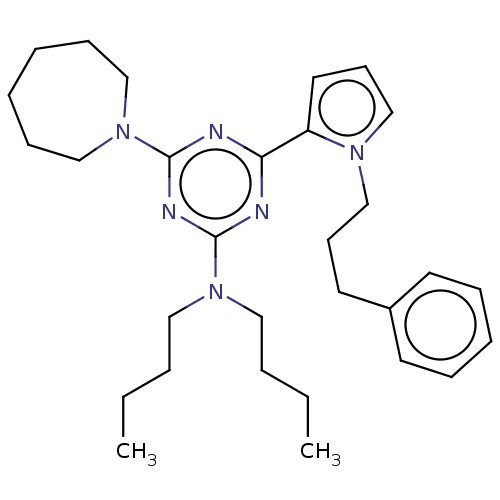
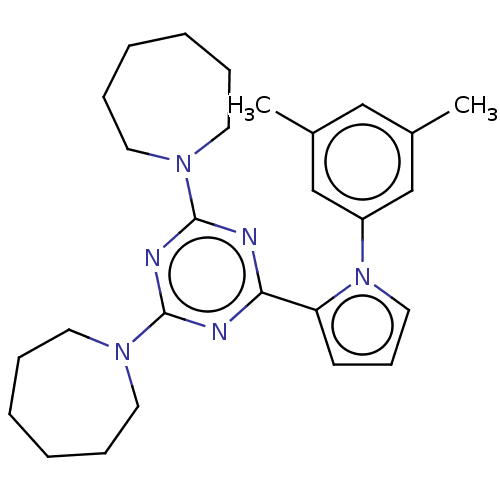
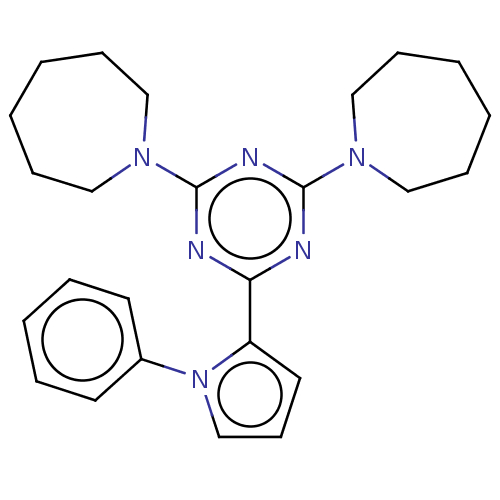
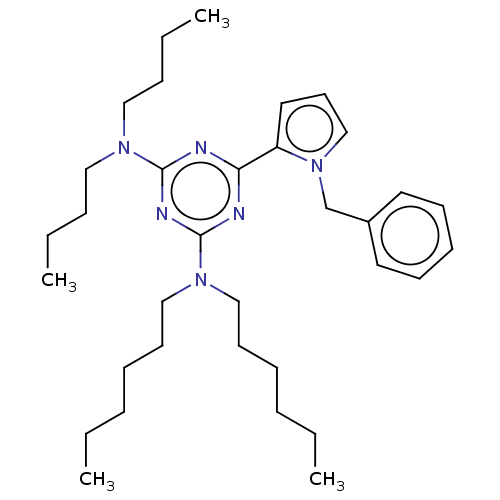
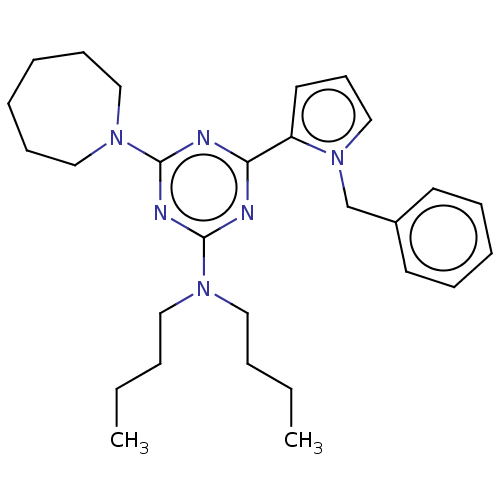
TargetProbable manganese-dependent inorganic pyrophosphatase(Staphylococcus aureus (strain MRSA252))
University of Kentucky
University of Kentucky
Affinity DataIC50: 9.70E+4nMpH: 7.5 T: 2°CAssay Description:The compounds were first mixed with tetrasodium PPi (final concentration of 100 μM) at concentrations specified in the text in the reaction buff...More data for this Ligand-Target Pair
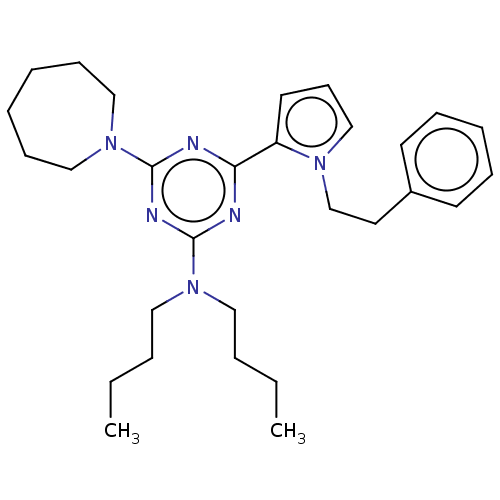
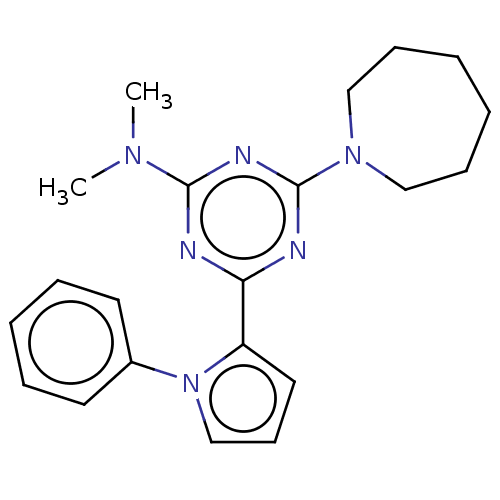
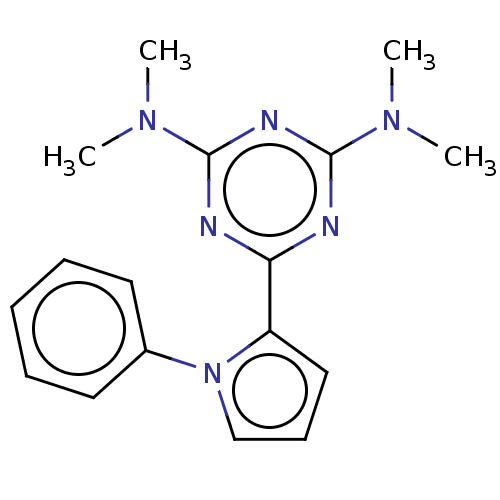
TargetProbable manganese-dependent inorganic pyrophosphatase(Staphylococcus aureus (strain MRSA252))
University of Kentucky
University of Kentucky
Affinity DataIC50: 1.09E+5nMpH: 7.5 T: 2°CAssay Description:The compounds were first mixed with tetrasodium PPi (final concentration of 100 μM) at concentrations specified in the text in the reaction buff...More data for this Ligand-Target Pair
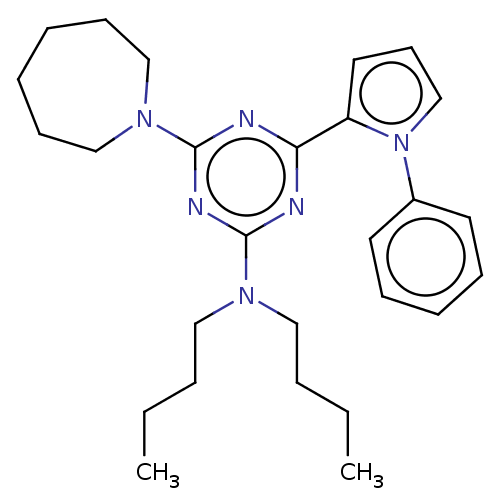
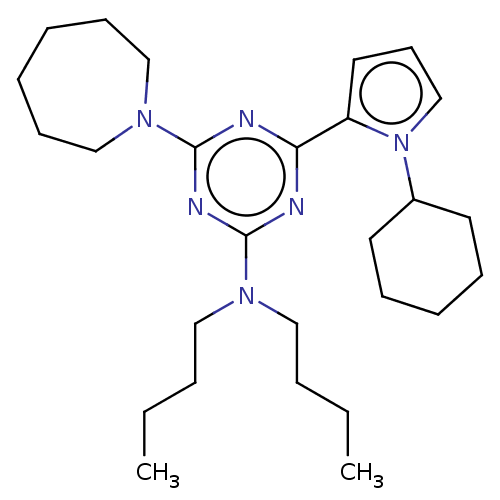
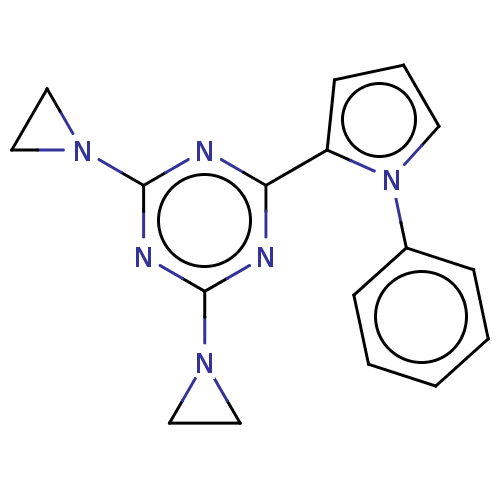
TargetProbable manganese-dependent inorganic pyrophosphatase(Staphylococcus aureus (strain MRSA252))
University of Kentucky
University of Kentucky
Affinity DataIC50: 1.55E+5nMpH: 7.5 T: 2°CAssay Description:The compounds were first mixed with tetrasodium PPi (final concentration of 100 μM) at concentrations specified in the text in the reaction buff...More data for this Ligand-Target Pair
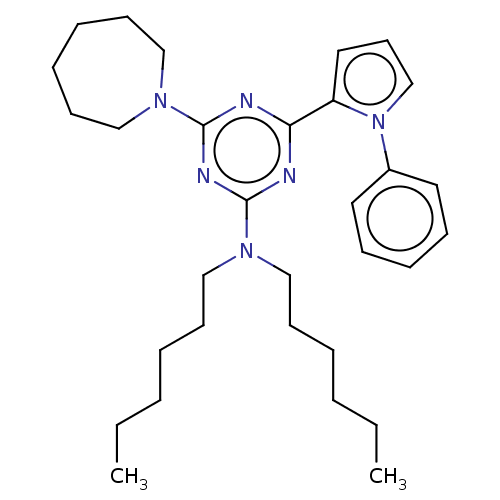
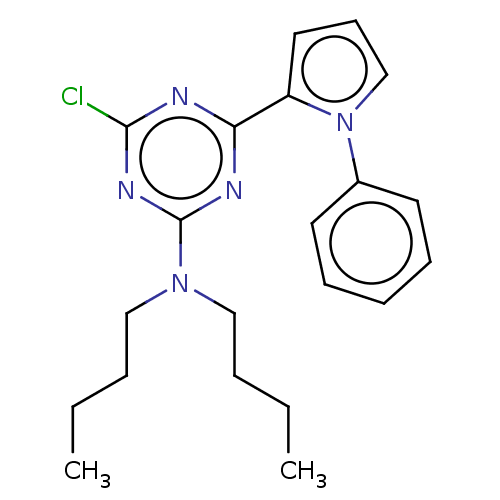
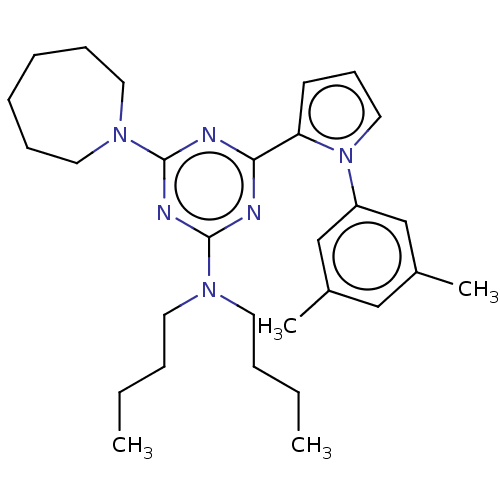
TargetProbable manganese-dependent inorganic pyrophosphatase(Staphylococcus aureus (strain MRSA252))
University of Kentucky
University of Kentucky
Affinity DataIC50: 2.69E+5nMpH: 7.5 T: 2°CAssay Description:The compounds were first mixed with tetrasodium PPi (final concentration of 100 μM) at concentrations specified in the text in the reaction buff...More data for this Ligand-Target Pair
TargetProbable manganese-dependent inorganic pyrophosphatase(Staphylococcus aureus (strain MRSA252))
University of Kentucky
University of Kentucky
Affinity DataIC50: 3.00E+5nMpH: 7.5 T: 2°CAssay Description:The compounds were first mixed with tetrasodium PPi (final concentration of 100 μM) at concentrations specified in the text in the reaction buff...More data for this Ligand-Target Pair
TargetProbable manganese-dependent inorganic pyrophosphatase(Staphylococcus aureus (strain MRSA252))
University of Kentucky
University of Kentucky
Affinity DataIC50: 3.14E+5nMpH: 7.5 T: 2°CAssay Description:The compounds were first mixed with tetrasodium PPi (final concentration of 100 μM) at concentrations specified in the text in the reaction buff...More data for this Ligand-Target Pair
TargetProbable manganese-dependent inorganic pyrophosphatase(Staphylococcus aureus (strain MRSA252))
University of Kentucky
University of Kentucky
Affinity DataIC50: 3.60E+5nMpH: 7.5 T: 2°CAssay Description:The compounds were first mixed with tetrasodium PPi (final concentration of 100 μM) at concentrations specified in the text in the reaction buff...More data for this Ligand-Target Pair
TargetProbable manganese-dependent inorganic pyrophosphatase(Staphylococcus aureus (strain MRSA252))
University of Kentucky
University of Kentucky
Affinity DataIC50: 6.00E+5nMpH: 7.5 T: 2°CAssay Description:The compounds were first mixed with tetrasodium PPi (final concentration of 100 μM) at concentrations specified in the text in the reaction buff...More data for this Ligand-Target Pair
TargetProbable manganese-dependent inorganic pyrophosphatase(Staphylococcus aureus (strain MRSA252))
University of Kentucky
University of Kentucky
Affinity DataIC50: 6.00E+5nMpH: 7.5 T: 2°CAssay Description:The compounds were first mixed with tetrasodium PPi (final concentration of 100 μM) at concentrations specified in the text in the reaction buff...More data for this Ligand-Target Pair
TargetProbable manganese-dependent inorganic pyrophosphatase(Staphylococcus aureus (strain MRSA252))
University of Kentucky
University of Kentucky
Affinity DataIC50: 6.00E+5nMpH: 7.5 T: 2°CAssay Description:The compounds were first mixed with tetrasodium PPi (final concentration of 100 μM) at concentrations specified in the text in the reaction buff...More data for this Ligand-Target Pair
TargetProbable manganese-dependent inorganic pyrophosphatase(Staphylococcus aureus (strain MRSA252))
University of Kentucky
University of Kentucky
Affinity DataIC50: 6.00E+5nMpH: 7.5 T: 2°CAssay Description:The compounds were first mixed with tetrasodium PPi (final concentration of 100 μM) at concentrations specified in the text in the reaction buff...More data for this Ligand-Target Pair
TargetProbable manganese-dependent inorganic pyrophosphatase(Staphylococcus aureus (strain MRSA252))
University of Kentucky
University of Kentucky
Affinity DataIC50: 6.00E+5nMpH: 7.5 T: 2°CAssay Description:The compounds were first mixed with tetrasodium PPi (final concentration of 100 μM) at concentrations specified in the text in the reaction buff...More data for this Ligand-Target Pair
TargetProbable manganese-dependent inorganic pyrophosphatase(Staphylococcus aureus (strain MRSA252))
University of Kentucky
University of Kentucky
Affinity DataIC50: 6.00E+5nMpH: 7.5 T: 2°CAssay Description:The compounds were first mixed with tetrasodium PPi (final concentration of 100 μM) at concentrations specified in the text in the reaction buff...More data for this Ligand-Target Pair
TargetProbable manganese-dependent inorganic pyrophosphatase(Staphylococcus aureus (strain MRSA252))
University of Kentucky
University of Kentucky
Affinity DataIC50: 6.00E+5nMpH: 7.5 T: 2°CAssay Description:The compounds were first mixed with tetrasodium PPi (final concentration of 100 μM) at concentrations specified in the text in the reaction buff...More data for this Ligand-Target Pair