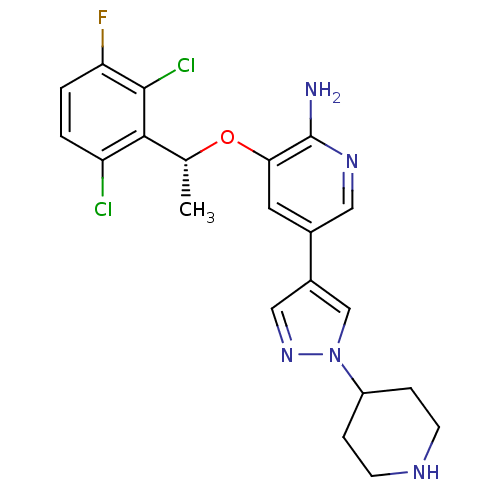
BDBM50306682 (R)-3-(1-(2,6-dichloro-3-fluorophenyl)ethoxy)-5-(1-(piperidin-4-yl)-1H-pyrazol-4-yl)pyridin-2-amine::3-(2,6-dichloro-3-fluorobenzyloxy)-5-(1-(piperidin-4-yl)-1H-pyrazol-4-yl)pyridin-2-amine::3-[(1R)-1-(2,6-dichloro-3-fluorophenyl)ethoxy]-5-(1-piperidin-4-yl-1H-pyrazol-4-yl)pyridin-2-amine::CHEMBL601719::CRIZOTINIB::PF-2341066::US10370379, Crizotinib::US10543199, Compound Crizotinib::US10780082, Compound Crizotinib::US11059827, Compound Crizotinib::US11517561, Compound Crizotinib::US9126941, PF-2341066::US9199944, Crizotinib::US9226923, Crizotinib
SMILES C[C@@H](Oc1cc(cnc1N)-c1cnn(c1)C1CCNCC1)c1c(Cl)ccc(F)c1Cl
InChI Key InChIKey=KTEIFNKAUNYNJU-GFCCVEGCSA-N
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 4 hits for monomerid = 50306682
Found 4 hits for monomerid = 50306682
Affinity DataIC50: >2.5nMAssay Description:Inhibition of N-terminal NH-tagged and avi-tagged dephosphorylated c-MET D1228V mutant (956 to 1390 residues) (unknown origin) expressed in sf21 cell...More data for this Ligand-Target Pair
Affinity DataIC50: >2.5nMAssay Description:Inhibition of wild type N-terminal NH-tagged and avi-tagged dephosphorylated c-MET (956 to 1390 residues) (unknown origin) expressed in sf21 cells us...More data for this Ligand-Target Pair
Affinity DataIC50: 11nMAssay Description:Inhibition of c-MET (unknown origin) assessed as reduction in ADP production incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 200nMAssay Description:Inhibition of MST1R (unknown origin)More data for this Ligand-Target Pair
 3D Structure (crystal)
3D Structure (crystal)