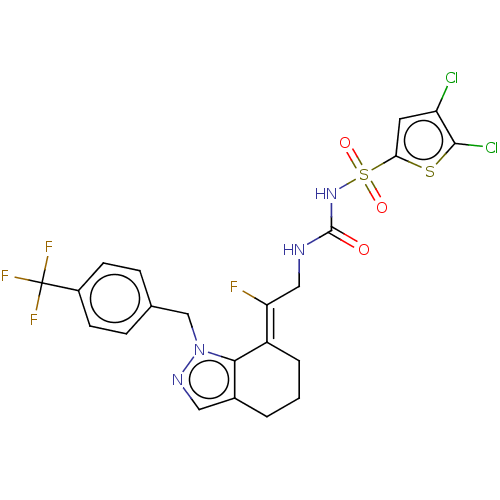
BDBM50580036 CHEMBL5086151
SMILES F\C(CNC(=O)NS(=O)(=O)c1cc(Cl)c(Cl)s1)=C1\CCCc2cnn(Cc3ccc(cc3)C(F)(F)F)c12
InChI Key InChIKey=DFKICTCSFWHWMW-UHFFFAOYSA-N
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 2 hits for monomerid = 50580036
Found 2 hits for monomerid = 50580036
Affinity DataKi: 7nMAssay Description:Displacement of [3H]-PGE2 from human EP3 receptor assessed as inhibition constant incubated for 2 hrs by TopCount scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataIC50: 52nMAssay Description:Antagonist activity at human EP3 receptor expressed in CHO-K1 cells assessed as reduction in sulprostone induced inhibition of forskolin stimulated c...More data for this Ligand-Target Pair