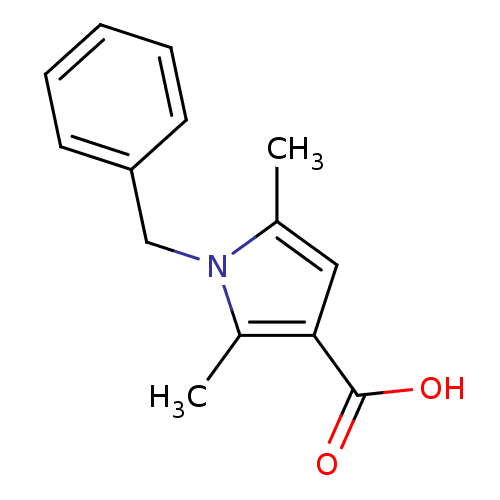
BDBM41732 1-benzyl-2,5-dimethyl-pyrrole-3-carboxylic acid::1-benzyl-2,5-dimethylpyrrole-3-carboxylic acid::2,5-dimethyl-1-(phenylmethyl)-3-pyrrolecarboxylic acid::2,5-dimethyl-1-(phenylmethyl)pyrrole-3-carboxylic acid::SR-01000106152-2::cid_935740
SMILES Cc1cc(C(O)=O)c(C)n1Cc1ccccc1
InChI Key InChIKey=YBCBUHIBNNSACV-UHFFFAOYSA-N
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 2 hits for monomerid = 41732
Found 2 hits for monomerid = 41732
Affinity DataIC50: 1.30E+4nMpH: 6.2 T: 2°CAssay Description:13 fragments with strong inhibition in the single concentration point assay were chose from the micelle screen for further analysis using a biochemic...More data for this Ligand-Target Pair
Affinity DataIC50: 1.40E+4nMAssay Description:Inhibition of human Notum (S81 to T451 residues) Cys330Ser mutant expressed in HEK293S GnTI cells using OPTS as substrate incubated for 40 mins by fl...More data for this Ligand-Target Pair