BDBM107244 US8598345, 200::US8598345, Table 1-30/2-4
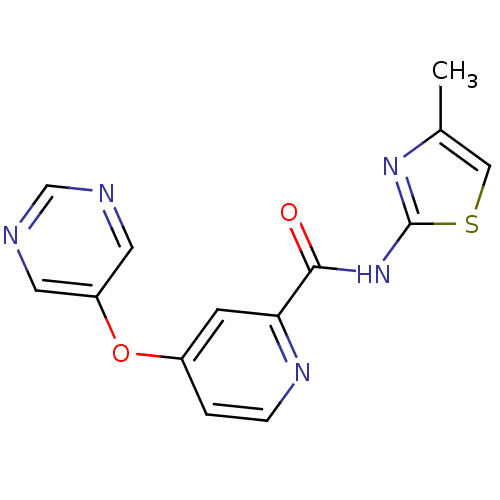
SMILES Cc1csc(NC(=O)c2cc(Oc3cncnc3)ccn2)n1
InChI Key InChIKey=HQSAJPPYHDESIU-UHFFFAOYSA-N
Data 3 IC50
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 3 hits for monomerid = 107244
Found 3 hits for monomerid = 107244
Affinity DataIC50: 100nMAssay Description:Inhibition of CYP1A2 in human liver microsomes using tacrine as substrate preincubated for 5 mins followed by NADPH addition measured after 8 mins by...More data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 5(Rat)
Vanderbilt University Institute of Imaging Science
Curated by ChEMBL
Vanderbilt University Institute of Imaging Science
Curated by ChEMBL
Affinity DataIC50: 200nMAssay Description:Negative allosteric modulation of rat mGlu5 receptor expressed in HEK293A cells assessed as inhibition of glutamate induced-calcium mobilization prei...More data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 5(Rat)
Vanderbilt University Institute of Imaging Science
Curated by ChEMBL
Vanderbilt University Institute of Imaging Science
Curated by ChEMBL
Affinity DataIC50: 200nMAssay Description:Negative allosteric modulation of rat mGlu5 receptor expressed in HEK293A cells assessed as inhibition of glutamate induced-calcium mobilization prei...More data for this Ligand-Target Pair