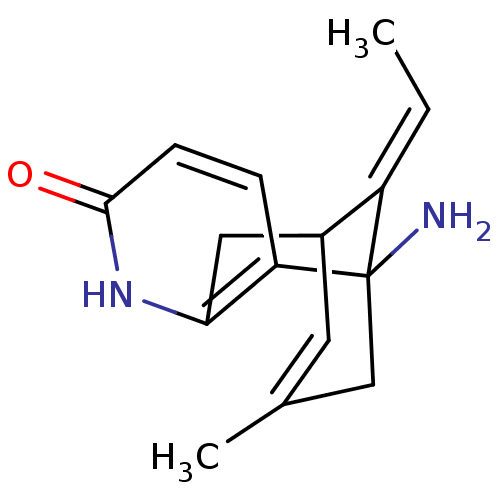
BDBM10441 (+)-Huperzine A::(+/-)Huperzine A::(-)-Huperzine A::(13E)-1-amino-13-ethylidene-11-methyl-6-azatricyclo[7.3.1.0^{2,7}]trideca-2(7),3,10-trien-5-one::(1R,13E)-1-amino-13-ethylidene-11-methyl-6-azatricyclo[7.3.1.0^{2,7}]trideca-2(7),3,10-trien-5-one::(1S,13E)-1-amino-13-ethylidene-11-methyl-6-azatricyclo[7.3.1.0^{2,7}]trideca-2(7),3,10-trien-5-one::(5S,11E)-5-amino-11-ethylidene-7-methyl-5,6,9,10-tetrahydro-5,9-methanocycloocta[b]pyridin-2(1H)-one::Huperzine A
SMILES C\C=C1/C2Cc3[nH]c(=O)ccc3C1(N)CC(C)=C2
InChI Key
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 17 hits for monomerid = 10441
Found 17 hits for monomerid = 10441
Affinity DataKi: 4.60nM ΔG°: -11.4kcal/molepH: 7.0 T: 2°CAssay Description:Assays were conducted at 25 C in 20 mM sodium phosphate buffer (pH 7.0) and 0.01% bovine serum albumin unless otherwise noted. AChE concentrations we...More data for this Ligand-Target Pair
Affinity DataKi: 47nM ΔG°: -9.99kcal/molepH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by E llman. Inhibition of enzyme activity was measured over a substrate c...More data for this Ligand-Target Pair
Affinity DataIC50: 47nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The absorbance changes at 412 nm were recorded for 5 min with ...More data for this Ligand-Target Pair
Affinity DataIC50: 47nMpH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The absorbance changes at 412 nm were recorded for 5 min with ...More data for this Ligand-Target Pair
Affinity DataKi: 47.1nM ΔG°: -9.89kcal/mole IC50: 114nMpH: 7.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at ...More data for this Ligand-Target Pair
Affinity DataIC50: 72.4nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataKi: 175nM ΔG°: -9.21kcal/molepH: 7.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at ...More data for this Ligand-Target Pair
Affinity DataKi: 175nM ΔG°: -9.12kcal/mole IC50: 414nMpH: 7.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at ...More data for this Ligand-Target Pair
Affinity DataIC50: 260nMpH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 648nMpH: 8.0Assay Description:The reaction mixture was consisted of 100 µL of the total volume. To the solution of PBS (pH 8.0, 30 µL) consisted of KH2PO4 and K2HPO4 in 96-well pl...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+3nM ΔG°: >-8.18kcal/molepH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by E llman. Inhibition of enzyme activity was measured over a substrate c...More data for this Ligand-Target Pair
Affinity DataKi: 4.30E+3nM ΔG°: -7.32kcal/molepH: 7.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The absorbance changes at 412 nm were recorded for 5 min with ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The absorbance changes at 412 nm were recorded for 5 min with ...More data for this Ligand-Target Pair
Affinity DataIC50: 7.02E+4nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataKi: 4.30E+6nMAssay Description:Inhibition of Torpedo californica AChEMore data for this Ligand-Target Pair