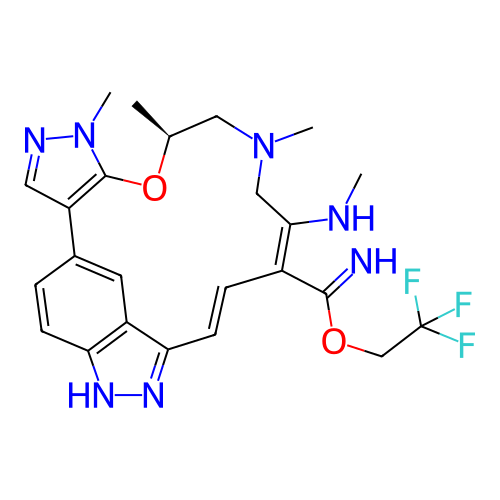
BDBM790046 (10S,17E)-8,10,12,14-tetramethyl-16- (2,2,2-trifluoroethoxy)-2,10,11,12,13,14- hexahydro-8H-3,5-ethenotripyrazolo[3,4- f:3',4'-j:4$#8243;,3$#8243;- n][1,4]oxazacyclopentadecine::US20250353864, Example 162
SMILES CN/C1=C(C(=N)OCC(F)(F)F)/C=C/c2n[nH]c3ccc(cc23)-c2cnn(C)c2O[C@@H](C)CN(C)C1
InChI Key InChIKey=WRNLVCNAJKXXBB-UHFFFAOYSA-N
Data 11 IC50
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 11 hits for monomerid = 790046
Found 11 hits for monomerid = 790046
Affinity DataIC50: 0.0700nMAssay Description:Table 3: Kinase protein and substrate were pre-diluted in the HEPES assay buffer (100 mM HEPES, pH 7.5, 0.01% Triton X-100, 0.1% BSA, 5 mM MgCl2, 1 m...More data for this Ligand-Target Pair
Affinity DataIC50: 0.110nMAssay Description:Table 3: Kinase protein and substrate were pre-diluted in the HEPES assay buffer (100 mM HEPES, pH 7.5, 0.01% Triton X-100, 0.1% BSA, 5 mM MgCl2, 1 m...More data for this Ligand-Target Pair
Affinity DataIC50: 0.25nMAssay Description:Table 3: Kinase protein and substrate were pre-diluted in the HEPES assay buffer (100 mM HEPES, pH 7.5, 0.01% Triton X-100, 0.1% BSA, 5 mM MgCl2, 1 m...More data for this Ligand-Target Pair
Affinity DataIC50: 0.900nMAssay Description:Table 1: The inhibitory activities against EGFR WT, EGFR (D770_N771insNPG), EGFR Δ746-750 mutant, EGFR Δ746-750/C797S mutant, EGFR L858R mutant, EGFR...More data for this Ligand-Target Pair
Affinity DataIC50: 1.40nMAssay Description:Table 1: The inhibitory activities against EGFR WT, EGFR (D770_N771insNPG), EGFR Δ746-750 mutant, EGFR Δ746-750/C797S mutant, EGFR L858R mutant, EGFR...More data for this Ligand-Target Pair
Affinity DataIC50: 30.6nMAssay Description:Table 1: The inhibitory activities against EGFR WT, EGFR (D770_N771insNPG), EGFR Δ746-750 mutant, EGFR Δ746-750/C797S mutant, EGFR L858R mutant, EGFR...More data for this Ligand-Target Pair
TargetEpidermal growth factor receptor [1-745,751-1210,C797S](Human)
Blossomhill Therapeutics
US Patent
Blossomhill Therapeutics
US Patent
Affinity DataIC50: 33.5nMAssay Description:Table 1: The inhibitory activities against EGFR WT, EGFR (D770_N771insNPG), EGFR Δ746-750 mutant, EGFR Δ746-750/C797S mutant, EGFR L858R mutant, EGFR...More data for this Ligand-Target Pair
Affinity DataIC50: 47.4nMAssay Description:Table 1: The inhibitory activities against EGFR WT, EGFR (D770_N771insNPG), EGFR Δ746-750 mutant, EGFR Δ746-750/C797S mutant, EGFR L858R mutant, EGFR...More data for this Ligand-Target Pair
Affinity DataIC50: 238nMAssay Description:Table 1: The inhibitory activities against EGFR WT, EGFR (D770_N771insNPG), EGFR Δ746-750 mutant, EGFR Δ746-750/C797S mutant, EGFR L858R mutant, EGFR...More data for this Ligand-Target Pair
TargetEpidermal growth factor receptor [1-770,'NPG',771-1210](Human)
Blossomhill Therapeutics
US Patent
Blossomhill Therapeutics
US Patent
Affinity DataIC50: 523nMAssay Description:Table 1: The inhibitory activities against EGFR WT, EGFR (D770_N771insNPG), EGFR Δ746-750 mutant, EGFR Δ746-750/C797S mutant, EGFR L858R mutant, EGFR...More data for this Ligand-Target Pair
Affinity DataIC50: 784nMAssay Description:Table 3: Kinase protein and substrate were pre-diluted in the HEPES assay buffer (100 mM HEPES, pH 7.5, 0.01% Triton X-100, 0.1% BSA, 5 mM MgCl2, 1 m...More data for this Ligand-Target Pair