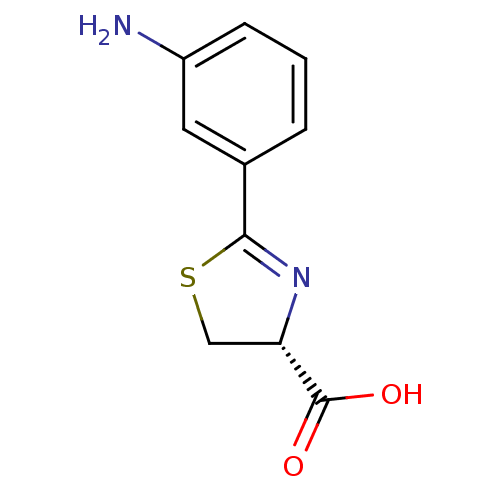
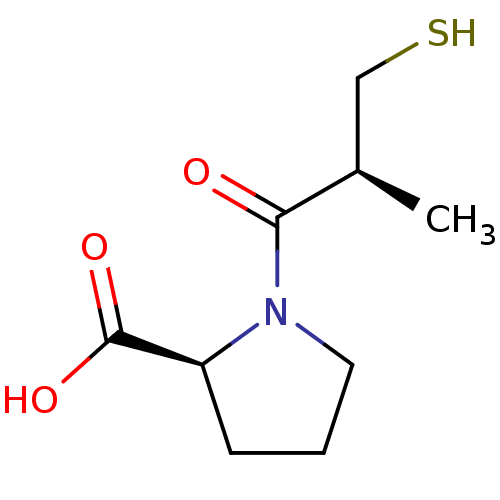
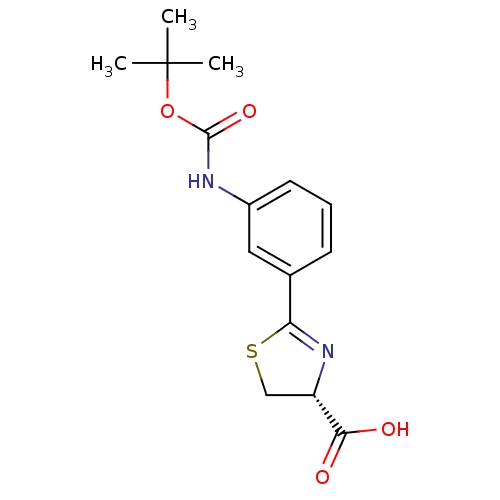
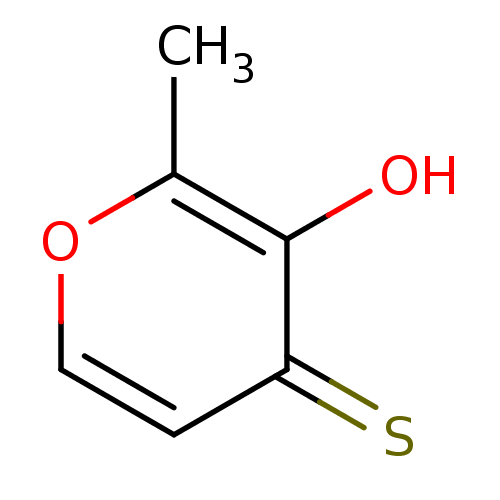
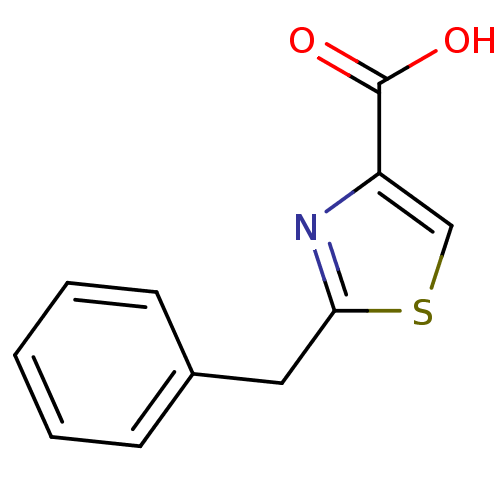
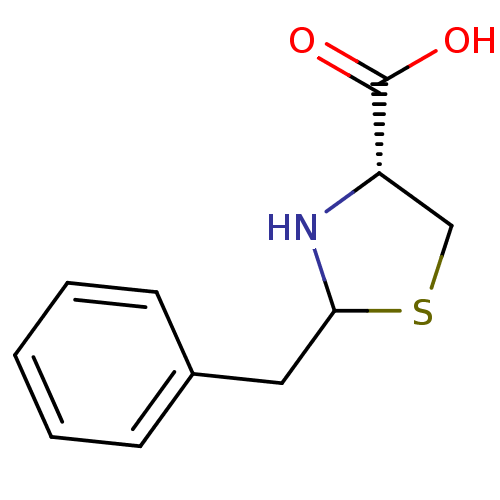
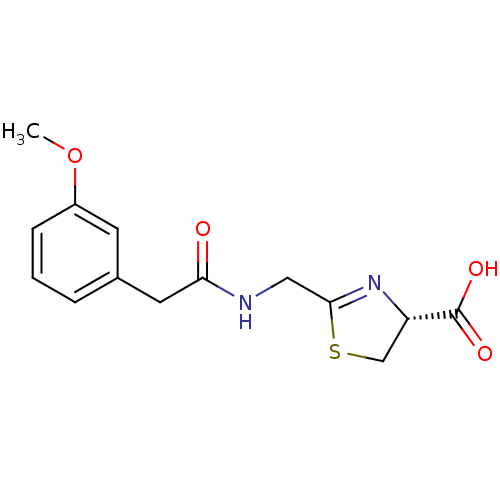
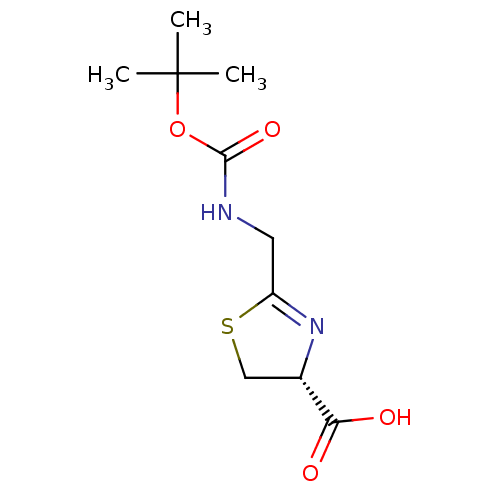
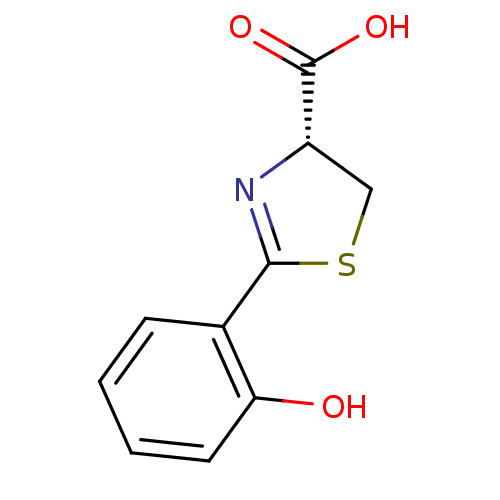
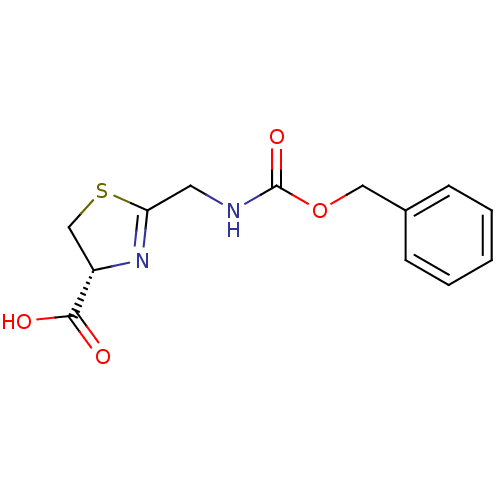
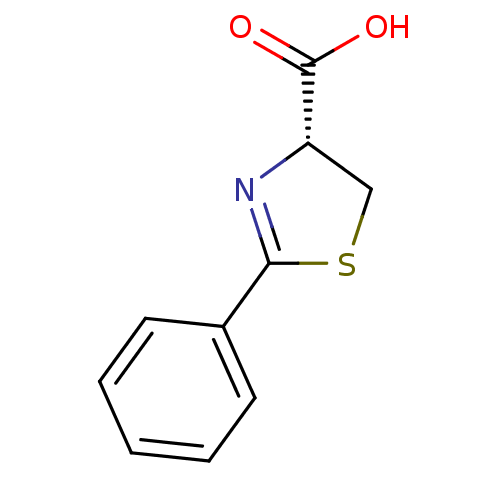
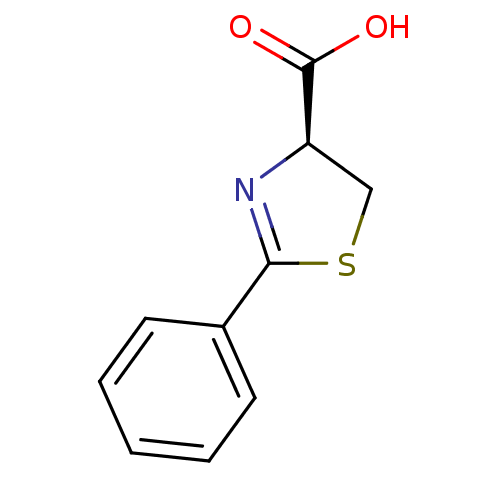
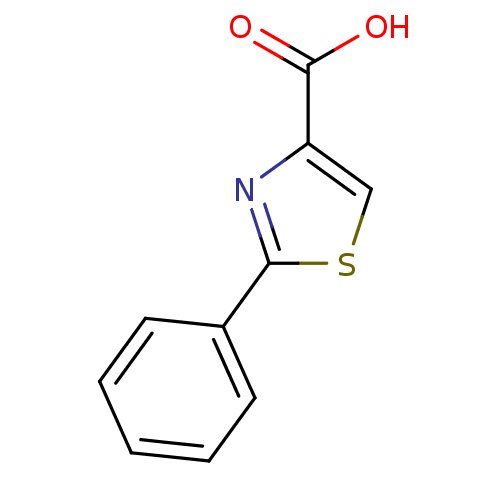
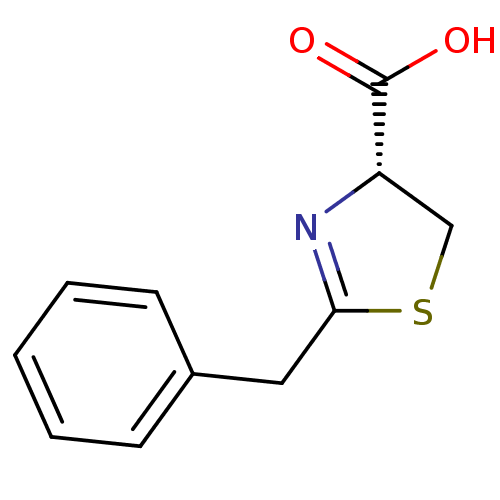
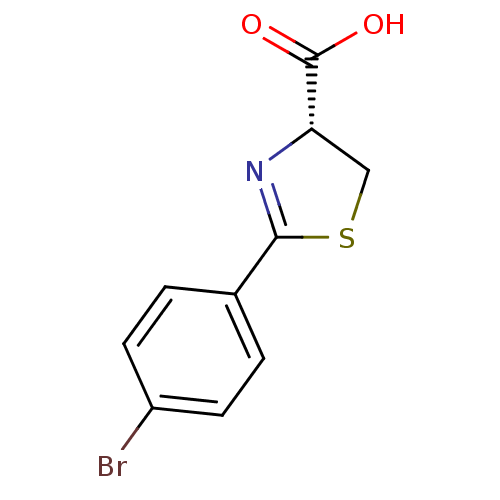
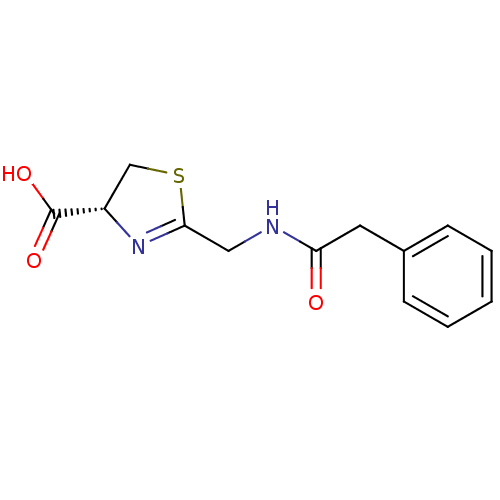
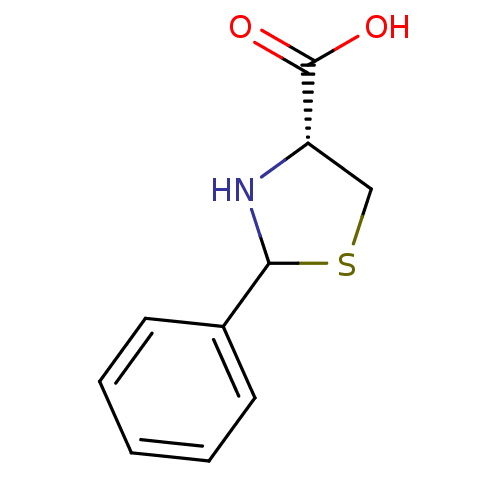
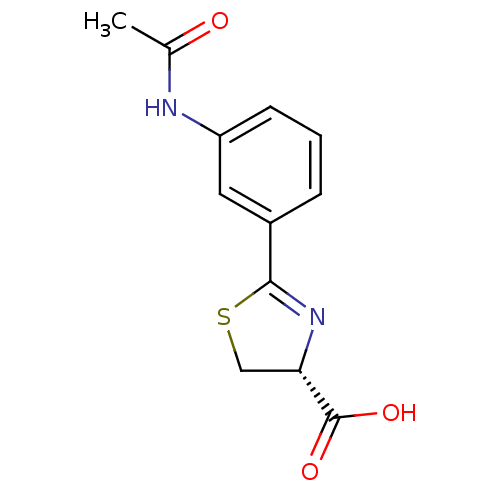
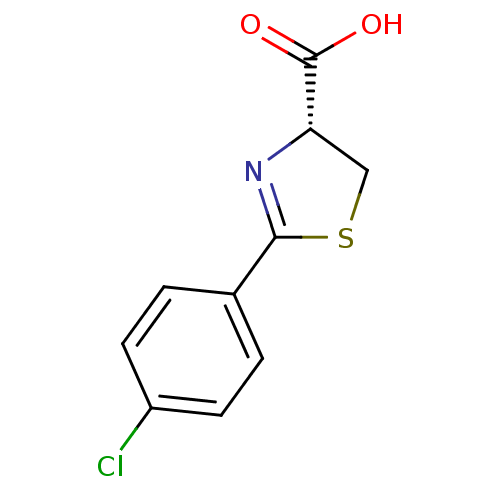
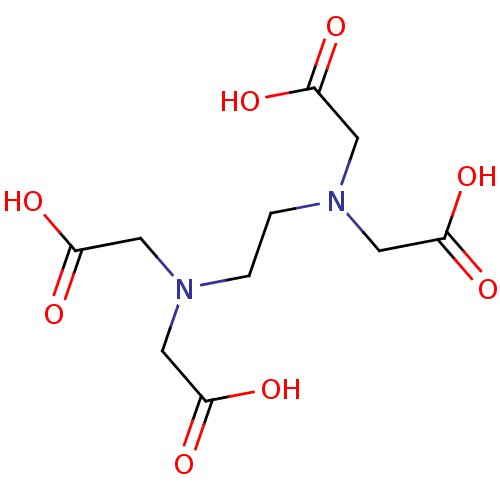
Affinity DataIC50: 4.90E+3nMAssay Description:Inhibition of Bacillus anthracis MBL Bla2 using nitrocefin as substrate preincubated for 20 mins by UV-spectrophotometric analysisMore data for this Ligand-Target Pair
Affinity DataKi: 5.10E+3nMAssay Description:Inhibition of Bacillus anthracis MBL Bla2 using nitrocefin as substrate preincubated for 20 mins by UV-spectrophotometric analysisMore data for this Ligand-Target Pair
Affinity DataKi: 1.79E+4nMAssay Description:Inhibition of Bacillus anthracis MBL Bla2 using nitrocefin as substrate preincubated for 20 mins by UV-spectrophotometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 2.58E+4nMAssay Description:Inhibition of Bacillus anthracis MBL Bla2 using nitrocefin as substrate preincubated for 20 mins by UV-spectrophotometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 7.41E+4nMAssay Description:Inhibition of Bacillus anthracis MBL Bla2 using nitrocefin as substrate preincubated for 20 mins by UV-spectrophotometric analysisMore data for this Ligand-Target Pair
Affinity DataKi: 8.50E+4nMpH: 7.0Assay Description:Progress curves for slow-binding inhibition analysis were obtained by adding nitrocefin (final concentration 36 μM) to a solution containing enz...More data for this Ligand-Target Pair
Affinity DataIC50: 1.59E+5nMpH: 7.0Assay Description:Bla2, in 50 mM MOPS and 30% (v/v) glycerol, was assayed for substrate activity by adding enzyme at a final concentration of 0.13 μg/mL to soluti...More data for this Ligand-Target Pair
Affinity DataIC50: 1.68E+5nMpH: 7.0Assay Description:Bla2, in 50 mM MOPS and 30% (v/v) glycerol, was assayed for substrate activity by adding enzyme at a final concentration of 0.13 μg/mL to soluti...More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+5nMAssay Description:Inhibition of Bacillus anthracis MBL Bla2 using nitrocefin as substrate preincubated for 20 mins by UV-spectrophotometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+5nMAssay Description:Inhibition of Bacillus anthracis MBL Bla2 using nitrocefin as substrate preincubated for 20 mins by UV-spectrophotometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+5nMAssay Description:Inhibition of Bacillus anthracis MBL Bla2 using nitrocefin as substrate preincubated for 20 mins by UV-spectrophotometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+5nMAssay Description:Inhibition of Bacillus anthracis MBL Bla2 using nitrocefin as substrate preincubated for 20 mins by UV-spectrophotometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+5nMAssay Description:Inhibition of Bacillus anthracis MBL Bla2 using nitrocefin as substrate preincubated for 20 mins by UV-spectrophotometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+5nMAssay Description:Inhibition of Bacillus anthracis MBL Bla2 using nitrocefin as substrate preincubated for 20 mins by UV-spectrophotometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+5nMAssay Description:Inhibition of Bacillus anthracis MBL Bla2 using nitrocefin as substrate preincubated for 20 mins by UV-spectrophotometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+5nMAssay Description:Inhibition of Bacillus anthracis MBL Bla2 using nitrocefin as substrate preincubated for 20 mins by UV-spectrophotometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+5nMAssay Description:Inhibition of Bacillus anthracis MBL Bla2 using nitrocefin as substrate preincubated for 20 mins by UV-spectrophotometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+5nMAssay Description:Inhibition of Bacillus anthracis MBL Bla2 using nitrocefin as substrate preincubated for 20 mins by UV-spectrophotometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+5nMAssay Description:Inhibition of Bacillus anthracis MBL Bla2 using nitrocefin as substrate preincubated for 20 mins by UV-spectrophotometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+5nMAssay Description:Inhibition of Bacillus anthracis MBL Bla2 using nitrocefin as substrate preincubated for 20 mins by UV-spectrophotometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+5nMAssay Description:Inhibition of Bacillus anthracis MBL Bla2 using nitrocefin as substrate preincubated for 20 mins by UV-spectrophotometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+5nMAssay Description:Inhibition of Bacillus anthracis MBL Bla2 using nitrocefin as substrate preincubated for 20 mins by UV-spectrophotometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+5nMAssay Description:Inhibition of Bacillus anthracis MBL Bla2 using nitrocefin as substrate preincubated for 20 mins by UV-spectrophotometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 2.04E+5nMpH: 7.0Assay Description:Bla2, in 50 mM MOPS and 30% (v/v) glycerol, was assayed for substrate activity by adding enzyme at a final concentration of 0.13 μg/mL to soluti...More data for this Ligand-Target Pair
Affinity DataIC50: 2.35E+5nMpH: 7.0Assay Description:Bla2, in 50 mM MOPS and 30% (v/v) glycerol, was assayed for substrate activity by adding enzyme at a final concentration of 0.13 μg/mL to soluti...More data for this Ligand-Target Pair
Affinity DataIC50: 2.90E+5nMpH: 7.0Assay Description:Bla2, in 50 mM MOPS and 30% (v/v) glycerol, was assayed for substrate activity by adding enzyme at a final concentration of 0.13 μg/mL to soluti...More data for this Ligand-Target Pair
Affinity DataIC50: 3.67E+5nMpH: 7.0Assay Description:Bla2, in 50 mM MOPS and 30% (v/v) glycerol, was assayed for substrate activity by adding enzyme at a final concentration of 0.13 μg/mL to soluti...More data for this Ligand-Target Pair
Affinity DataIC50: 1.06E+6nMAssay Description:Inhibition of Bacillus anthracis MBL Bla2 using nitrocefin as substrate after 12 hrs by UV-spectrophotometric analysisMore data for this Ligand-Target Pair