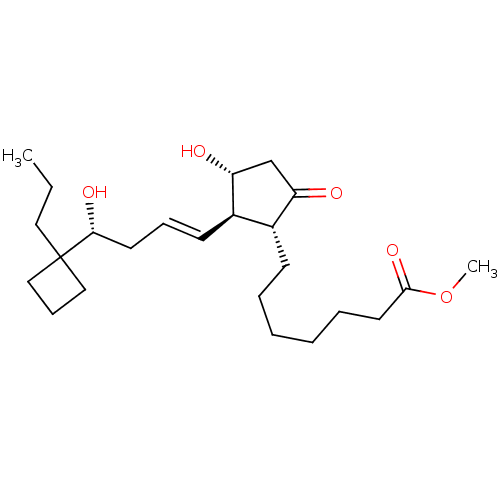
BDBM85602 (R)-BUTAPROST::Butaprost (Free Acid)::CAS_69648-38-0
SMILES CCCC1(CCC1)[C@H](O)C\C=C\[C@H]1[C@H](O)CC(=O)[C@@H]1CCCCCCC(=O)OC
InChI Key InChIKey=XRISENIKJUKIHD-UHFFFAOYSA-N
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 14 hits for monomerid = 85602
Found 14 hits for monomerid = 85602
Affinity DataEC50: 33nMAssay Description:Agonist activity at EP2 receptor (unknown origin) by functional assayMore data for this Ligand-Target Pair
Affinity DataIC50: 37nMAssay Description:Agonist activity at prostanoid IP receptor (unknown origin) by functional assayMore data for this Ligand-Target Pair
TargetProstaglandin E2 receptor EP3 subtype(Human)
Merck Frosst Centre For Therapeutic Research
Curated by PDSP Ki Database
Merck Frosst Centre For Therapeutic Research
Curated by PDSP Ki Database
Affinity DataKi: 2.40E+3nMAssay Description:Binding affinity to EP2 receptor (unknown origin) by competitive binding assayMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:Binding affinity to EP1 receptor (unknown origin)More data for this Ligand-Target Pair
TargetProstaglandin F2-alpha receptor(Human)
Merck Frosst Centre For Therapeutic Research
Curated by PDSP Ki Database
Merck Frosst Centre For Therapeutic Research
Curated by PDSP Ki Database
Affinity DataKi: >1.00E+4nMAssay Description:Binding affinity to EP4 receptor (unknown origin)More data for this Ligand-Target Pair
TargetSolute carrier organic anion transporter family member 2A1(Human)
Merck Frosst Centre For Therapeutic Research
Curated by PDSP Ki Database
Merck Frosst Centre For Therapeutic Research
Curated by PDSP Ki Database
TargetHematopoietic prostaglandin D synthase(Human)
Merck Frosst Centre For Therapeutic Research
Curated by PDSP Ki Database
Merck Frosst Centre For Therapeutic Research
Curated by PDSP Ki Database
Affinity DataKi: >1.00E+4nMAssay Description:Binding affinity to prostanoid IP receptor (unknown origin)More data for this Ligand-Target Pair
TargetProstaglandin E2 receptor EP3 subtype(Human)
Merck Frosst Centre For Therapeutic Research
Curated by PDSP Ki Database
Merck Frosst Centre For Therapeutic Research
Curated by PDSP Ki Database
Affinity DataKi: >1.00E+4nMAssay Description:Binding affinity to EP3 receptor (unknown origin)More data for this Ligand-Target Pair