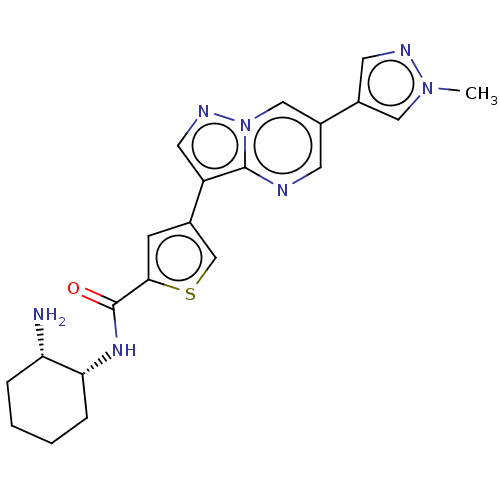
Affinity DataIC50: 2nMAssay Description:Inhibition of MARK4 in mouse cortical neurons assessed as inhibition of okadaic acid-induced tau phosphorylation at S262 residue preincubated for 15 ...More data for this Ligand-Target Pair
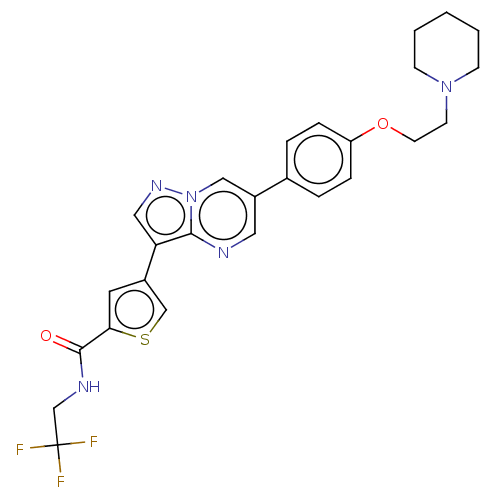
Affinity DataIC50: 50nMAssay Description:Inhibition of MARK4 in mouse cortical neurons assessed as inhibition of okadaic acid-induced tau phosphorylation at S262 residue preincubated for 15 ...More data for this Ligand-Target Pair
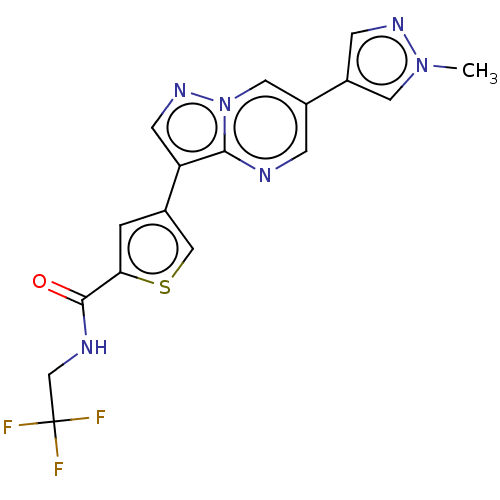
Affinity DataIC50: 60nMAssay Description:Inhibition of MARK4 in mouse cortical neurons assessed as inhibition of okadaic acid-induced tau phosphorylation at S262 residue preincubated for 15 ...More data for this Ligand-Target Pair
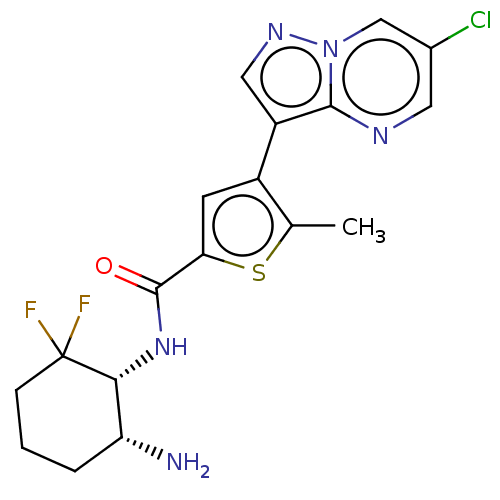
Affinity DataIC50: 280nMAssay Description:Inhibition of MARK4 in mouse cortical neurons assessed as inhibition of okadaic acid-induced tau phosphorylation at S262 residue preincubated for 15 ...More data for this Ligand-Target Pair