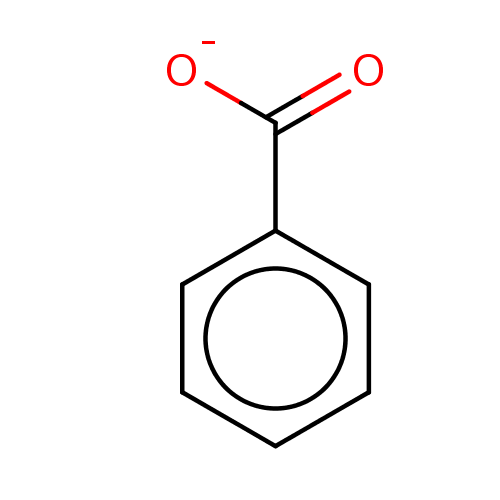
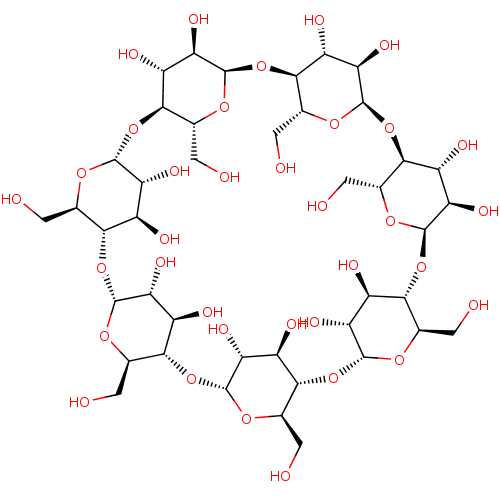
BDBM36181 SODIUM BENZOATE::benzoate::benzoic acid::benzoic acid-d5
SMILES [O-]C(=O)c1ccccc1
InChI Key InChIKey=WPYMKLBDIGXBTP-UHFFFAOYSA-M
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 22 hits for monomerid = 36181
Found 22 hits for monomerid = 36181
Affinity DataKi: 2.00E+3nMAssay Description:Competitive inhibition of human recombinant DAAO by Michaelis-Menten plot analysis in presence of D-serineMore data for this Ligand-Target Pair
Affinity DataIC50: 2.09E+3nMAssay Description:Competitive inhibition of human recombinant DAAO after 1 hr by coupled enzyme assay in presence of D-serineMore data for this Ligand-Target Pair
Affinity DataKd: 2.20E+3nMAssay Description:Binding affinity to human recombinant DAAO at 492 nm spectral modification by spectrophotometric analysis in presence of FADMore data for this Ligand-Target Pair
Affinity DataKd: 2.30E+3nMAssay Description:Binding affinity to human recombinant DAAO by stopped flow spectrophotometric analysis in presence of FADMore data for this Ligand-Target Pair
Affinity DataKi: 7.00E+3nMAssay Description:Inhibition of human recombinant DAO expressed in Escherichia coli BL21(DE3) by oxygraphic assayMore data for this Ligand-Target Pair
Affinity DataKd: 9.00E+3nMAssay Description:Binding affinity to human recombinant DAAO at 442 nm spectral modification by spectrophotometric analysis in presence of FADMore data for this Ligand-Target Pair
Affinity DataEC50: >1.00E+4nMAssay Description:Positive allosteric modulation of human mGluR5a transfected in HEK293 cells by calcium mobilization assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.09E+4nMAssay Description:Inhibition of human recombinant N-terminal His-tagged DAO expressed in Escherichia coli BL21(DE3) using D-serine as substrate by Lineweaver-Burk plot...More data for this Ligand-Target Pair
Affinity DataKi: 1.38E+4nMAssay Description:Inhibition of mouse Oat6-mediated [3H]ES uptake in Xenopus oocytes after 1 hrMore data for this Ligand-Target Pair
Affinity DataKi: 2.44E+4nMAssay Description:Inhibition of Candida albicans Nce103More data for this Ligand-Target Pair
Affinity DataKi: 3.00E+4nMAssay Description:Inhibition of human carbonic anhydrase2More data for this Ligand-Target Pair
Affinity DataKi: 3.00E+4nMAssay Description:Binding affinity against human cytosolic Carbonic anhydrase IIMore data for this Ligand-Target Pair
Affinity DataIC50: 4.69E+4nMAssay Description:Inhibition of human recombinant N-terminal His-tagged DAO expressed in Escherichia coli BL21(DE3) using D-serine as substrate by colorimetric assayMore data for this Ligand-Target Pair
TargetCarbonic anhydrase [110-330](Coastal plain yellowtops)
Istituto Di Biostrutture E Bioimmagini-Cnr
Istituto Di Biostrutture E Bioimmagini-Cnr
Affinity DataKi: 6.45E+4nM ΔG°: -5.62kcal/molepH: 8.3 T: 2°CAssay Description:Inhibition constants of carboxylate inhibitors against F. bidentis CA I were determined by a stopped flow CO2 hydration assay at 20 °C.More data for this Ligand-Target Pair
Affinity DataIC50: 6.57E+4nMAssay Description:Inhibition of human recombinant N-terminal His-tagged DAO expressed in Escherichia coli BL21(DE3) using D-alanine as substrate by colorimetric assayMore data for this Ligand-Target Pair
Affinity DataKi: 7.47E+4nMAssay Description:Binding affinity against human cytosolic Carbonic anhydrase IVMore data for this Ligand-Target Pair
Affinity DataKi: 2.53E+5nMAssay Description:Inhibition of mouse Oat1-mediated [3H]PAH uptake in Xenopus oocytes after 1 hrMore data for this Ligand-Target Pair
Affinity DataKi: 7.30E+5nMAssay Description:Inhibition of human cytosolic carbonic anhydrase 1 by stopped flow CO2 hydration assayMore data for this Ligand-Target Pair
Affinity DataKi: 7.01E+6nMAssay Description:Binding affinity against human cytosolic Carbonic anhydrase VMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+7nMAssay Description:Inhibition of human recombinant N-terminal His-tagged DDO expressed in Escherichia coli BL21(DE3) using D-aspartate as substrate by colorimetric assa...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+7nMAssay Description:Inhibition of human recombinant N-terminal His-tagged serine racemase expressed in Escherichia coli BL21(DE3) using L-serine as substrate after 10 mi...More data for this Ligand-Target Pair
Affinity DataKi: >1.50E+8nMAssay Description:Binding affinity against human cytosolic Carbonic anhydrase IXMore data for this Ligand-Target Pair
Activity Spreadsheet -- ITC Data from BindingDB
 Found 5 hits for monomerid = 36181
Found 5 hits for monomerid = 36181
Host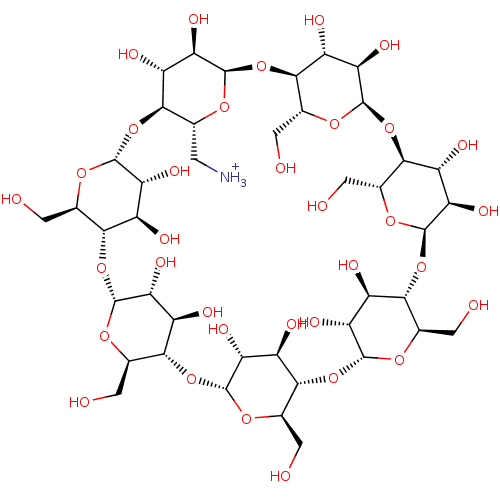 BDBM36127(6-NH2-beta-cyclodextrin | mono(6-amino-6-deoxy)-be...)
BDBM36127(6-NH2-beta-cyclodextrin | mono(6-amino-6-deoxy)-be...)
Japan Science and Technology Agency
 BDBM36127(6-NH2-beta-cyclodextrin | mono(6-amino-6-deoxy)-be...)
BDBM36127(6-NH2-beta-cyclodextrin | mono(6-amino-6-deoxy)-be...)Japan Science and Technology Agency
ITC DataΔG°: -2.51kcal/mole −TΔS°: -0.669kcal/mole ΔH°: -1.83kcal/mole logk: 69
pH: 6.9 T: 25.00°C
pH: 6.9 T: 25.00°C
Host BDBM36127(6-NH2-beta-cyclodextrin | mono(6-amino-6-deoxy)-be...)
BDBM36127(6-NH2-beta-cyclodextrin | mono(6-amino-6-deoxy)-be...)
Japan Science and Technology Agency
 BDBM36127(6-NH2-beta-cyclodextrin | mono(6-amino-6-deoxy)-be...)
BDBM36127(6-NH2-beta-cyclodextrin | mono(6-amino-6-deoxy)-be...)Japan Science and Technology Agency
ITC DataΔG°: -2.58kcal/mole −TΔS°: -0.762kcal/mole ΔH°: -1.83kcal/mole logk: 78
pH: 6.9 T: 25.00°C
pH: 6.9 T: 25.00°C
Host BDBM36127(6-NH2-beta-cyclodextrin | mono(6-amino-6-deoxy)-be...)
BDBM36127(6-NH2-beta-cyclodextrin | mono(6-amino-6-deoxy)-be...)
Japan Science and Technology Agency
 BDBM36127(6-NH2-beta-cyclodextrin | mono(6-amino-6-deoxy)-be...)
BDBM36127(6-NH2-beta-cyclodextrin | mono(6-amino-6-deoxy)-be...)Japan Science and Technology Agency
ITC DataΔG°: -2.46kcal/mole −TΔS°: -0.577kcal/mole ΔH°: -1.89kcal/mole logk: 64
pH: 6.9 T: 25.00°C
pH: 6.9 T: 25.00°C
Host BDBM36127(6-NH2-beta-cyclodextrin | mono(6-amino-6-deoxy)-be...)
BDBM36127(6-NH2-beta-cyclodextrin | mono(6-amino-6-deoxy)-be...)
Japan Science and Technology Agency
 BDBM36127(6-NH2-beta-cyclodextrin | mono(6-amino-6-deoxy)-be...)
BDBM36127(6-NH2-beta-cyclodextrin | mono(6-amino-6-deoxy)-be...)Japan Science and Technology Agency
ITC DataΔG°: -2.55kcal/mole −TΔS°: -0.741kcal/mole ΔH°: -1.82kcal/mole logk: 74
pH: 6.9 T: 25.00°C
pH: 6.9 T: 25.00°C
ITC DataΔG°: -1.64kcal/mole −TΔS°: 0.869kcal/mole ΔH°: -2.51kcal/mole logk: 15.9
pH: 6.9 T: 25.00°C
pH: 6.9 T: 25.00°C