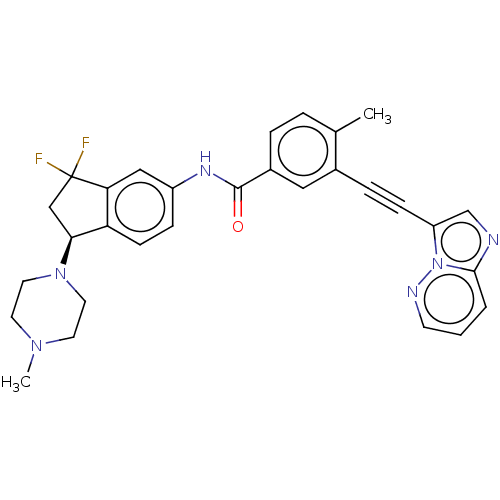
BDBM50582559 CHEMBL5093894
SMILES CN1CCN(CC1)[C@H]1CC(F)(F)c2cc(NC(=O)c3ccc(C)c(c3)C#Cc3cnc4cccnn34)ccc12
InChI Key InChIKey=BPLVEFPXYUJTQJ-UHFFFAOYSA-N
Data 11 IC50
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 11 hits for monomerid = 50582559
Found 11 hits for monomerid = 50582559
TargetProto-oncogene tyrosine-protein kinase Src(Human)
Peking Union Medical College
Curated by ChEMBL
Peking Union Medical College
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Inhibition of human c-SRC using poly [Glu:Tyr] as substrate incubated for 30 mins followed by 33P-ATP addition and measured after 2 hrs by liquid sci...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Inhibition of human FGFR1 using KKKSPGEYVNIEFG as substrate incubated for 30 mins followed by 33P-ATP addition and measured after 2 hrs by liquid sci...More data for this Ligand-Target Pair
TargetReceptor-type tyrosine-protein kinase FLT3(Human)
Peking Union Medical College
Curated by ChEMBL
Peking Union Medical College
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Inhibition of human FLT3 using EAIYAAPFAKKK as substrate incubated for 30 mins followed by 33P-ATP addition and measured after 2 hrs by liquid scinti...More data for this Ligand-Target Pair
TargetVascular endothelial growth factor receptor 2(Human)
Peking Union Medical College
Curated by ChEMBL
Peking Union Medical College
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Inhibition of human KDR/VEGFR2 using poly [Glu:Tyr] as substrate incubated for 30 mins followed by 33P-ATP addition and measured after 2 hrs by liqui...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Inhibition of human LYN using poly [Glu:Tyr] as substrate incubated for 30 mins followed by 33P-ATP addition and measured after 2 hrs by liquid scint...More data for this Ligand-Target Pair
TargetPlatelet-derived growth factor receptor alpha(Human)
Peking Union Medical College
Curated by ChEMBL
Peking Union Medical College
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Inhibition of human PDGFRa using poly [Glu:Tyr] as substrate incubated for 30 mins followed by 33P-ATP addition and measured after 2 hrs by liquid sc...More data for this Ligand-Target Pair
Affinity DataIC50: 22nMAssay Description:Inhibition of FGFR1 (unknown origin) using FAM-P22 peptide as substrate preincubated with compound for 10 mins followed by substrate addition and ATP...More data for this Ligand-Target Pair
TargetFibroblast growth factor receptor 2(Human)
Peking Union Medical College and Chinese Academy of Medical Sciences
Curated by ChEMBL
Peking Union Medical College and Chinese Academy of Medical Sciences
Curated by ChEMBL
Affinity DataIC50: 22nMAssay Description:Inhibition of FGFR2 (unknown origin) using FAM-P22 peptide as substrate preincubated with compound for 10 mins followed by substrate addition and ATP...More data for this Ligand-Target Pair
TargetFibroblast growth factor receptor 3(Human)
Peking Union Medical College and Chinese Academy of Medical Sciences
Curated by ChEMBL
Peking Union Medical College and Chinese Academy of Medical Sciences
Curated by ChEMBL
Affinity DataIC50: 52nMAssay Description:Inhibition of FGFR3 (unknown origin) using FAM-P22 peptide as substrate preincubated with compound for 10 mins followed by substrate addition and ATP...More data for this Ligand-Target Pair
TargetFibroblast growth factor receptor 4(Human)
Peking Union Medical College and Chinese Academy of Medical Sciences
Curated by ChEMBL
Peking Union Medical College and Chinese Academy of Medical Sciences
Curated by ChEMBL
Affinity DataIC50: 124nMAssay Description:Inhibition of FGFR4 (unknown origin) using FAM-P22 peptide as substrate preincubated with compound for 10 mins followed by substrate addition and ATP...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Peking Union Medical College
Curated by ChEMBL
Peking Union Medical College
Curated by ChEMBL
Affinity DataIC50: 1.11E+4nMAssay Description:Inhibition of hERGMore data for this Ligand-Target Pair