BDBM50344263 (+)-1-methyl-16-azatetracyclo[7.6.1.02,7.010,15]hexadeca-2(7),3,5,10(15),11,13-hexaene::(+)-MK-801::(+)MK-801::(+/-) MK-8011-methyl-(9R,1R)-16-azatetracyclo[7.6.1.02,7.010,15]hexadeca-2(7),3,5,10(15),11,13-hexaene::(+/-)-1-methyl-16-azatetracyclo[7.6.1.02,7.010,15]hexadeca-2(7),3,5,10(15),11,13-hexaene::(+/-)-MK801::(-)-1-methyl-16-azatetracyclo[7.6.1.02,7.010,15]hexadeca-2(7),3,5,10(15),11,13-hexaene::(-)-MK801::(1S,9R)-1-methyl-16-azatetracyclo[7.6.1.0^{2,7}.0^{10,15}]hexadeca-2(7),3,5,10(15),11,13-hexaene::(1S,9R)-1-methyl-16-azatetracyclo[7.6.1.0^{2,7}.0^{10,15}]hexadeca-2,4,6,10(15),11,13-hexaene::(5S,10R)-(+)-5-methyl-10,11-dihydro-5Hdibenzo[a,d]cyclohepten-5,10-imine::(5S,10S)-(+)-5-methyl-10,11-dihydro-5Hdibenzo[a,d]cyclohepten-5,10-imine::(Dizocilpine)1-methyl-16-azatetracyclo[7.6.1.02,7.010,15]hexadeca-2(7),3,5,10(15),11,13-hexaene::(MK-801)1-methyl-16-azatetracyclo[7.6.1.02,7.010,15]hexadeca-2(7),3,5,10(15),11,13-hexaene::1-methyl-16-azatetracyclo[7.6.1.02,7.010,15]hexadeca-2(7),3,5,10(15),11,13-hexaene::1-methyl-16-azatetracyclo[7.6.1.02,7.010,15]hexadeca-2(7),3,5,10(15),11,13-hexaene(MK-801)::1-methyl-16-azatetracyclo[7.6.1.02,7.010,15]hexadeca-2(7),3,5,10,12,14-hexaene::1-methyl-16-azatetracyclo[7.6.1.02,7.010,15]hexadeca-2,4,6,10(15),11,13-hexaene::10,11-Dihydro-5-methyl-5H-dibenzo[a,d]cyclohepten-5,10-imine.(MK-801)::CHEMBL284237::MK-801::MK-801 (Dizocilpine)::dizocilpine
SMILES C[C@@]12c3ccccc3C[C@@H](N1)c4c2cccc4
InChI Key InChIKey=LBOJYSIDWZQNJS-UHFFFAOYSA-N
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 12 hits for monomerid = 50344263
Found 12 hits for monomerid = 50344263
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
University of Illinois At Chicago
Curated by ChEMBL
University of Illinois At Chicago
Curated by ChEMBL
National Institute On Drug Abuse
Curated by ChEMBL
National Institute On Drug Abuse
Curated by ChEMBL
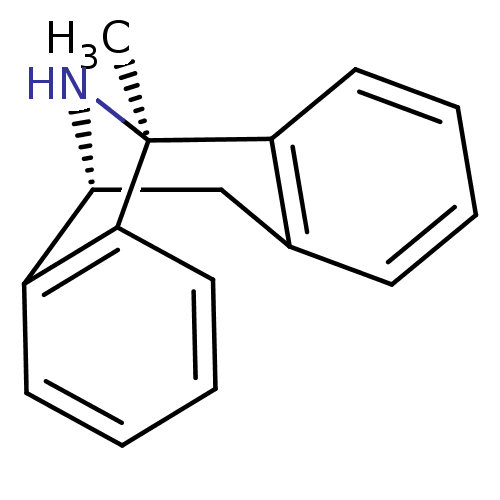
 3D Structure (crystal)
3D Structure (crystal)