40E+5nMAssay Description:Inhibition of Desulfovibrio desulfuricans (strain G20) CutC assessed as reduction in TMA by spectrophotometryMore data for this Ligand-Target Pair
TargetCholine trimethylamine-lyase(Desulfovibrio alaskensis (strain ATCC BAA 1058 / D...)
Harvard University
Curated by ChEMBL
Harvard University
Curated by ChEMBL
Affinity DataIC50: 1.67E+5nMAssay Description:Inhibition of N-terminal His6-tagged Desulfovibrio desulfuricans G20 CutC encoding 52 residues assessed as reduction in acetaldehyde using choline as...More data for this Ligand-Target Pair
TargetCholine trimethylamine-lyase(Desulfovibrio alaskensis (strain ATCC BAA 1058 / D...)
Harvard University
Curated by ChEMBL
Harvard University
Curated by ChEMBL
Affinity DataIC50: 1.99E+5nMAssay Description:Inhibition of N-terminal His6-tagged Desulfovibrio desulfuricans G20 CutC encoding 52 residues assessed as reduction in acetaldehyde using choline as...More data for this Ligand-Target Pair
TargetCholine trimethylamine-lyase(Desulfovibrio alaskensis (strain ATCC BAA 1058 / D...)
Harvard University
Curated by ChEMBL
Harvard University
Curated by ChEMBL
Affinity DataIC50: 2.00E+5nMAssay Description:Inhibition of Desulfovibrio desulfuricans (strain G20) CutC assessed as reduction in TMA by spectrophotometryMore data for this Ligand-Target Pair
TargetCholine trimethylamine-lyase(Desulfovibrio alaskensis (strain ATCC BAA 1058 / D...)
Harvard University
Curated by ChEMBL
Harvard University
Curated by ChEMBL
Affinity DataIC50: 2.10E+5nMAssay Description:Inhibition of N-terminal His6-tagged Desulfovibrio desulfuricans G20 CutC encoding 52 residues assessed as reduction in acetaldehyde using choline as...More data for this Ligand-Target Pair
TargetCholine trimethylamine-lyase(Desulfovibrio alaskensis (strain ATCC BAA 1058 / D...)
Harvard University
Curated by ChEMBL
Harvard University
Curated by ChEMBL
Affinity DataIC50: 2.12E+5nMAssay Description:Inhibition of N-terminal His6-tagged Desulfovibrio desulfuricans G20 CutC encoding 52 residues assessed as reduction in acetaldehyde using choline as...More data for this Ligand-Target Pair
TargetCholine trimethylamine-lyase(Desulfovibrio alaskensis (strain ATCC BAA 1058 / D...)
Harvard University
Curated by ChEMBL
Harvard University
Curated by ChEMBL
Affinity DataIC50: 3.01E+5nMAssay Description:Inhibition of N-terminal His6-tagged Desulfovibrio desulfuricans G20 CutC encoding 52 residues assessed as reduction in acetaldehyde using choline as...More data for this Ligand-Target Pair
TargetCholine trimethylamine-lyase(Desulfovibrio alaskensis (strain ATCC BAA 1058 / D...)
Harvard University
Curated by ChEMBL
Harvard University
Curated by ChEMBL
Affinity DataIC50: 4.20E+5nMAssay Description:Inhibition of Desulfovibrio desulfuricans (strain G20) CutC assessed as reduction in TMA by spectrophotometryMore data for this Ligand-Target Pair
TargetCholine trimethylamine-lyase(Desulfovibrio alaskensis (strain ATCC BAA 1058 / D...)
Harvard University
Curated by ChEMBL
Harvard University
Curated by ChEMBL
Affinity DataIC50: 5.00E+5nMAssay Description:Inhibition of N-terminal His6-tagged Desulfovibrio desulfuricans G20 CutC encoding 52 residues assessed as reduction in acetaldehyde using choline as...More data for this Ligand-Target Pair
TargetCholine trimethylamine-lyase(Desulfovibrio alaskensis (strain ATCC BAA 1058 / D...)
Harvard University
Curated by ChEMBL
Harvard University
Curated by ChEMBL
Affinity DataIC50: 5.00E+5nMAssay Description:Inhibition of N-terminal His6-tagged Desulfovibrio desulfuricans G20 CutC encoding 52 residues assessed as reduction in acetaldehyde using choline as...More data for this Ligand-Target Pair
TargetCholine trimethylamine-lyase(Desulfovibrio alaskensis (strain ATCC BAA 1058 / D...)
Harvard University
Curated by ChEMBL
Harvard University
Curated by ChEMBL
Affinity DataIC50: 5.00E+5nMAssay Description:Inhibition of N-terminal His6-tagged Desulfovibrio desulfuricans G20 CutC encoding 52 residues assessed as reduction in acetaldehyde using choline as...More data for this Ligand-Target Pair
TargetCholine trimethylamine-lyase(Desulfovibrio alaskensis (strain ATCC BAA 1058 / D...)
Harvard University
Curated by ChEMBL
Harvard University
Curated by ChEMBL
Affinity DataIC50: 5.00E+5nMAssay Description:Inhibition of N-terminal His6-tagged Desulfovibrio desulfuricans G20 CutC encoding 52 residues assessed as reduction in acetaldehyde using choline as...More data for this Ligand-Target Pair
TargetCholine trimethylamine-lyase(Desulfovibrio alaskensis (strain ATCC BAA 1058 / D...)
Harvard University
Curated by ChEMBL
Harvard University
Curated by ChEMBL
Affinity DataIC50: 5.00E+5nMAssay Description:Inhibition of N-terminal His6-tagged Desulfovibrio desulfuricans G20 CutC encoding 52 residues assessed as reduction in acetaldehyde using choline as...More data for this Ligand-Target Pair
TargetCholine trimethylamine-lyase(Desulfovibrio alaskensis (strain ATCC BAA 1058 / D...)
Harvard University
Curated by ChEMBL
Harvard University
Curated by ChEMBL
Affinity DataIC50: 7.10E+5nMAssay Description:Inhibition of Desulfovibrio desulfuricans (strain G20) CutC assessed as reduction in TMA by spectrophotometryMore data for this Ligand-Target Pair
TargetCholine trimethylamine-lyase(Desulfovibrio alaskensis (strain ATCC BAA 1058 / D...)
Harvard University
Curated by ChEMBL
Harvard University
Curated by ChEMBL
Affinity DataIC50: 8.40E+5nMAssay Description:Inhibition of Desulfovibrio desulfuricans (strain G20) CutC assessed as reduction in TMA by spectrophotometryMore data for this Ligand-Target Pair
TargetCholine trimethylamine-lyase(Desulfovibrio alaskensis (strain ATCC BAA 1058 / D...)
Harvard University
Curated by ChEMBL
Harvard University
Curated by ChEMBL
Affinity DataIC50: 2.00E+6nMAssay Description:Inhibition of Desulfovibrio desulfuricans (strain G20) CutC assessed as reduction in TMA by spectrophotometryMore data for this Ligand-Target Pair
TargetCholine trimethylamine-lyase(Desulfovibrio alaskensis (strain ATCC BAA 1058 / D...)
Harvard University
Curated by ChEMBL
Harvard University
Curated by ChEMBL
Affinity DataIC50: 2.60E+6nMAssay Description:Inhibition of Desulfovibrio desulfuricans (strain G20) CutC assessed as reduction in TMA by spectrophotometryMore data for this Ligand-Target Pair
TargetCholine trimethylamine-lyase(Desulfovibrio alaskensis (strain ATCC BAA 1058 / D...)
Harvard University
Curated by ChEMBL
Harvard University
Curated by ChEMBL
Affinity DataIC50: 2.90E+6nMAssay Description:Inhibition of Desulfovibrio desulfuricans (strain G20) CutC assessed as reduction in TMA by spectrophotometryMore data for this Ligand-Target Pair
TargetCholine trimethylamine-lyase(Desulfovibrio alaskensis (strain ATCC BAA 1058 / D...)
Harvard University
Curated by ChEMBL
Harvard University
Curated by ChEMBL
Affinity DataIC50: 7.00E+6nMAssay Description:Inhibition of Desulfovibrio desulfuricans (strain G20) CutC assessed as reduction in TMA by spectrophotometryMore data for this Ligand-Target Pair
TargetCholine trimethylamine-lyase(Desulfovibrio alaskensis (strain ATCC BAA 1058 / D...)
Harvard University
Curated by ChEMBL
Harvard University
Curated by ChEMBL
Affinity DataIC50: 1.00E+7nMAssay Description:Inhibition of Desulfovibrio desulfuricans (strain G20) CutC assessed as reduction in TMA by spectrophotometryMore data for this Ligand-Target Pair
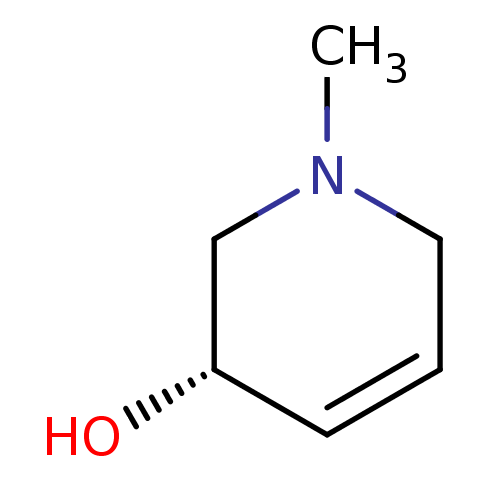
 3D Structure (crystal)
3D Structure (crystal)