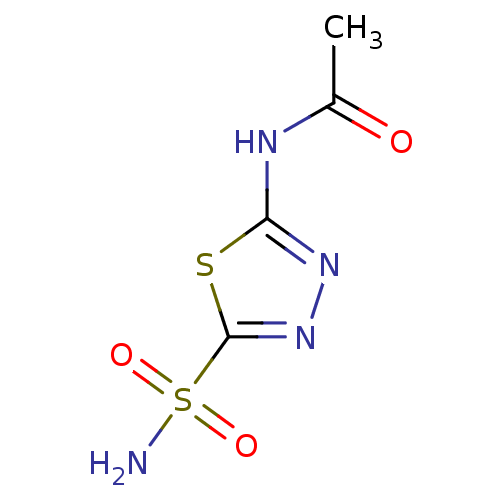
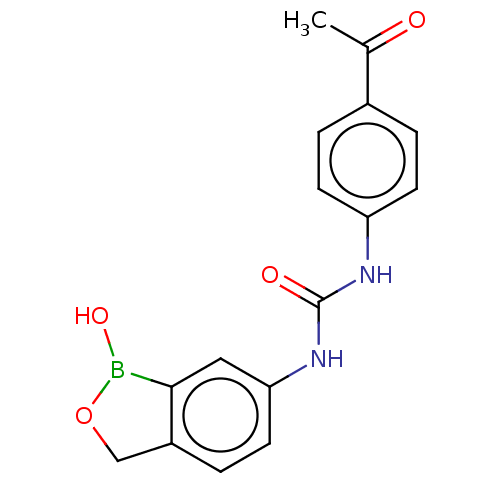
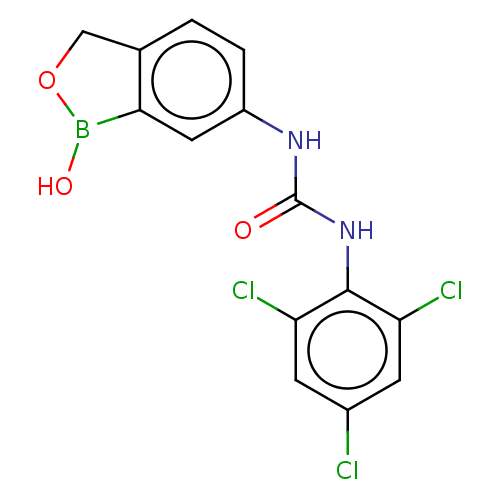
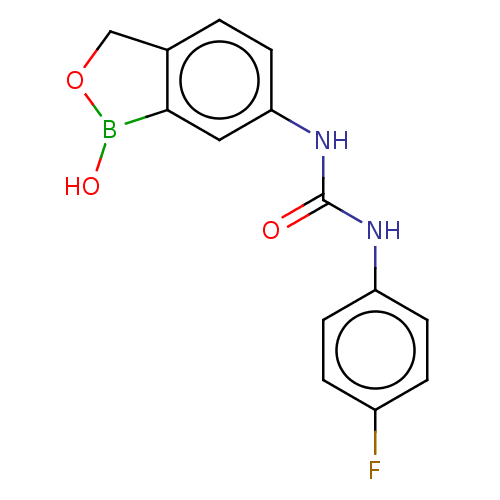
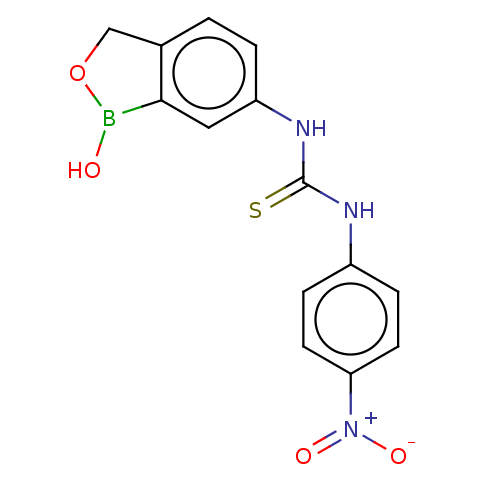
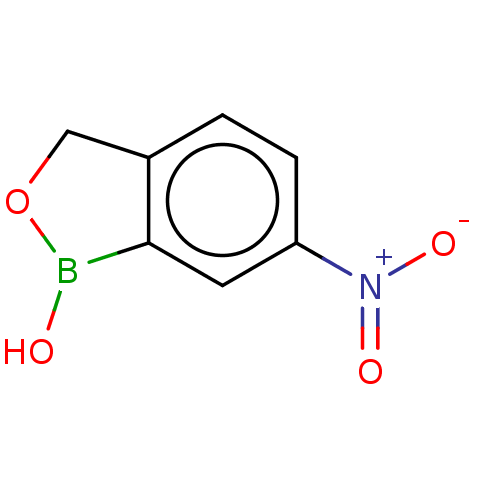
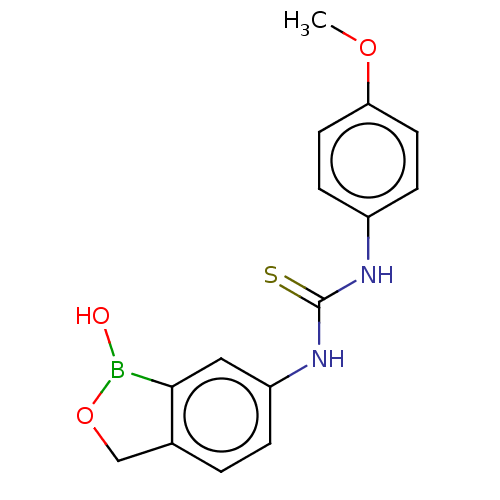
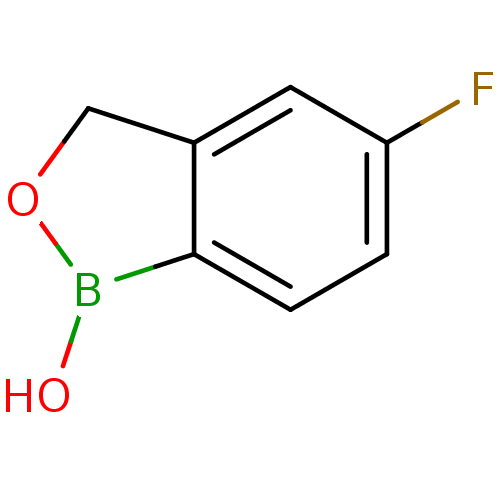
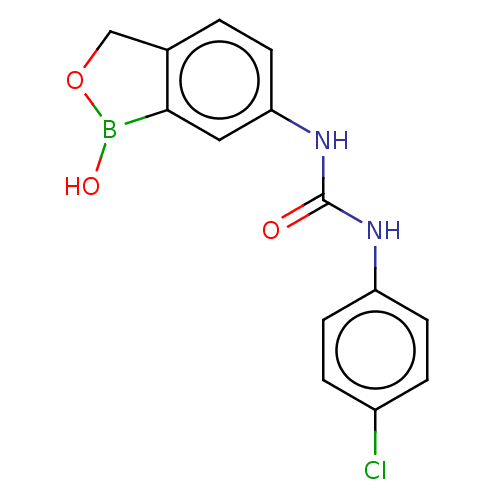
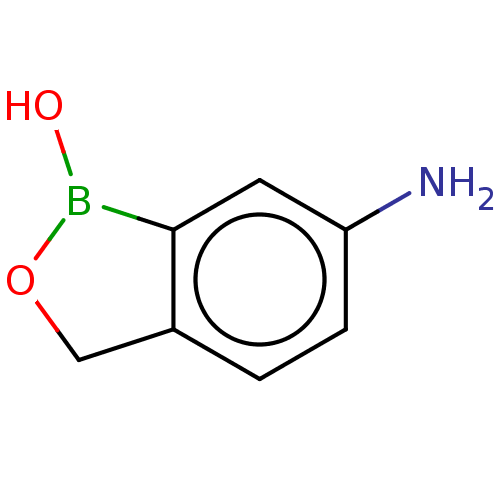
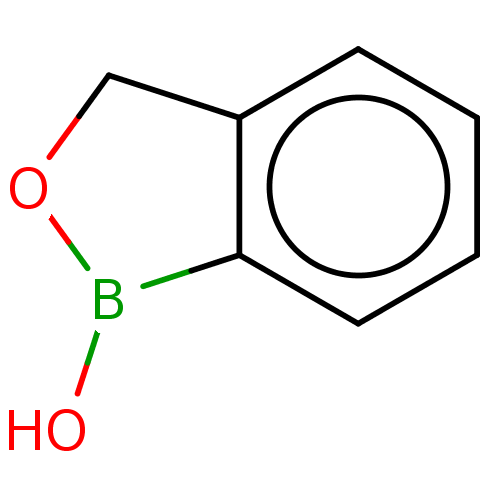
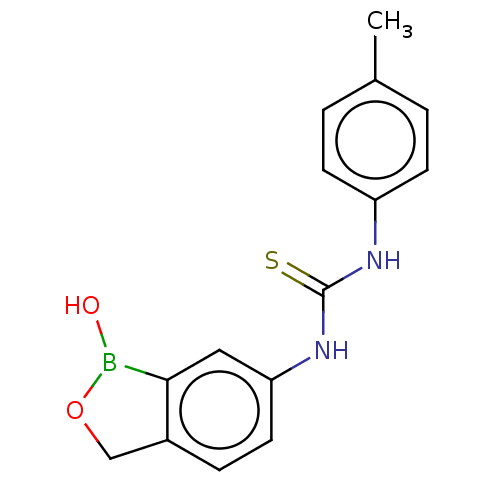
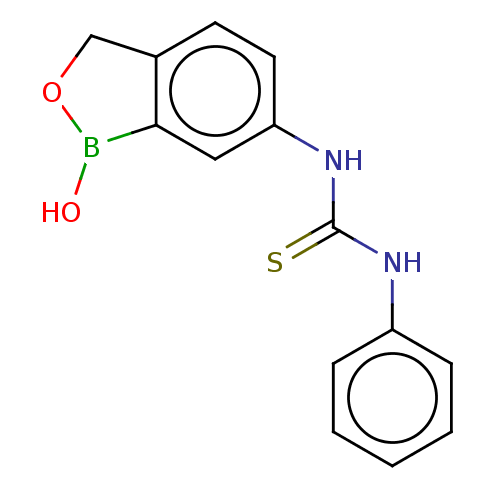
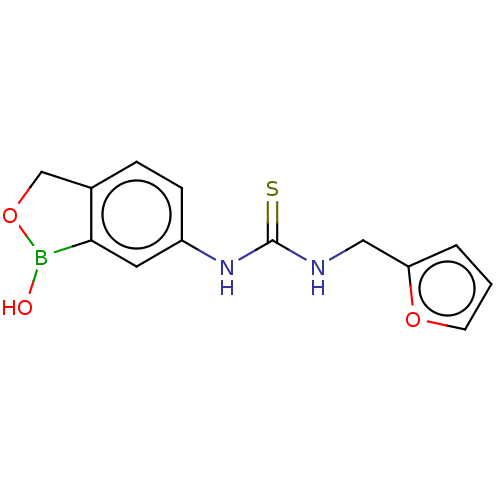
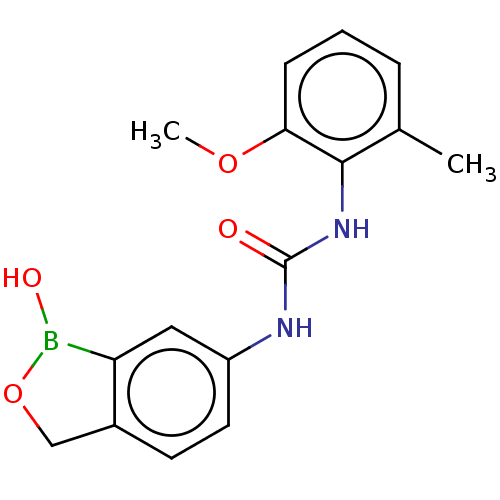
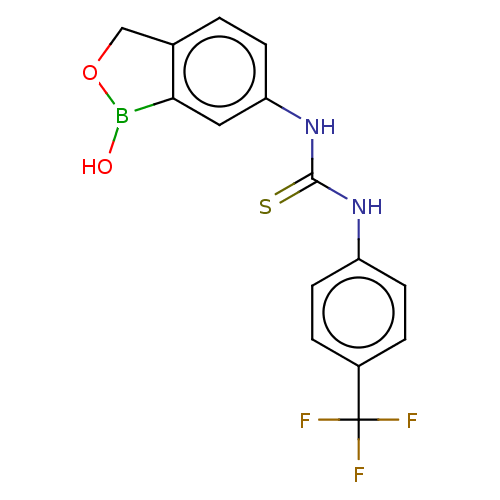
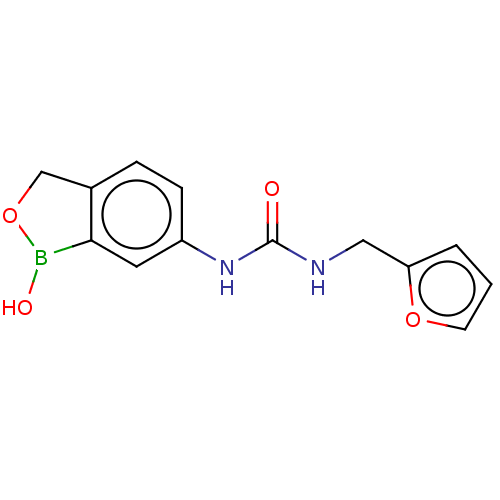
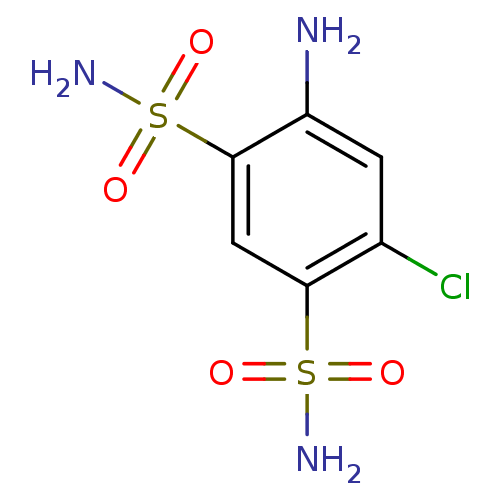
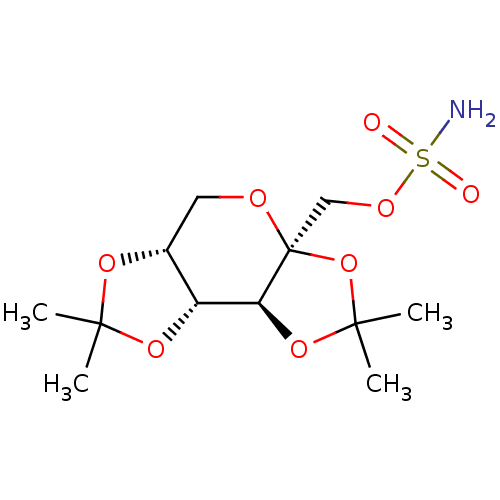
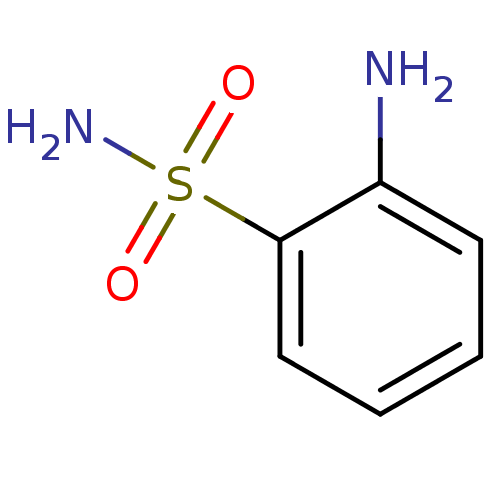
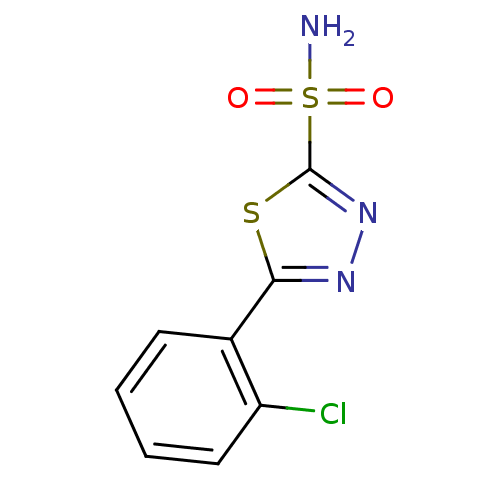
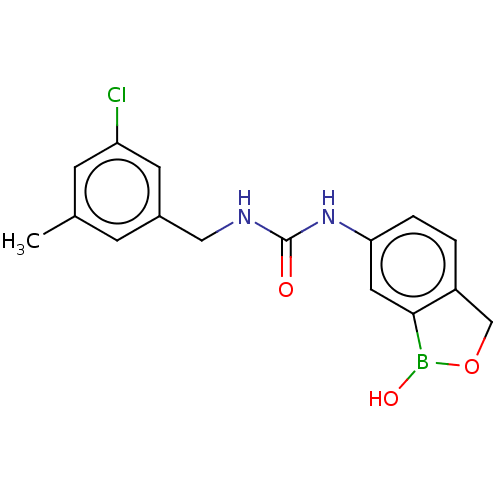
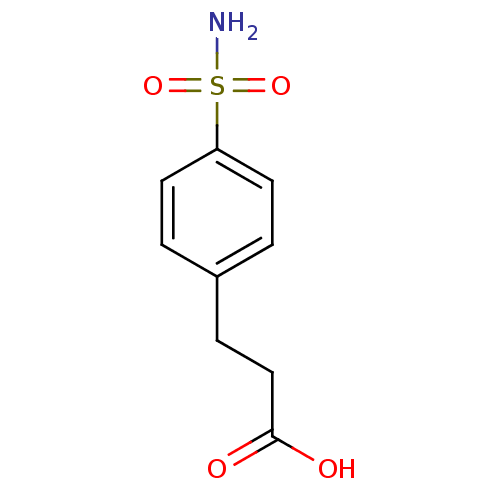
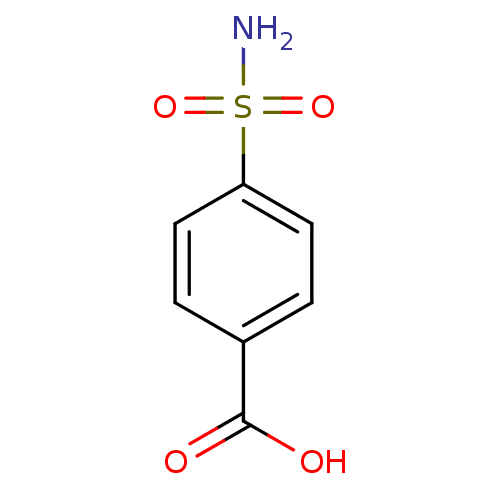
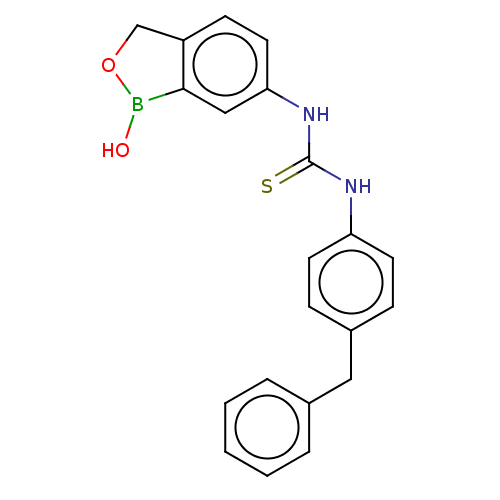
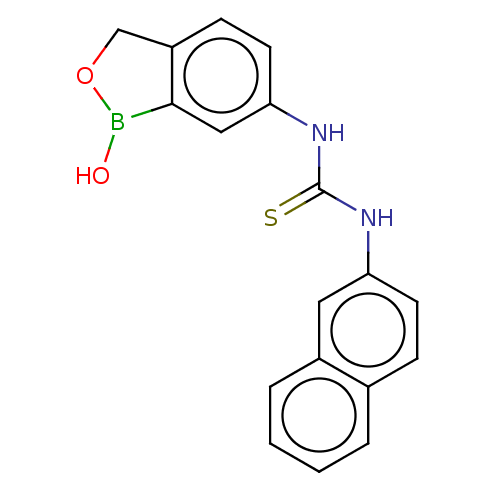
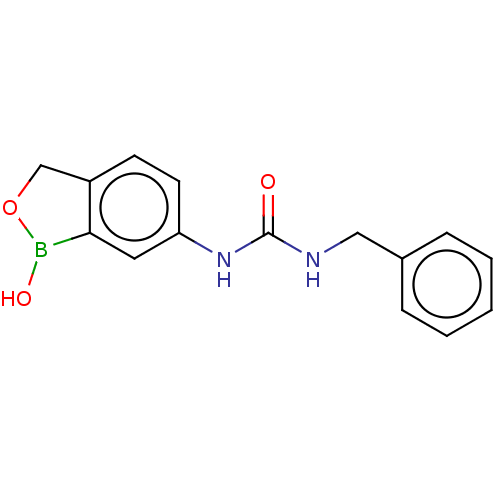
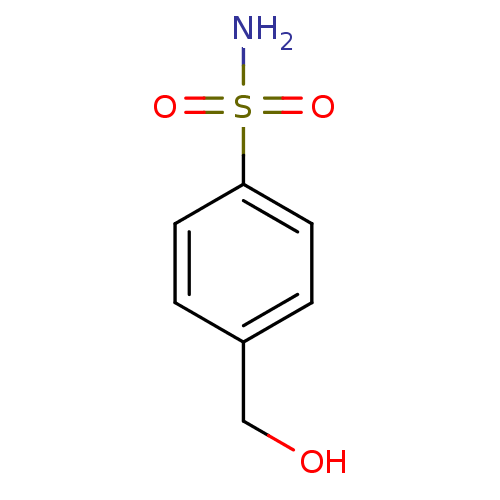
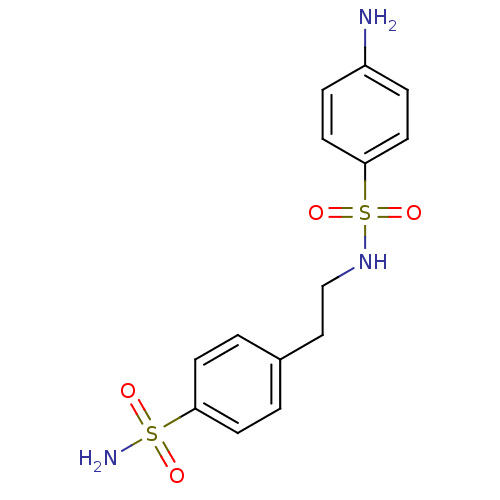
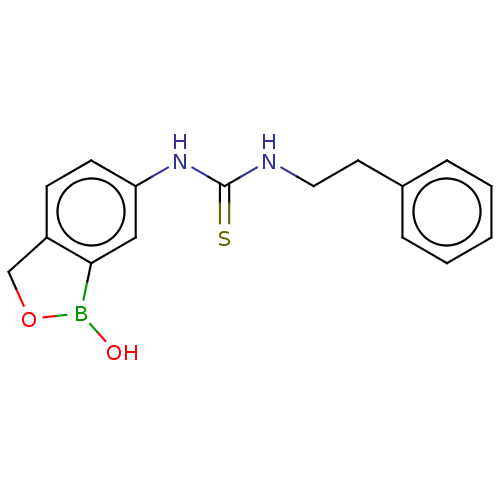
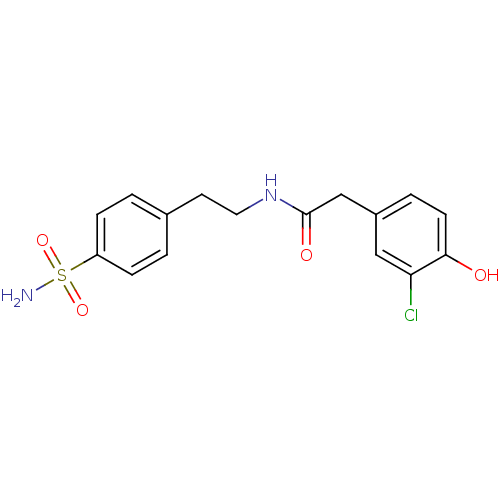
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
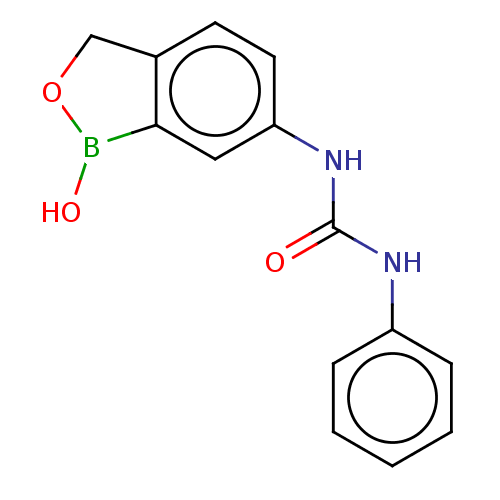
Affinity DataKi: 10nMAssay Description:Inhibition of Cryptococcus neoformans recombinant Can2 beta-carbonic anhydrase after 15 mins by stopped flow CO2 hydration assayMore data for this Ligand-Target Pair
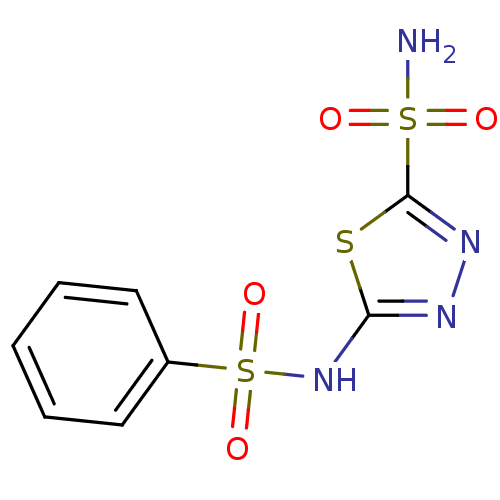
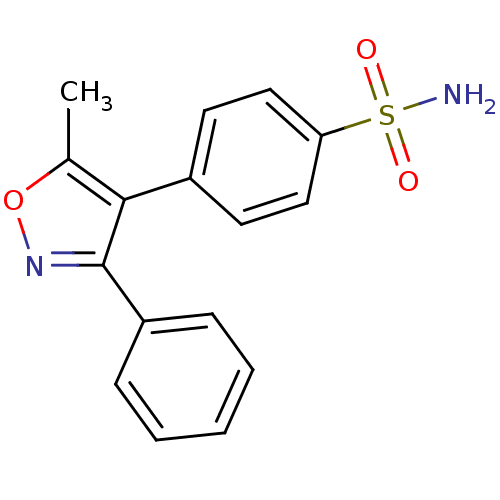
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
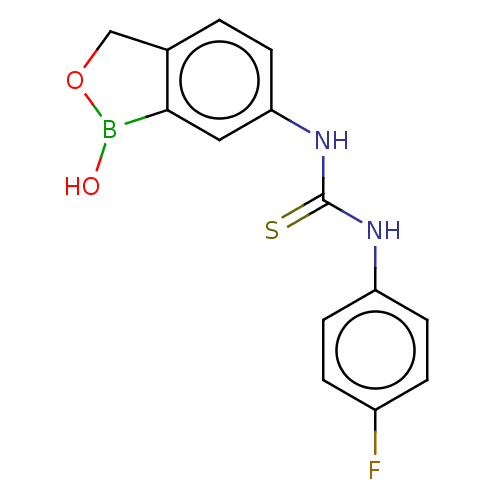
Affinity DataKi: 10nMAssay Description:Inhibition of Cryptococcus neoformans Can2 incubated for 15 mins prior to testing by stopped flow CO2 hydrase assayMore data for this Ligand-Target Pair
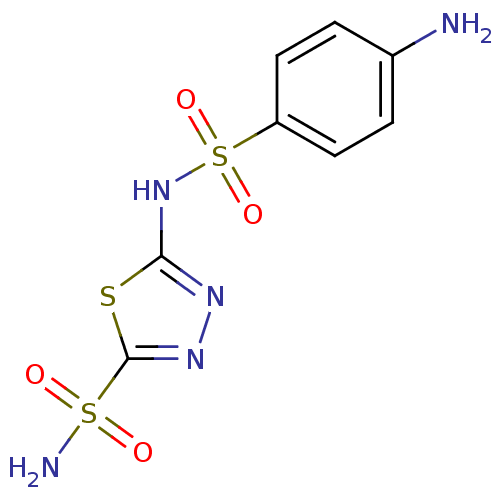
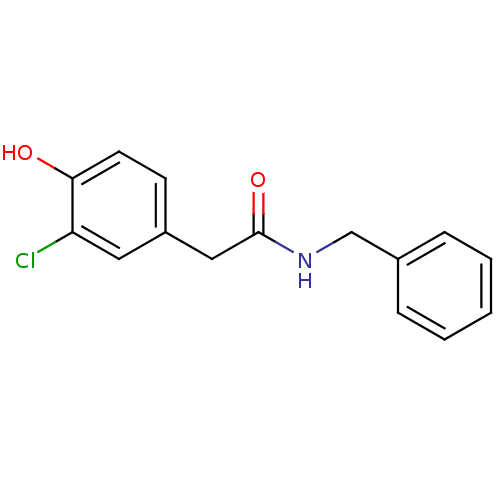
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
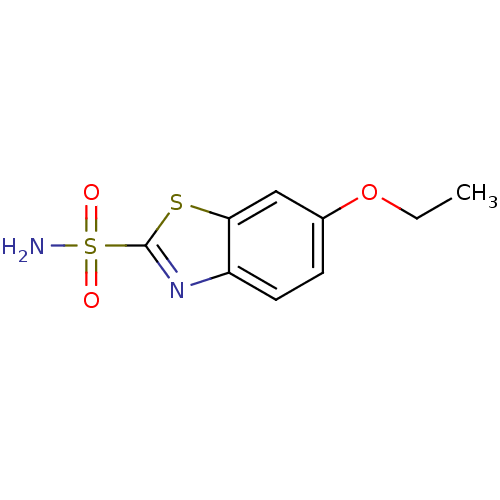
Affinity DataKi: 10.5nMAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
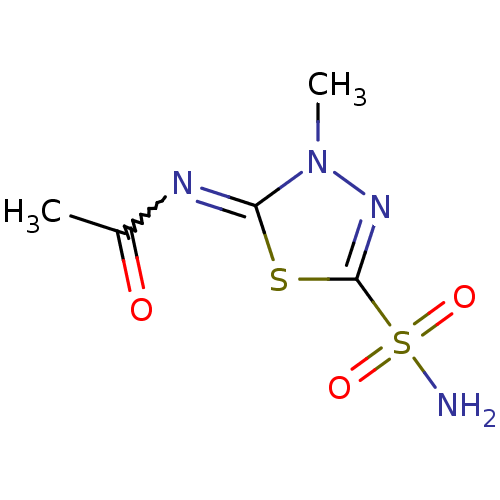
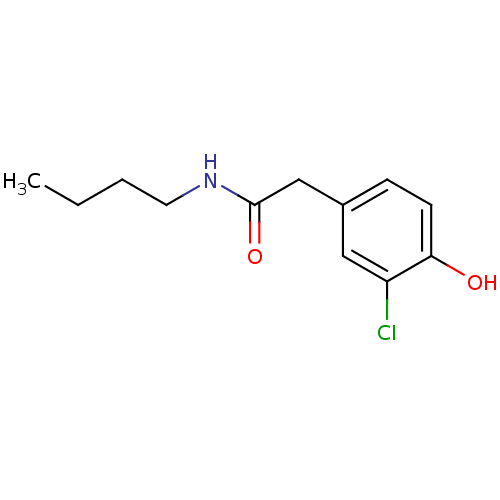
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
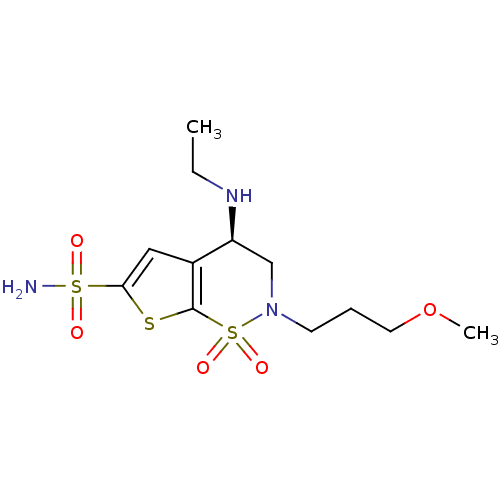
Affinity DataKi: 11nMAssay Description:Inhibition of Cryptococcus neoformans var. grubii beta-carbonic anhydrase incubated for 15 mins prior to testing by stopped-flow CO2 hydration assayMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 11nMAssay Description:Inhibition of Cryptococcus neoformans beta-carbonic anhydrase Can2More data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 23nMAssay Description:Inhibition of Cryptococcus neoformans var. grubii beta-carbonic anhydrase incubated for 15 mins prior to testing by stopped-flow CO2 hydration assayMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 23nMAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 42nMAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 63nMAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 65nMAssay Description:Inhibition of Cryptococcus neoformans Can2 incubated for 15 mins prior to testing by stopped flow CO2 hydrase assayMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 73nMAssay Description:Inhibition of Cryptococcus neoformans Can2 incubated for 15 mins prior to testing by stopped flow CO2 hydrase assayMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 78nMAssay Description:Inhibition of Cryptococcus neoformans Can2 incubated for 15 mins prior to testing by stopped flow CO2 hydrase assayMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 79nMAssay Description:Inhibition of Cryptococcus neoformans Can2 incubated for 15 mins prior to testing by stopped flow CO2 hydrase assayMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 81nMAssay Description:Inhibition of Cryptococcus neoformans Can2 incubated for 15 mins prior to testing by stopped flow CO2 hydrase assayMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 84nMAssay Description:Inhibition of Cryptococcus neoformans Can2 incubated for 15 mins prior to testing by stopped flow CO2 hydrase assayMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 86nMAssay Description:Inhibition of Cryptococcus neoformans Can2 incubated for 15 mins prior to testing by stopped flow CO2 hydrase assayMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 87nMAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 87nMAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 88nMAssay Description:Inhibition of Cryptococcus neoformans Can2 incubated for 15 mins prior to testing by stopped flow CO2 hydrase assayMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 89nMAssay Description:Inhibition of Cryptococcus neoformans Can2 incubated for 15 mins prior to testing by stopped flow CO2 hydrase assayMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 89nMAssay Description:Inhibition of Cryptococcus neoformans Can2 incubated for 15 mins prior to testing by stopped flow CO2 hydrase assayMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 92nMAssay Description:Inhibition of Cryptococcus neoformans Can2 incubated for 15 mins prior to testing by stopped flow CO2 hydrase assayMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 93nMAssay Description:Inhibition of Cryptococcus neoformans Can2 incubated for 15 mins prior to testing by stopped flow CO2 hydrase assayMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 174nMAssay Description:Inhibition of Cryptococcus neoformans Can2 incubated for 15 mins prior to testing by stopped flow CO2 hydrase assayMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 181nMAssay Description:Inhibition of Cryptococcus neoformans Can2 incubated for 15 mins prior to testing by stopped flow CO2 hydrase assayMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 198nMAssay Description:Inhibition of Cryptococcus neoformans Can2 incubated for 15 mins prior to testing by stopped flow CO2 hydrase assayMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 236nMAssay Description:Inhibition of Cryptococcus neoformans Can2 incubated for 15 mins prior to testing by stopped flow CO2 hydrase assayMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 248nMAssay Description:Inhibition of Cryptococcus neoformans Can2 incubated for 15 mins prior to testing by stopped flow CO2 hydrase assayMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 275nMAssay Description:Inhibition of Cryptococcus neoformans Can2 incubated for 15 mins prior to testing by stopped flow CO2 hydrase assayMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 283nMAssay Description:Inhibition of Cryptococcus neoformans Can2 incubated for 15 mins prior to testing by stopped flow CO2 hydrase assayMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 300nMAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 367nMAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 370nMAssay Description:Inhibition of Cryptococcus neoformans recombinant Can2 beta-carbonic anhydrase after 15 mins by stopped flow CO2 hydration assayMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 379nMAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 379nMAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 400nMAssay Description:Inhibition of Cryptococcus neoformans Can2 incubated for 15 mins prior to testing by stopped flow CO2 hydrase assayMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 440nMAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 484nMAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 537nMAssay Description:Inhibition of Cryptococcus neoformans Can2 incubated for 15 mins prior to testing by stopped flow CO2 hydrase assayMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 573nMAssay Description:Inhibition of Cryptococcus neoformans Can2 incubated for 15 mins prior to testing by stopped flow CO2 hydrase assayMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 602nMAssay Description:Inhibition of Cryptococcus neoformans Can2 incubated for 15 mins prior to testing by stopped flow CO2 hydrase assayMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 605nMAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 623nMAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 624nMAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 676nMAssay Description:Inhibition of Cryptococcus neoformans Can2 incubated for 15 mins prior to testing by stopped flow CO2 hydrase assayMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 704nMAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 711nMAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA-catalyzed CO2 hydration activity. Phenol red has been used as indic...More data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 720nMAssay Description:Inhibition of Cryptococcus neoformans recombinant Can2 beta-carbonic anhydrase after 15 mins by stopped flow CO2 hydration assayMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 730nMAssay Description:Inhibition of Cryptococcus neoformans recombinant Can2 beta-carbonic anhydrase after 15 mins by stopped flow CO2 hydration assayMore data for this Ligand-Target Pair
TargetCarbonic anhydrase(Cryptococcus neoformans var. grubii (Filobasidiell...)
Griffith University
Curated by ChEMBL
Griffith University
Curated by ChEMBL
Affinity DataKi: 740nMAssay Description:Inhibition of Cryptococcus neoformans recombinant Can2 beta-carbonic anhydrase after 15 mins by stopped flow CO2 hydration assayMore data for this Ligand-Target Pair