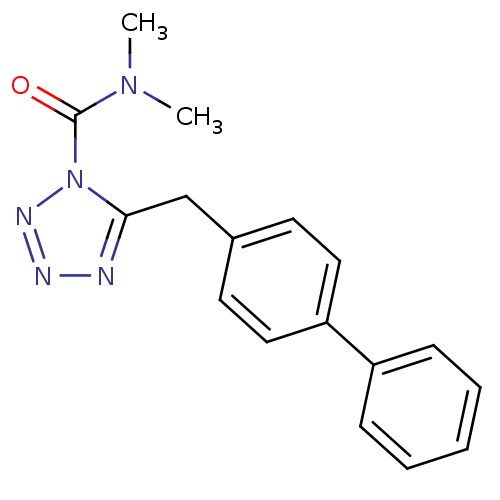
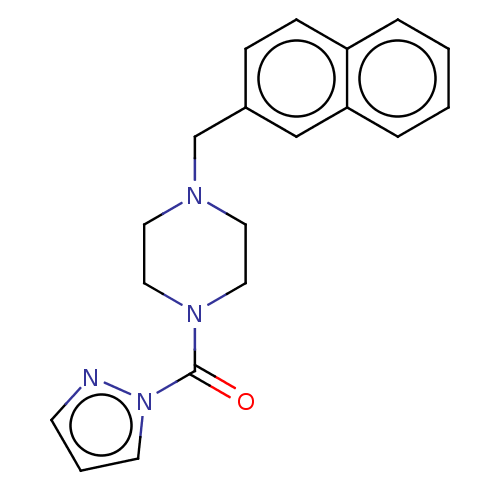
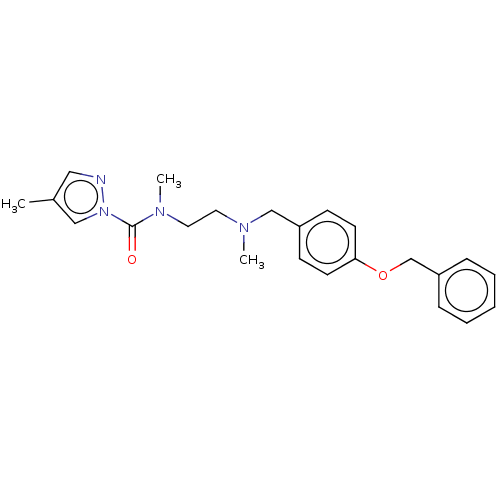
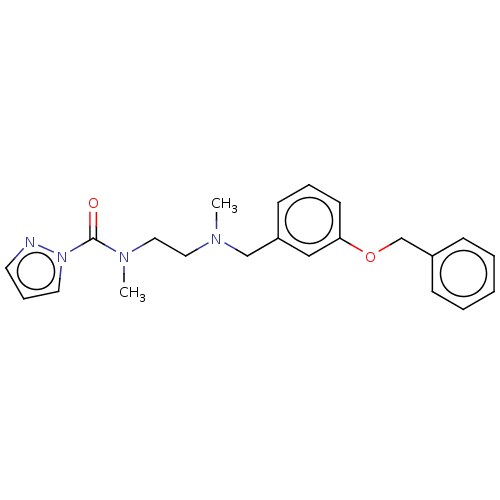
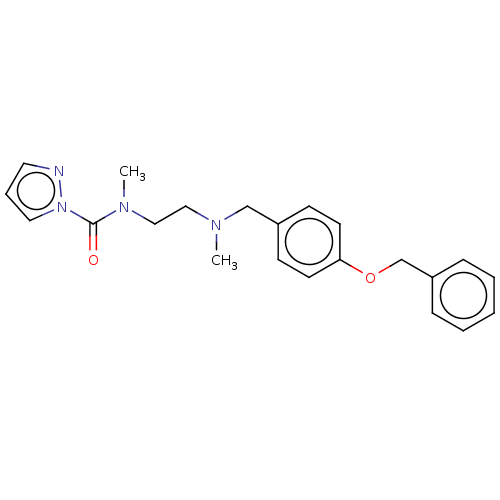
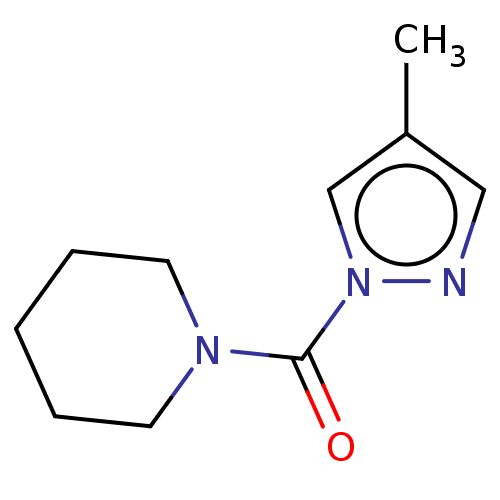
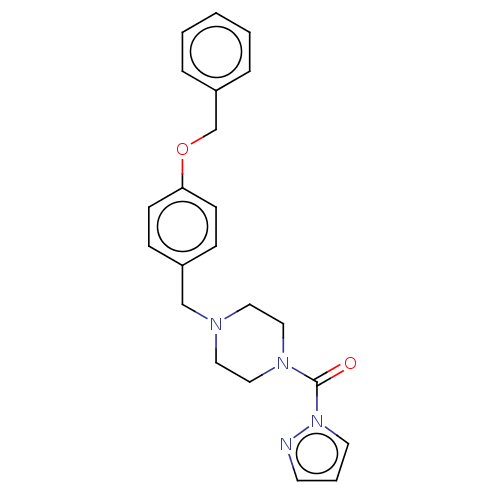
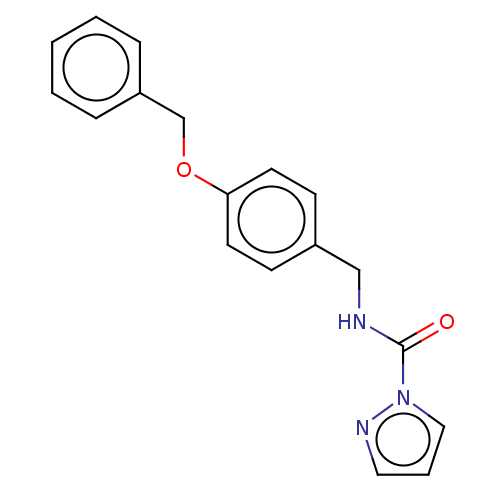
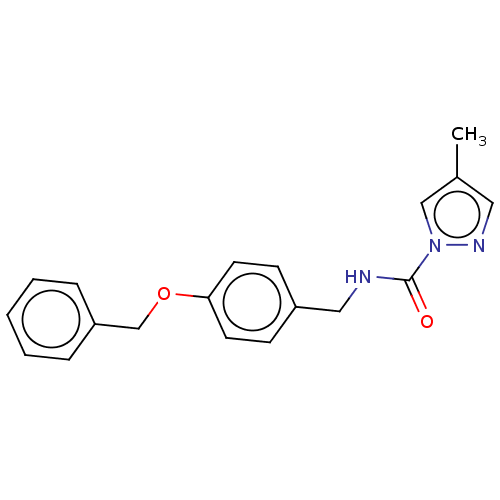
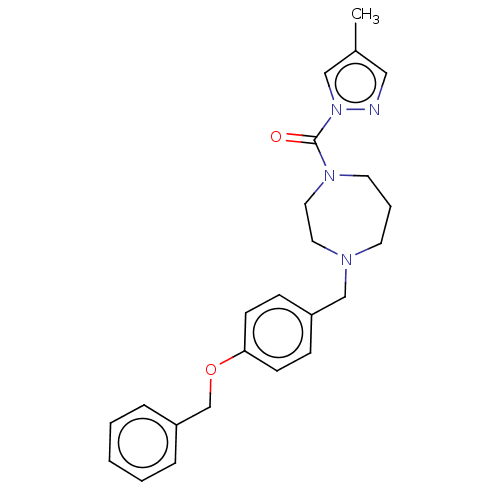
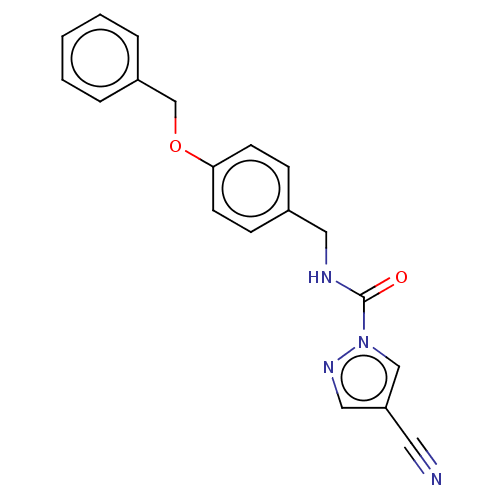
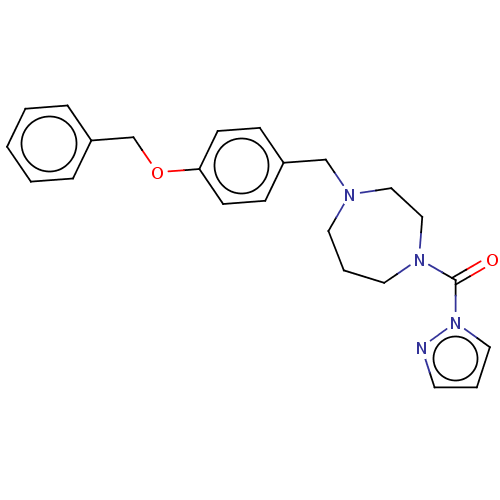
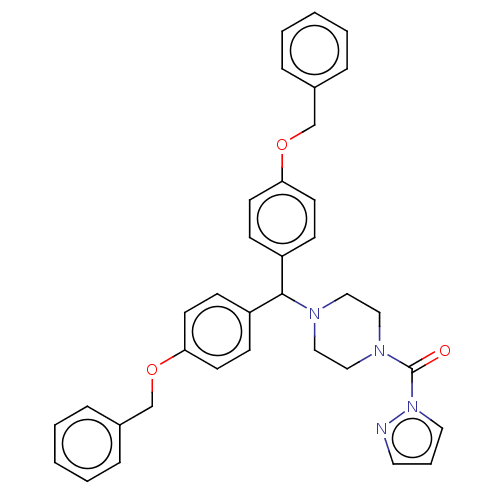
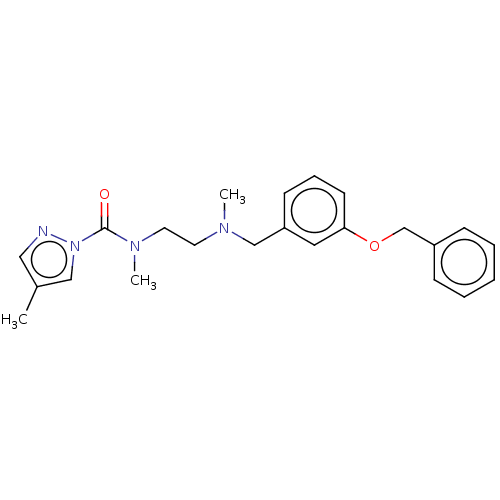
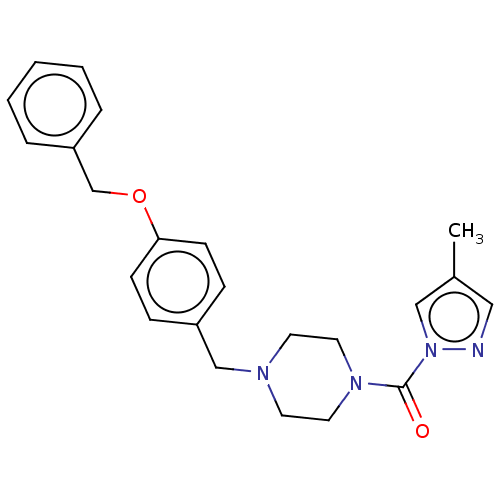
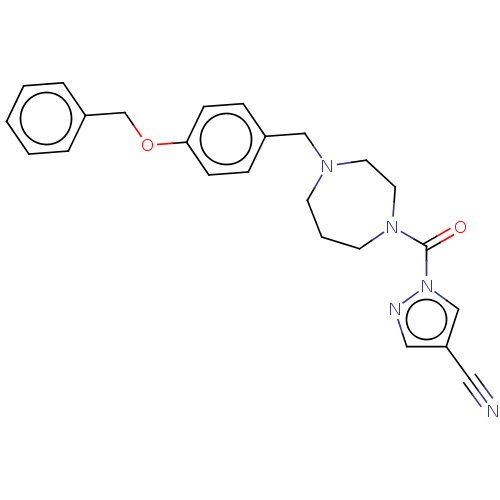
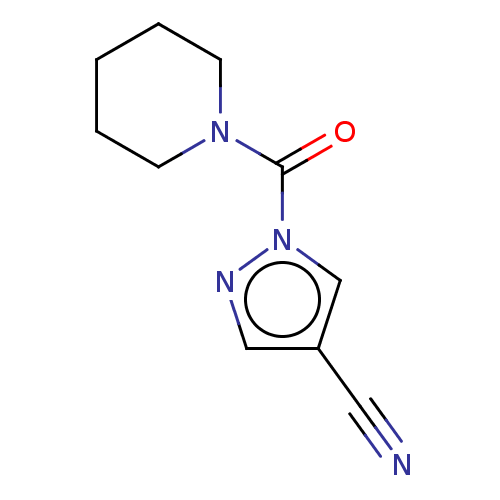
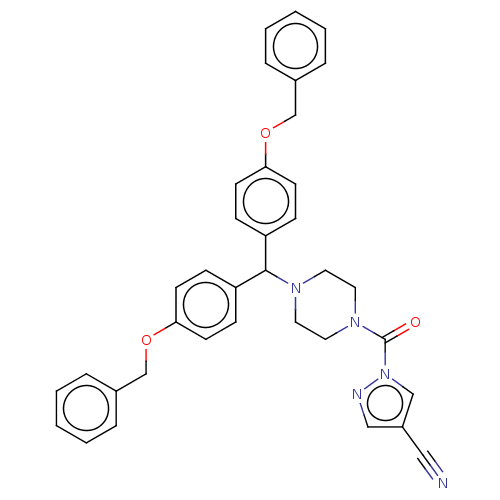
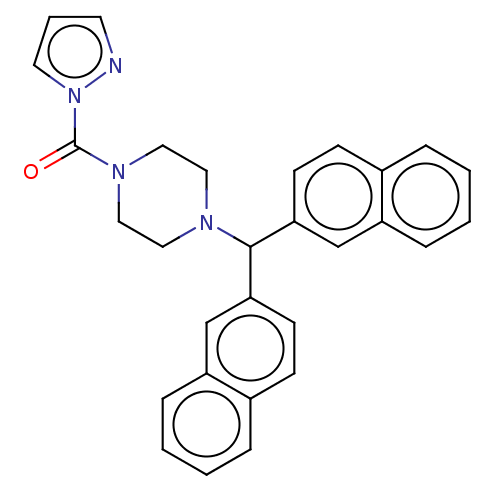
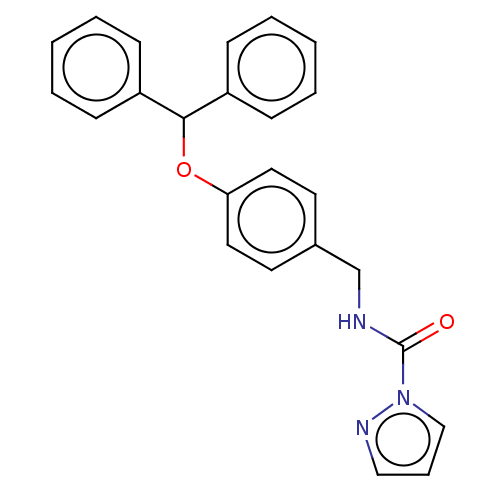
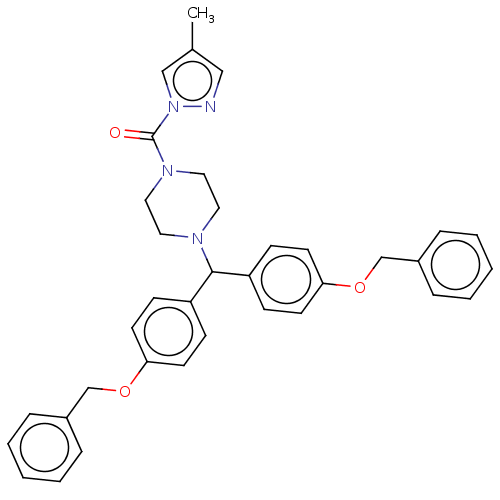
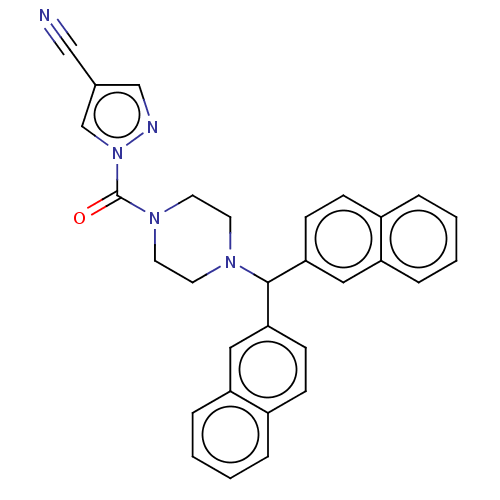
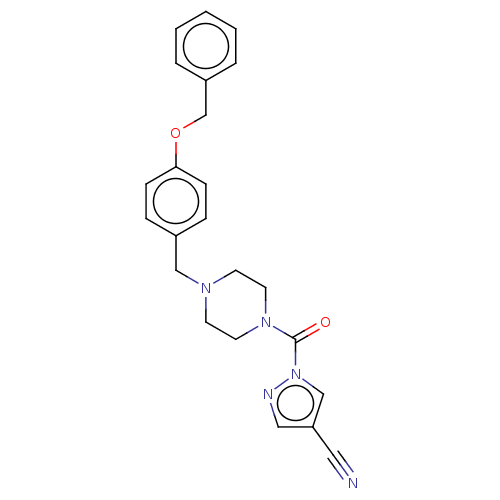
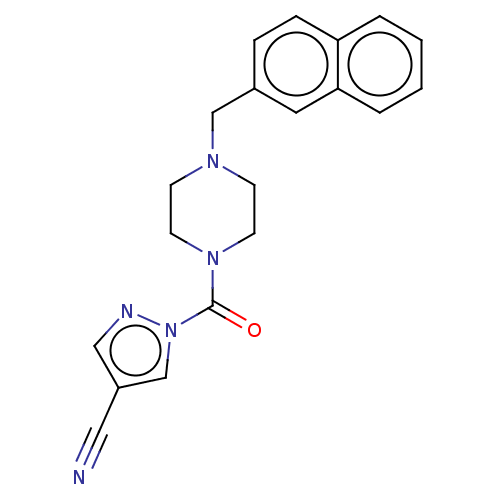
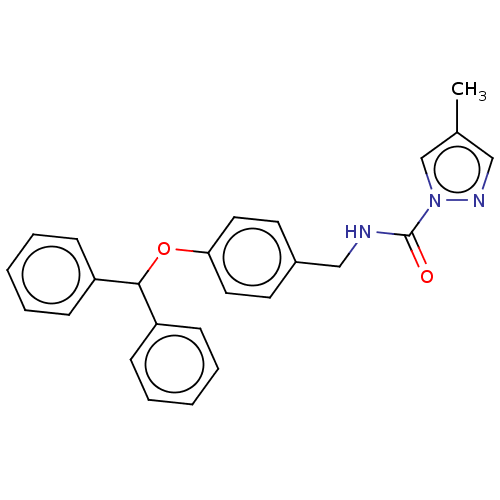
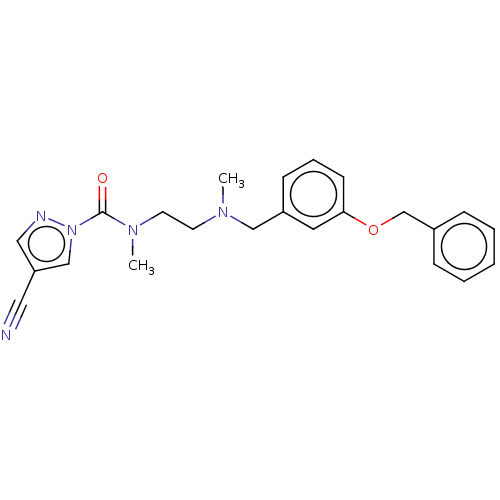
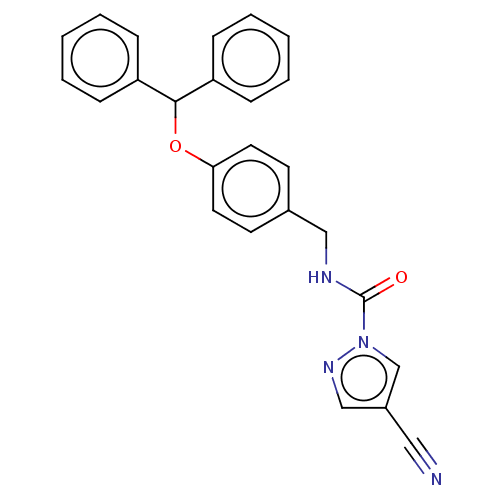
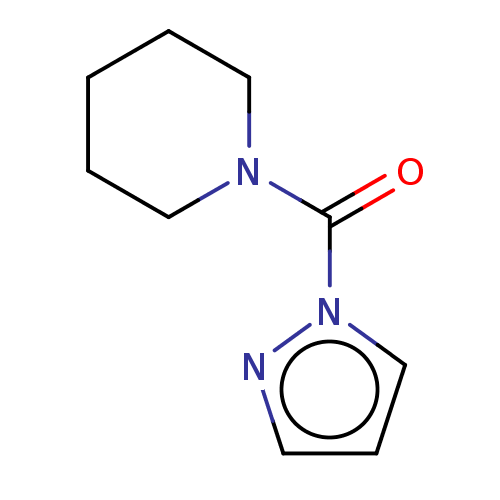
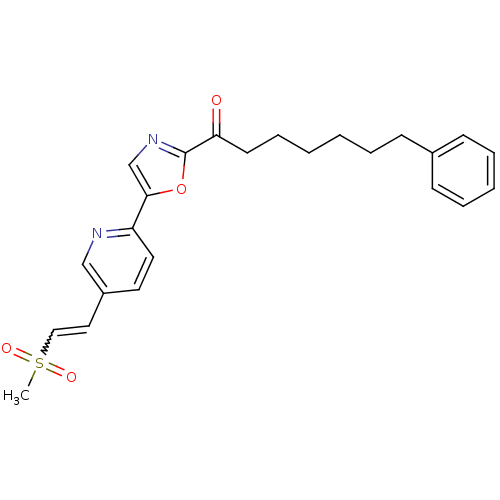
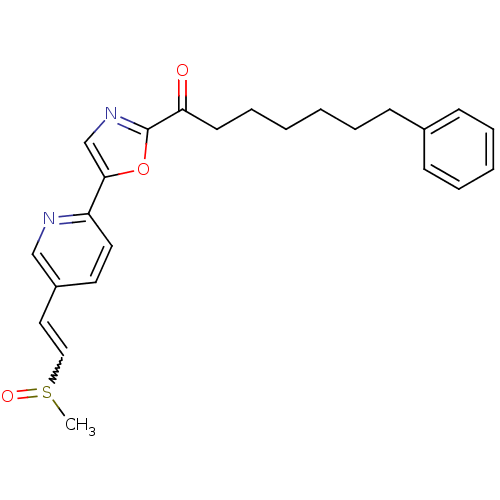
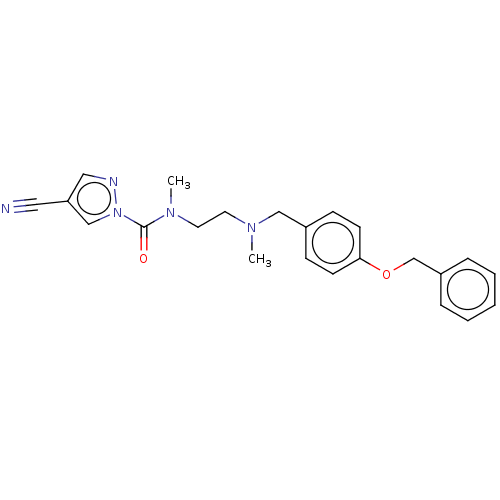
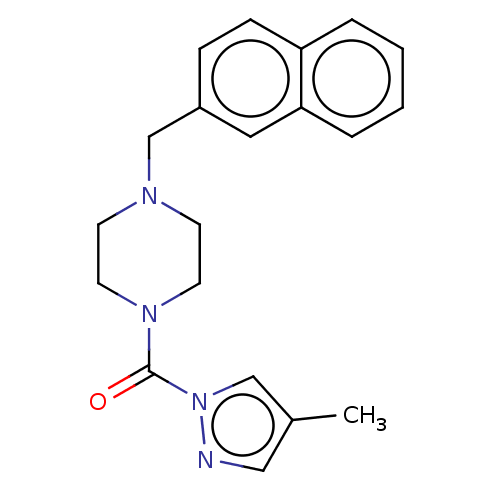
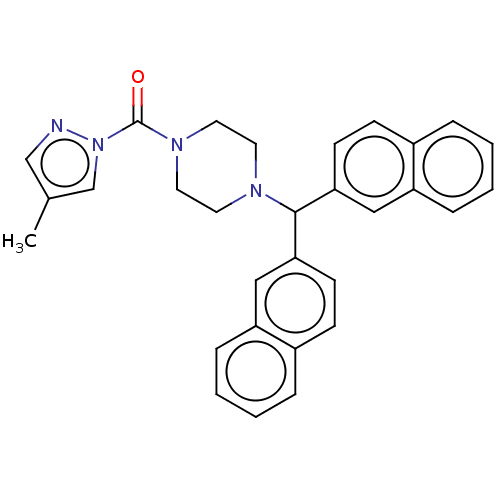
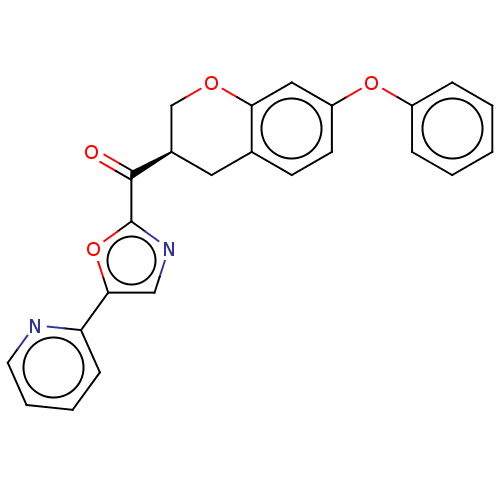
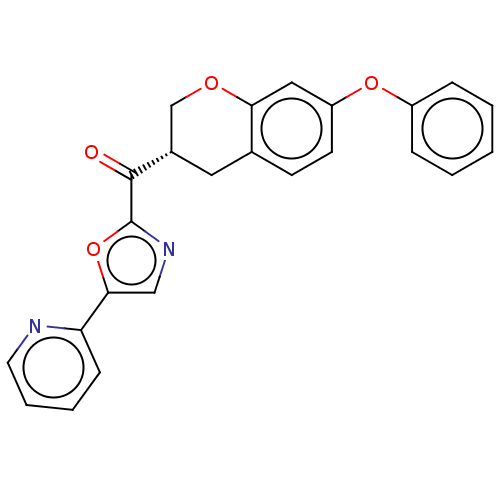
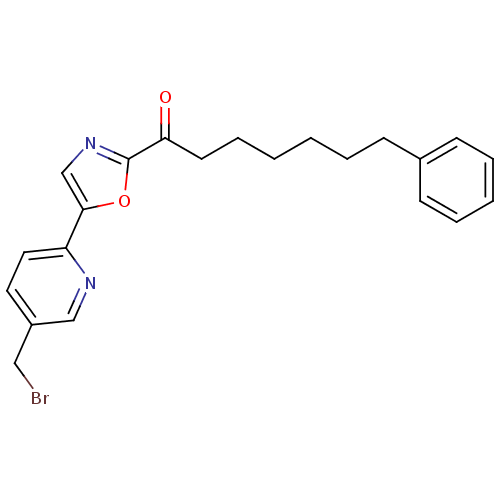
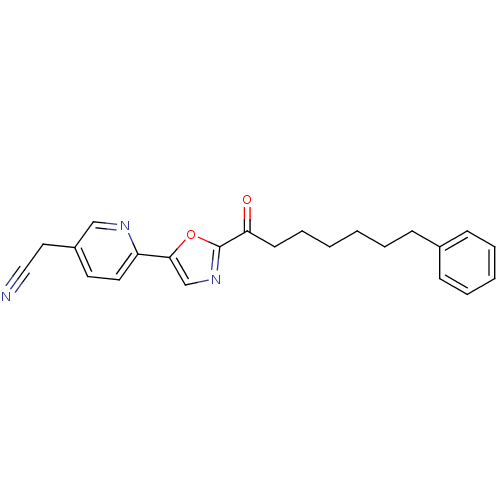
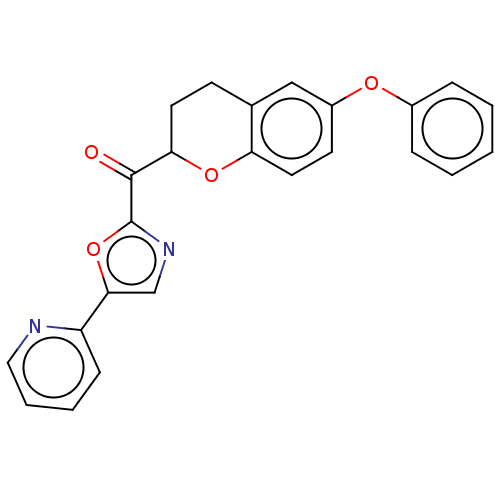
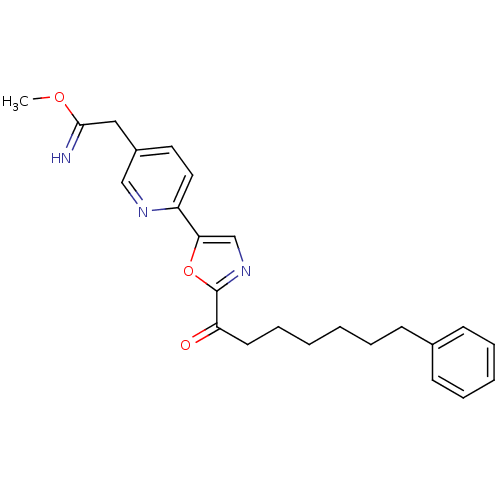
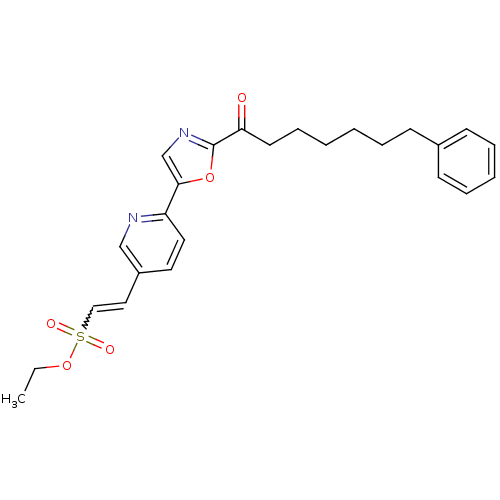
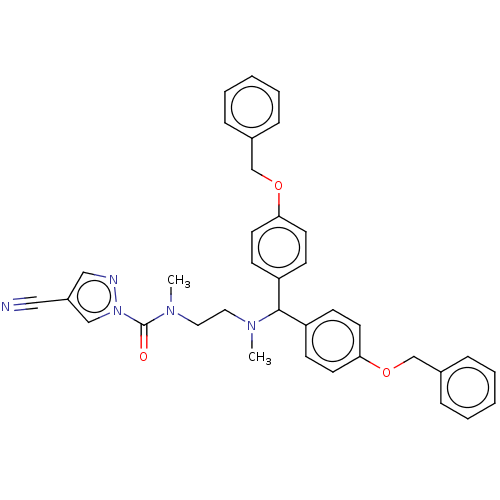
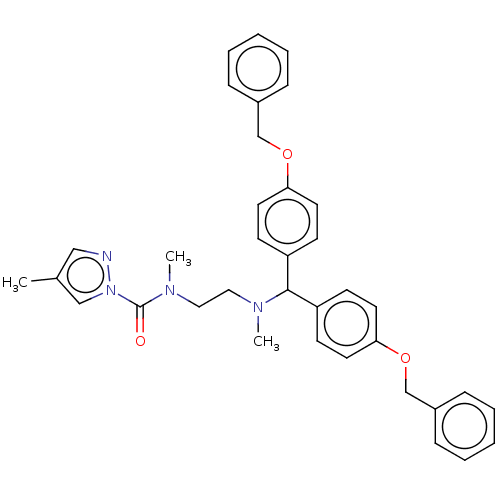
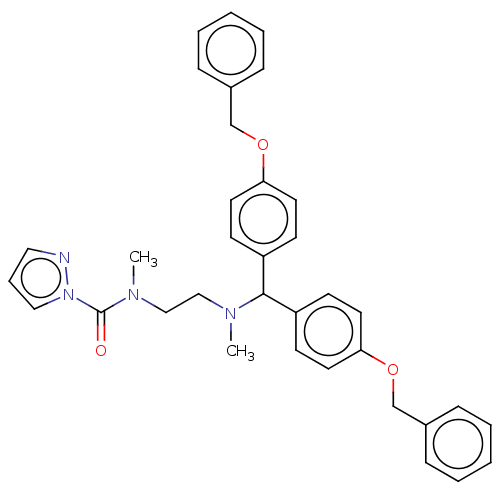
Affinity DataIC50: 8.20nMpH: 7.4 T: 2°CAssay Description:Serine hydrolase targets were recombinantly expressed in COS-7 cells by transient transfection. IC50 values were obtained by competitive ABPP with F...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of FP-rhodamine probe binding to KIAA1363 in mouse brain membrane proteome preincubated for 1 hr followed by FP-rhodamine addition and mea...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of FP-rhodamine probe binding to KIAA1363 in mouse brain membrane proteome preincubated for 1 hr followed by FP-rhodamine addition and mea...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of FP-rhodamine probe binding to KIAA1363 in mouse brain membrane proteome preincubated for 1 hr followed by FP-rhodamine addition and mea...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of FP-rhodamine probe binding to KIAA1363 in mouse brain membrane proteome preincubated for 1 hr followed by FP-rhodamine addition and mea...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of FP-rhodamine probe binding to KIAA1363 in mouse brain membrane proteome preincubated for 1 hr followed by FP-rhodamine addition and mea...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of FP-rhodamine probe binding to KIAA1363 in mouse brain membrane proteome preincubated for 1 hr followed by FP-rhodamine addition and mea...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of FP-rhodamine probe binding to KIAA1363 in mouse brain membrane proteome preincubated for 1 hr followed by FP-rhodamine addition and mea...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of FP-rhodamine probe binding to KIAA1363 in mouse brain membrane proteome preincubated for 1 hr followed by FP-rhodamine addition and mea...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of FP-rhodamine probe binding to KIAA1363 in mouse brain membrane proteome preincubated for 1 hr followed by FP-rhodamine addition and mea...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of FP-rhodamine probe binding to KIAA1363 in mouse brain membrane proteome preincubated for 1 hr followed by FP-rhodamine addition and mea...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of FP-rhodamine probe binding to KIAA1363 in mouse brain membrane proteome preincubated for 1 hr followed by FP-rhodamine addition and mea...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of FP-rhodamine probe binding to KIAA1363 in mouse brain membrane proteome preincubated for 1 hr followed by FP-rhodamine addition and mea...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of FP-rhodamine probe binding to KIAA1363 in mouse brain membrane proteome preincubated for 1 hr followed by FP-rhodamine addition and mea...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of FP-rhodamine probe binding to KIAA1363 in mouse brain membrane proteome preincubated for 1 hr followed by FP-rhodamine addition and mea...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of FP-rhodamine probe binding to KIAA1363 in mouse brain membrane proteome preincubated for 1 hr followed by FP-rhodamine addition and mea...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of FP-rhodamine probe binding to KIAA1363 in mouse brain membrane proteome preincubated for 1 hr followed by FP-rhodamine addition and mea...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of FP-rhodamine probe binding to KIAA1363 in mouse brain membrane proteome preincubated for 1 hr followed by FP-rhodamine addition and mea...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of FP-rhodamine probe binding to KIAA1363 in mouse brain membrane proteome preincubated for 1 hr followed by FP-rhodamine addition and mea...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of FP-rhodamine probe binding to KIAA1363 in mouse brain membrane proteome preincubated for 1 hr followed by FP-rhodamine addition and mea...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of FP-rhodamine probe binding to KIAA1363 in mouse brain membrane proteome preincubated for 1 hr followed by FP-rhodamine addition and mea...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of FP-rhodamine probe binding to KIAA1363 in mouse brain membrane proteome preincubated for 1 hr followed by FP-rhodamine addition and mea...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of FP-rhodamine probe binding to KIAA1363 in mouse brain membrane proteome preincubated for 1 hr followed by FP-rhodamine addition and mea...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of FP-rhodamine probe binding to KIAA1363 in mouse brain membrane proteome preincubated for 1 hr followed by FP-rhodamine addition and mea...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of FP-rhodamine probe binding to KIAA1363 in mouse brain membrane proteome preincubated for 1 hr followed by FP-rhodamine addition and mea...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of FP-rhodamine probe binding to KIAA1363 in mouse brain membrane proteome preincubated for 1 hr followed by FP-rhodamine addition and mea...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of FP-rhodamine probe binding to KIAA1363 in mouse brain membrane proteome preincubated for 1 hr followed by FP-rhodamine addition and mea...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of FP-rhodamine probe binding to KIAA1363 in mouse brain membrane proteome preincubated for 1 hr followed by FP-rhodamine addition and mea...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Competitive inhibition of mouse brain membrane KIAA1363 preincubated for 6 hrs followed by FP-rhodamine addition measured after 10 mins by fluorescen...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Competitive inhibition of mouse brain membrane KIAA1363 preincubated for 6 hrs followed by FP-rhodamine addition measured after 10 mins by fluorescen...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of FP-rhodamine probe binding to KIAA1363 in mouse brain membrane proteome preincubated for 1 hr followed by FP-rhodamine addition and mea...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of FP-rhodamine probe binding to KIAA1363 in mouse brain membrane proteome preincubated for 1 hr followed by FP-rhodamine addition and mea...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of FP-rhodamine probe binding to KIAA1363 in mouse brain membrane proteome preincubated for 1 hr followed by FP-rhodamine addition and mea...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of mouse brain KIAA1363 incubated for 10 mins prior to rhodamine-tagged fluorophosphonate addition measured after 10 mins by SDS-PAGEMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of mouse brain KIAA1363 incubated for 10 mins prior to rhodamine-tagged fluorophosphonate addition measured after 10 mins by SDS-PAGEMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Competitive inhibition of mouse brain membrane KIAA1363 preincubated for 6 hrs followed by FP-rhodamine addition measured after 10 mins by fluorescen...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Competitive inhibition of mouse brain membrane KIAA1363 preincubated for 6 hrs followed by FP-rhodamine addition measured after 10 mins by fluorescen...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of mouse brain KIAA1363 incubated for 10 mins prior to rhodamine-tagged fluorophosphonate addition measured after 10 mins by SDS-PAGEMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Competitive inhibition of mouse brain membrane KIAA1363 preincubated for 6 hrs followed by FP-rhodamine addition measured after 10 mins by fluorescen...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of mouse brain KIAA1363 incubated for 10 mins prior to rhodamine-tagged fluorophosphonate addition measured after 10 mins by SDS-PAGEMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Competitive inhibition of mouse brain membrane KIAA1363 preincubated for 6 hrs followed by FP-rhodamine addition measured after 10 mins by fluorescen...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+6nMAssay Description:Inhibition of FP-rhodamine probe binding to KIAA1363 in mouse brain membrane proteome preincubated for 1 hr followed by FP-rhodamine addition and mea...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+6nMAssay Description:Inhibition of FP-rhodamine probe binding to KIAA1363 in mouse brain membrane proteome preincubated for 1 hr followed by FP-rhodamine addition and mea...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+6nMAssay Description:Inhibition of FP-rhodamine probe binding to KIAA1363 in mouse brain membrane proteome preincubated for 1 hr followed by FP-rhodamine addition and mea...More data for this Ligand-Target Pair