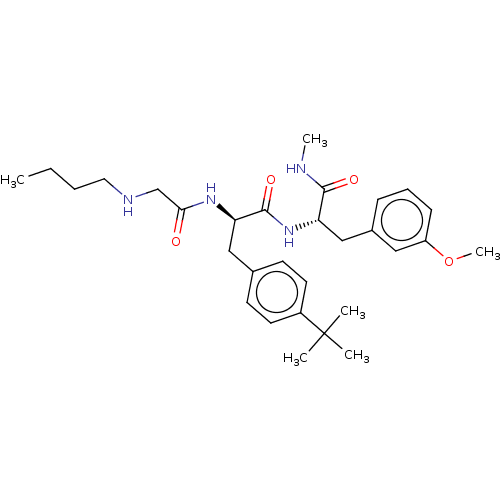
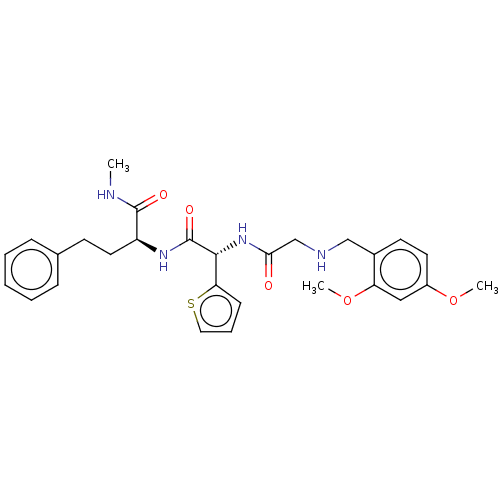
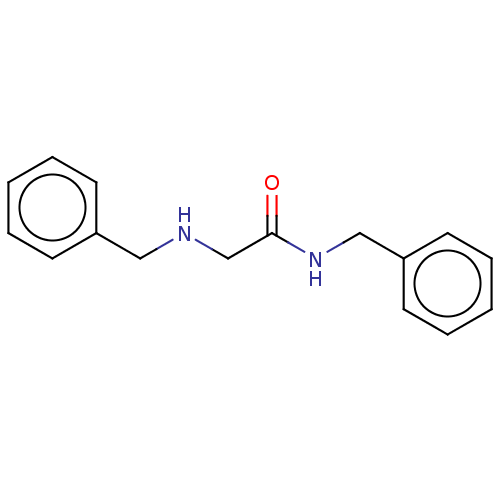
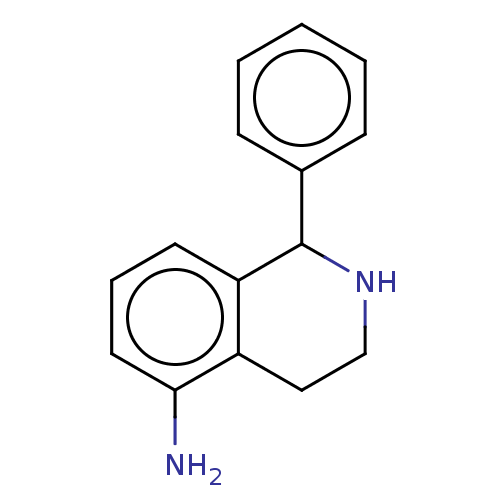
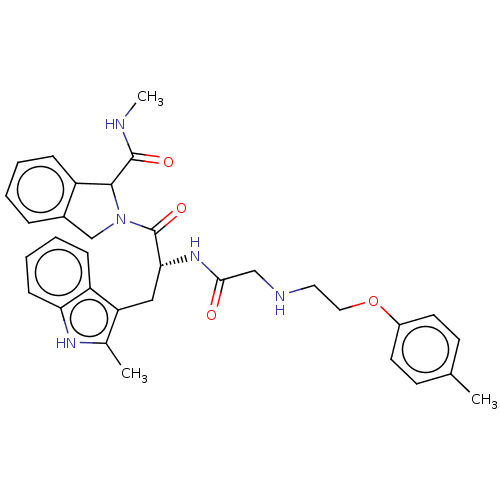
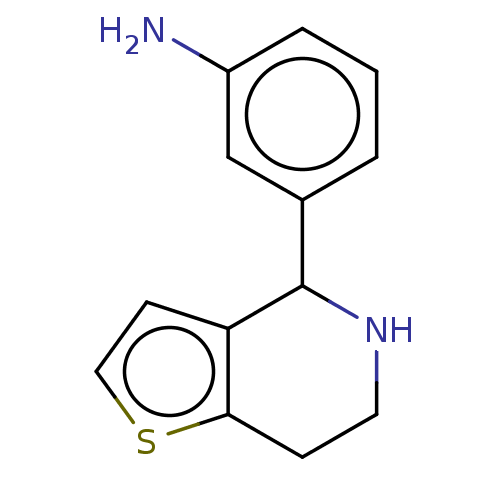
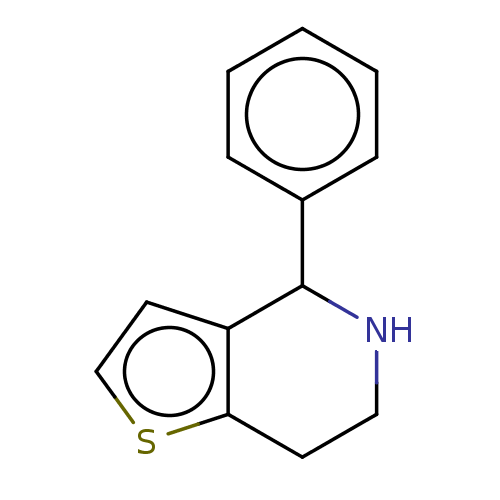
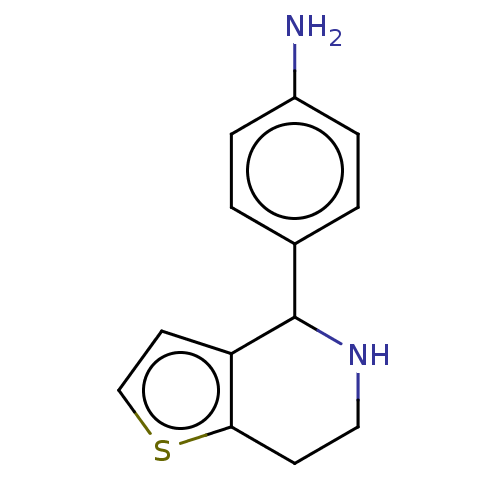
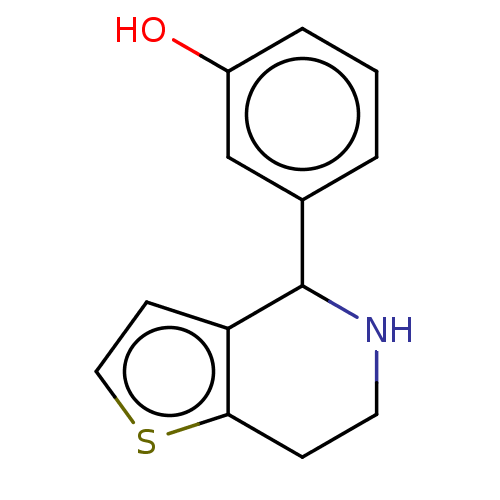
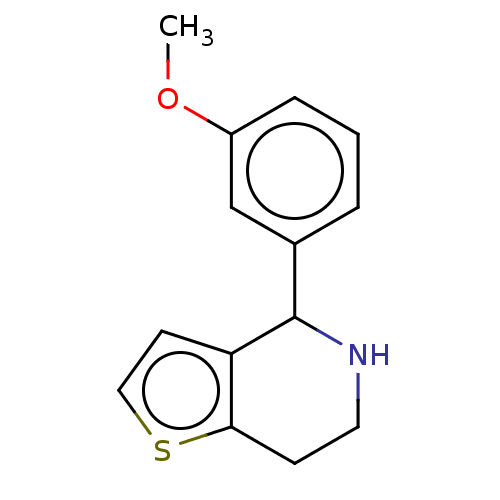
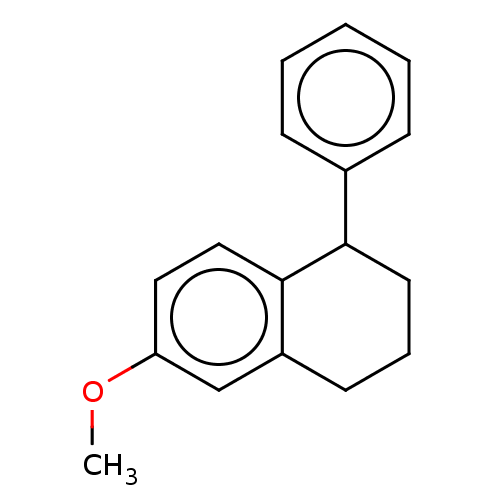
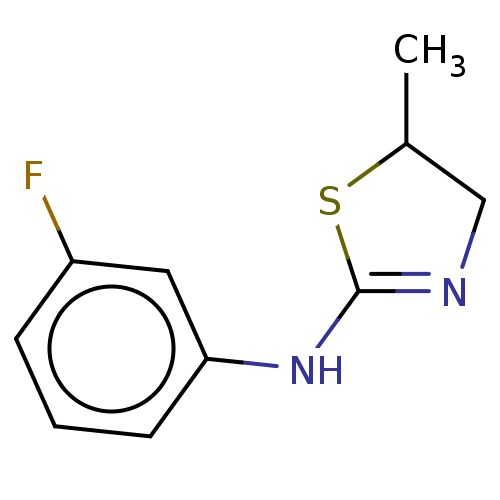
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
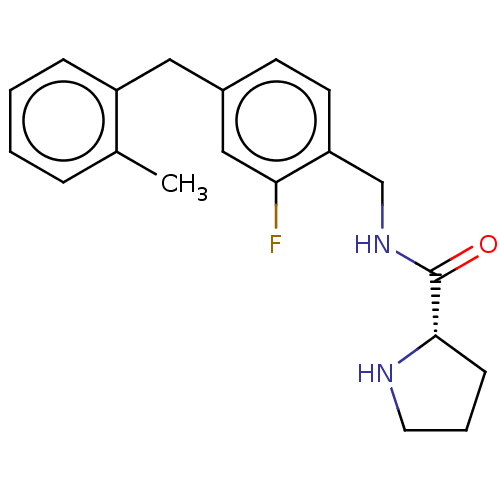
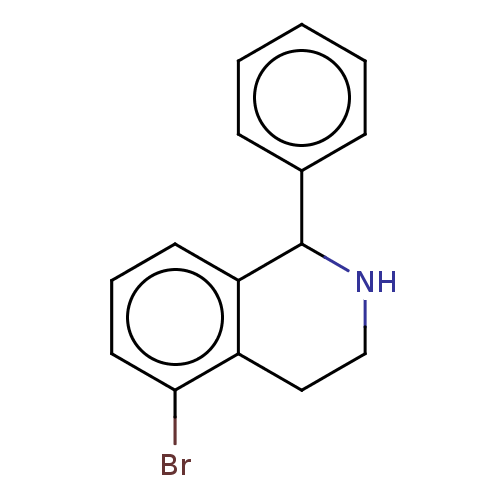
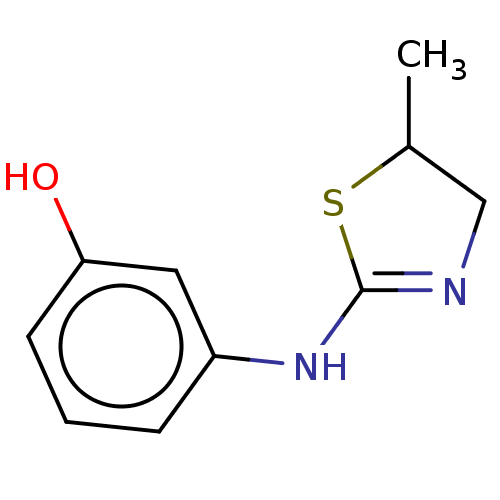
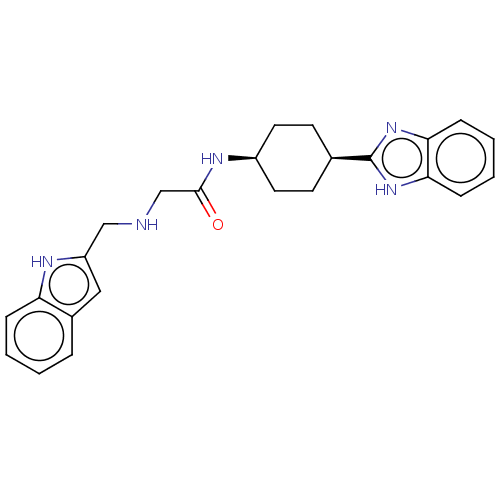
Affinity DataKd: 80nMAssay Description:Binding affinity to GID4 (unknown origin) assessed as dissociation constant by surface plasmon resonance assayMore data for this Ligand-Target Pair
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
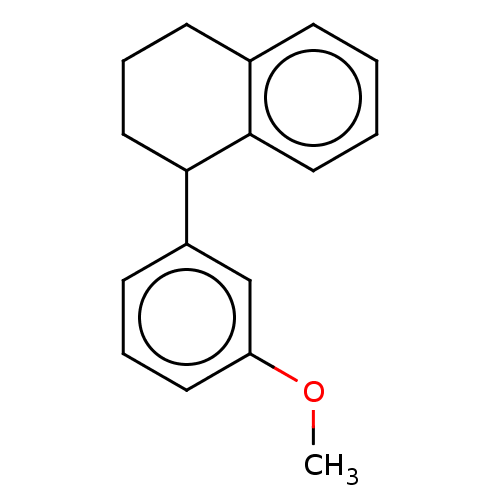
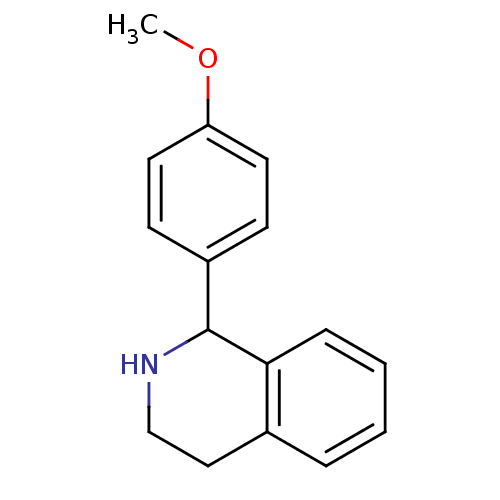
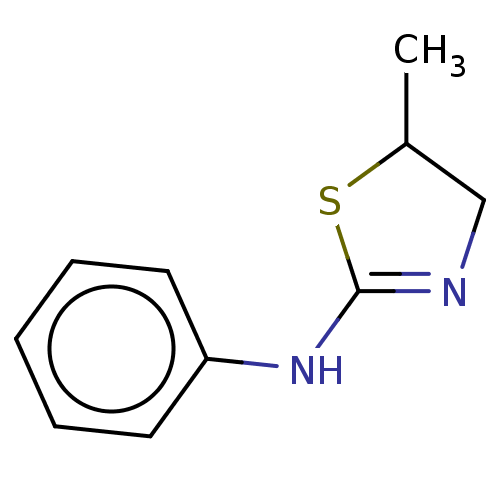
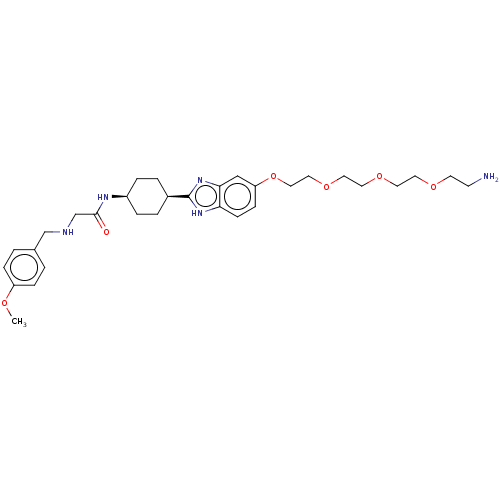
Affinity DataKd: 81nMAssay Description:Binding affinity to recombinant GID4 (unknown origin) assessed as dissociation constant by surface plasmon resonance assayMore data for this Ligand-Target Pair
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
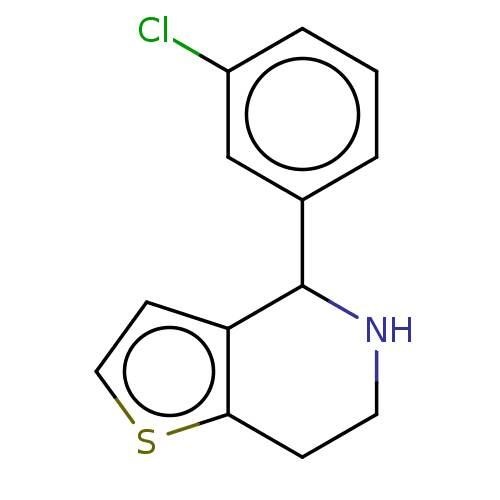
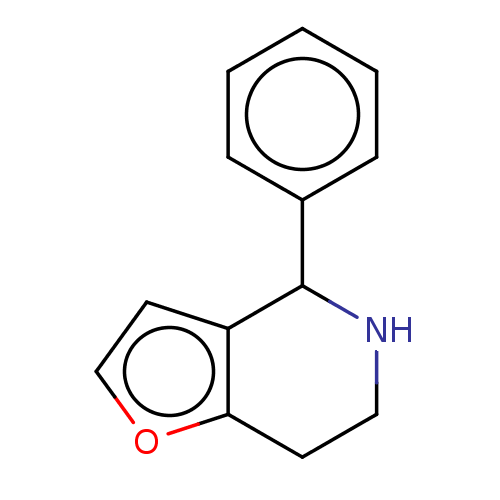
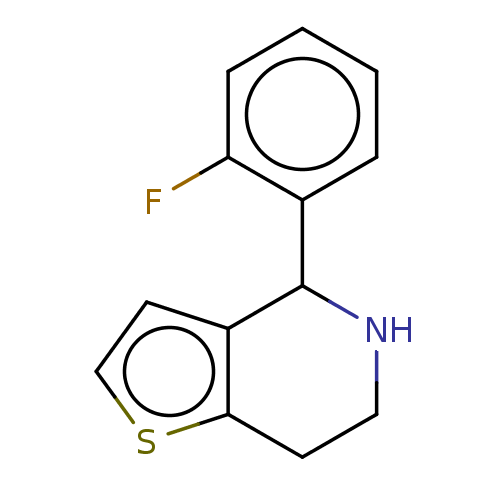
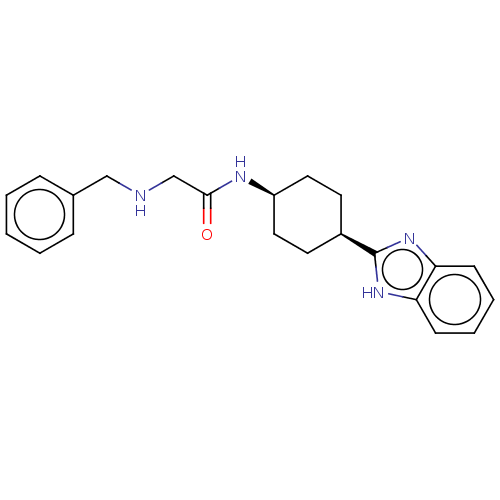
Affinity DataKd: 300nMAssay Description:Binding affinity to recombinant GID4 (unknown origin) assessed as dissociation constant by surface plasmon resonance assayMore data for this Ligand-Target Pair
Ligand InfoSimilars
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
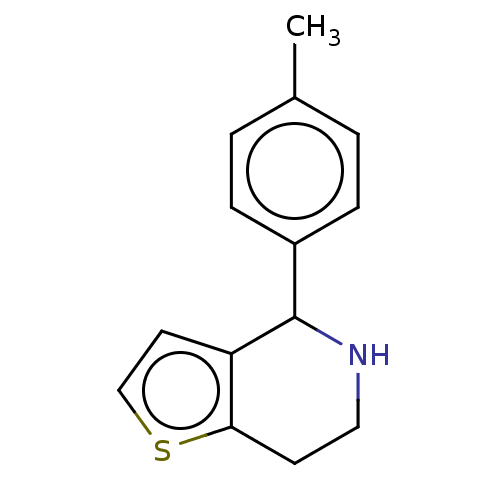
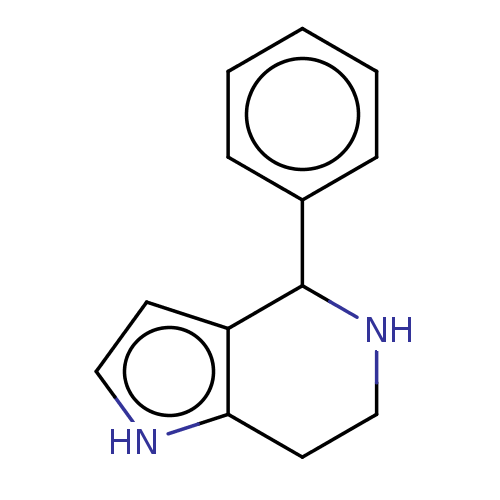
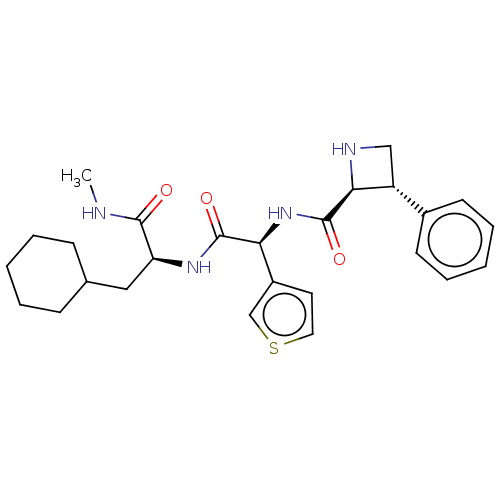
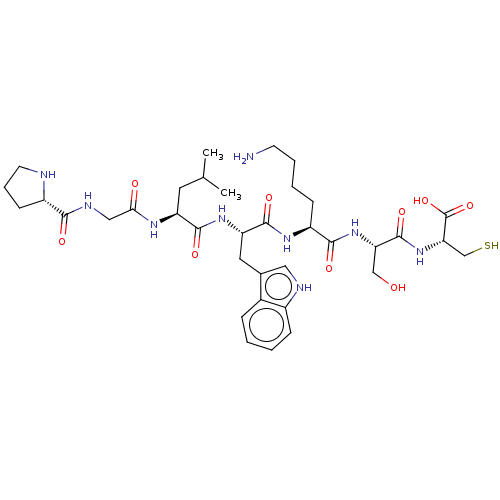
Affinity DataKd: 500nMAssay Description:Binding affinity to recombinant GID4 (unknown origin) assessed as dissociation constant by surface plasmon resonance assayMore data for this Ligand-Target Pair
Ligand InfoSimilars
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Affinity DataKd: 1.90E+3nMAssay Description:Binding affinity to GID4 (unknown origin) assessed as dissociation constantMore data for this Ligand-Target Pair
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Affinity DataKd: 2.30E+3nMAssay Description:Binding affinity to recombinant GID4 (unknown origin) assessed as dissociation constant by surface plasmon resonance assayMore data for this Ligand-Target Pair
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Affinity DataIC50: 2.50E+3nMAssay Description:Displacement of C-terminal NanoLuc-tagged MPGLWKS peptide from N-terminal HaloTag-tagged GID4 (unknown origin) expressed in HEK293T cells incubated f...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Affinity DataKd: 2.70E+3nMAssay Description:Binding affinity to recombinant GID4 (unknown origin) assessed as dissociation constant by surface plasmon resonance assayMore data for this Ligand-Target Pair
Ligand InfoSimilars
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Affinity DataKd: 3.40E+3nMAssay Description:Binding affinity to recombinant GID4 (unknown origin) assessed as dissociation constant by surface plasmon resonance assayMore data for this Ligand-Target Pair
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Affinity DataIC50: 4.10E+3nMAssay Description:Inhibition of GID4 (124 to 289) (unknown origin) expressed in Escherichia coli BL21(DE3) using PGLWKS-FITC peptide by competitive-fluorescence polari...More data for this Ligand-Target Pair
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Affinity DataIC50: 4.70E+3nMAssay Description:Inhibition of GID4 (124 to 289) (unknown origin) expressed in Escherichia coli BL21(DE3) using PGLWKS-FITC peptide by competitive-fluorescence polari...More data for this Ligand-Target Pair
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Affinity DataIC50: 4.80E+3nMAssay Description:Inhibition of GID4 (124 to 289) (unknown origin) expressed in Escherichia coli BL21(DE3) using PGLWKS-FITC peptide by competitive-fluorescence polari...More data for this Ligand-Target Pair
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Affinity DataIC50: 5.40E+3nMAssay Description:Inhibition of GID4 (124 to 289) (unknown origin) expressed in Escherichia coli BL21(DE3) using PGLWKS-FITC peptide by competitive-fluorescence polari...More data for this Ligand-Target Pair
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Affinity DataKd: 6.05E+3nMAssay Description:Binding affinity to His6-tagged GID4 (114 to 300 residues)(unknown origin) expressed in Escherichia coli Rosetta (DE3) assessed as dissociation const...More data for this Ligand-Target Pair
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Affinity DataKd: 9.00E+3nMAssay Description:Binding affinity to recombinant GID4 (unknown origin) assessed as dissociation constant by surface plasmon resonance assayMore data for this Ligand-Target Pair
Ligand InfoSimilars
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Affinity DataKd: 1.50E+4nMAssay Description:Binding affinity to recombinant GID4 (unknown origin) assessed as dissociation constant by surface plasmon resonance assayMore data for this Ligand-Target Pair
Ligand InfoSimilars
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Affinity DataKd: 1.70E+4nMAssay Description:Binding affinity to GID4 (124 to 289) (unknown origin) assessed as dissociation constant measured by isothermal titration calorimetry assayMore data for this Ligand-Target Pair
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Affinity DataIC50: 1.89E+4nMAssay Description:Inhibition of GID4 (124 to 289) (unknown origin) expressed in Escherichia coli BL21(DE3) using PGLWKS-FITC peptide by competitive-fluorescence polari...More data for this Ligand-Target Pair
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Affinity DataIC50: 2.14E+4nMAssay Description:Inhibition of GID4 (124 to 289) (unknown origin) expressed in Escherichia coli BL21(DE3) using PGLWKS-FITC peptide by competitive-fluorescence polari...More data for this Ligand-Target Pair
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Affinity DataKd: 2.40E+4nMAssay Description:Binding affinity to recombinant GID4 (unknown origin) assessed as dissociation constant by surface plasmon resonance assayMore data for this Ligand-Target Pair
Ligand InfoSimilars
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Affinity DataIC50: 3.55E+4nMAssay Description:Inhibition of GID4 (124 to 289) (unknown origin) expressed in Escherichia coli BL21(DE3) using PGLWKS-FITC peptide by competitive-fluorescence polari...More data for this Ligand-Target Pair
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Affinity DataIC50: 4.93E+4nMAssay Description:Inhibition of GID4 (124 to 289) (unknown origin) expressed in Escherichia coli BL21(DE3) using PGLWKS-FITC peptide by competitive-fluorescence polari...More data for this Ligand-Target Pair
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Affinity DataIC50: 5.50E+4nMAssay Description:Inhibition of GID4 (124 to 289) (unknown origin) expressed in Escherichia coli BL21(DE3) using PGLWKS-FITC peptide by competitive-fluorescence polari...More data for this Ligand-Target Pair
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Affinity DataIC50: 6.48E+4nMAssay Description:Inhibition of GID4 (124 to 289) (unknown origin) expressed in Escherichia coli BL21(DE3) using PGLWKS-FITC peptide by competitive-fluorescence polari...More data for this Ligand-Target Pair
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Affinity DataIC50: 8.98E+4nMAssay Description:Inhibition of GID4 (124 to 289) (unknown origin) expressed in Escherichia coli BL21(DE3) using PGLWKS-FITC peptide by competitive-fluorescence polari...More data for this Ligand-Target Pair
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Affinity DataIC50: 9.08E+4nMAssay Description:Inhibition of GID4 (124 to 289) (unknown origin) expressed in Escherichia coli BL21(DE3) using PGLWKS-FITC peptide by competitive-fluorescence polari...More data for this Ligand-Target Pair
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Affinity DataIC50: 1.08E+5nMAssay Description:Inhibition of GID4 (124 to 289) (unknown origin) expressed in Escherichia coli BL21(DE3) using PGLWKS-FITC peptide by competitive-fluorescence polari...More data for this Ligand-Target Pair
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Affinity DataKd: 1.10E+5nMAssay Description:Binding affinity to GID4 (124 to 289) (unknown origin) assessed as dissociation constant measured by isothermal titration calorimetry assayMore data for this Ligand-Target Pair
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Affinity DataIC50: 1.13E+5nMAssay Description:Inhibition of GID4 (124 to 289) (unknown origin) expressed in Escherichia coli BL21(DE3) using PGLWKS-FITC peptide by competitive-fluorescence polari...More data for this Ligand-Target Pair
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Affinity DataIC50: 1.49E+5nMAssay Description:Inhibition of GID4 (124 to 289) (unknown origin) expressed in Escherichia coli BL21(DE3) using PGLWKS-FITC peptide by competitive-fluorescence polari...More data for this Ligand-Target Pair
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Affinity DataIC50: 1.49E+5nMAssay Description:Inhibition of GID4 (124 to 289) (unknown origin) expressed in Escherichia coli BL21(DE3) using PGLWKS-FITC peptide by competitive-fluorescence polari...More data for this Ligand-Target Pair
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Affinity DataIC50: 1.88E+5nMAssay Description:Inhibition of GID4 (124 to 289) (unknown origin) expressed in Escherichia coli BL21(DE3) using PGLWKS-FITC peptide by competitive-fluorescence polari...More data for this Ligand-Target Pair
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Affinity DataIC50: 1.88E+5nMAssay Description:Inhibition of GID4 (124 to 289) (unknown origin) expressed in Escherichia coli BL21(DE3) using PGLWKS-FITC peptide by competitive-fluorescence polari...More data for this Ligand-Target Pair
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Affinity DataIC50: 2.35E+5nMAssay Description:Inhibition of GID4 (124 to 289) (unknown origin) expressed in Escherichia coli BL21(DE3) using PGLWKS-FITC peptide by competitive-fluorescence polari...More data for this Ligand-Target Pair
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of GID4 (124 to 289) (unknown origin) expressed in Escherichia coli BL21(DE3) using PGLWKS-FITC peptide by competitive-fluorescence polari...More data for this Ligand-Target Pair
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of GID4 (124 to 289) (unknown origin) expressed in Escherichia coli BL21(DE3) using PGLWKS-FITC peptide by competitive-fluorescence polari...More data for this Ligand-Target Pair
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of GID4 (124 to 289) (unknown origin) expressed in Escherichia coli BL21(DE3) using PGLWKS-FITC peptide by competitive-fluorescence polari...More data for this Ligand-Target Pair
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of GID4 (124 to 289) (unknown origin) expressed in Escherichia coli BL21(DE3) using PGLWKS-FITC peptide by competitive-fluorescence polari...More data for this Ligand-Target Pair
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of GID4 (124 to 289) (unknown origin) expressed in Escherichia coli BL21(DE3) using PGLWKS-FITC peptide by competitive-fluorescence polari...More data for this Ligand-Target Pair
TargetGlucose-induced degradation protein 4 homolog(Human)
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Lunenfeld-Tanenbaum Research Institute
Curated by ChEMBL
Affinity DataIC50: 2.64E+5nMAssay Description:Inhibition of GID4 (124 to 289) (unknown origin) expressed in Escherichia coli BL21(DE3) using PGLWKS-FITC peptide by competitive-fluorescence polari...More data for this Ligand-Target Pair
 3D Structure (crystal)
3D Structure (crystal)