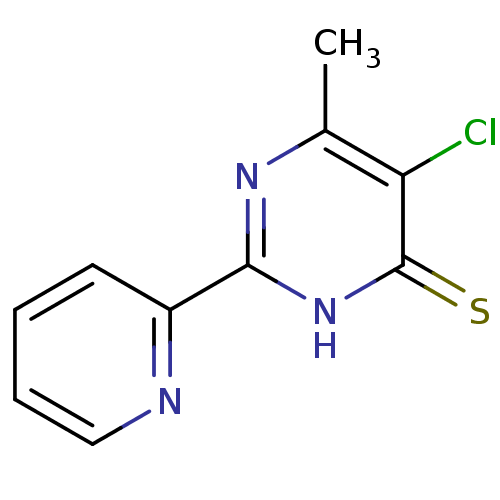
BDBM60271 5-chloranyl-6-methyl-2-pyridin-2-yl-1H-pyrimidine-4-thione::5-chloro-6-methyl-2-(2-pyridinyl)-1H-pyrimidine-4-thione::5-chloro-6-methyl-2-(2-pyridyl)-1H-pyrimidine-4-thione::5-chloro-6-methyl-2-(2-pyridyl)pyrimidine-4-thiol::5-chloro-6-methyl-2-pyridin-2-yl-1H-pyrimidine-4-thione::MLS000861172::SMR000459956::cid_2725954
SMILES Cc1nc([nH]c(=S)c1Cl)-c1ccccn1
InChI Key InChIKey=OMVHUYAQBLWYAA-UHFFFAOYSA-N
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 2 hits for monomerid = 60271
Found 2 hits for monomerid = 60271
Affinity DataEC50: 4.58E+3nMAssay Description:University of New Mexico Assay Overview: Assay Support: 1R03 MH086450-01 Project Title: Chemical Screen of TOR pathway GFP fusion proteins in S. ...More data for this Ligand-Target Pair
TargetKappa-type opioid receptor(Human)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 2.03E+4nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute (SBMRI, San Diego, C...More data for this Ligand-Target Pair