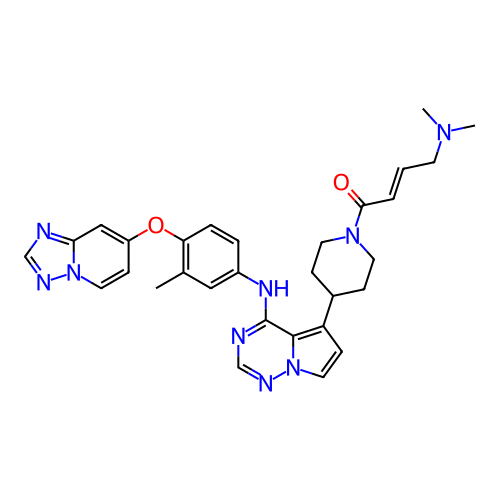
BDBM768387 (E)-1-(4-(4-((4- ([1,2,4]triazolo[1,5- a]pyridin-7-yloxy)-3- methylphenyl)amino) pyrrolo[2,1-f][1,2,4]triazin- 5-yl)piperidin-1-yl)-4- (dimethylamino)but-2- en-1-one::US20250276983, Example 21::US20250345339, Compound 5
SMILES Cc1cc(Nc2ncnn3ccc(C4CCN(C(=O)/C=C/CN(C)C)CC4)c23)ccc1Oc1ccn2ncnc2c1
InChI Key
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 8 hits for monomerid = 768387
Found 8 hits for monomerid = 768387
Affinity DataKi: 16nMAssay Description:The purpose of the wtERBB2, ERBB2-A775_G776insYVMA (ERBB2YVMA), wtEGFR biochemical assay was to evaluate the inhibition (% inhibition and IC50 values...More data for this Ligand-Target Pair
Affinity DataIC50: 200nMAssay Description:The purpose of the wtERBB2, ERBB2-A775_G776insYVMA (ERBB2YVMA), wtEGFR biochemical assay was to evaluate the inhibition (% inhibition and IC50 values...More data for this Ligand-Target Pair
Affinity DataIC50: 200nMAssay Description:The radioactive signal generated from the aliquot of buffer solution was proportional to the amount of radiolabeled substrate produced, and which is ...More data for this Ligand-Target Pair
Affinity DataIC50: 200nMAssay Description:The purpose of the wtERBB2, ERBB2-A775_G776insYVMA (ERBB2YVMA), wtEGFR biochemical assay was to evaluate the inhibition (% inhibition and IC50 values...More data for this Ligand-Target Pair
Affinity DataIC50: 200nMAssay Description:The radioactive signal generated from the aliquot of buffer solution was proportional to the amount of radiolabeled substrate produced, and which is ...More data for this Ligand-Target Pair
Affinity DataKi: 290nMAssay Description:The purpose of the wtERBB2, ERBB2-A775_G776insYVMA (ERBB2YVMA), wtEGFR biochemical assay was to evaluate the inhibition (% inhibition and IC50 values...More data for this Ligand-Target Pair
Affinity DataIC50: 350nMAssay Description:The purpose of the wtERBB2, ERBB2-A775_G776insYVMA (ERBB2YVMA), wtEGFR biochemical assay was to evaluate the inhibition (% inhibition and IC50 values...More data for this Ligand-Target Pair
Affinity DataIC50: 350nMAssay Description:The radioactive signal generated from the aliquot of buffer solution was proportional to the amount of radiolabeled substrate produced, and which is ...More data for this Ligand-Target Pair