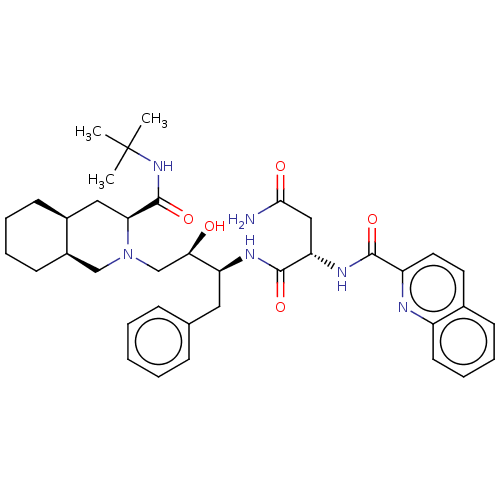
BDBM519 (2S)-N-[(2S,3R)-4-[(3S,4aS,8aS)-3-(tert-butylcarbamoyl)-decahydroisoquinolin-2-yl]-3-hydroxy-1-phenylbutan-2-yl]-2-(quinolin-2-ylformamido)butanediamide::CHEMBL114::Fortovase::Invirase::Ro 31-8959::SQV::Saquinavir
SMILES CC(C)(C)NC(=O)[C@@H]1C[C@@H]2CCCC[C@@H]2CN1C[C@@H](O)[C@H](Cc1ccccc1)NC(=O)[C@H](CC(N)=O)NC(=O)c1ccc2ccccc2n1
InChI Key InChIKey=QWAXKHKRTORLEM-UHFFFAOYSA-N
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 70 hits for monomerid = 519
Found 70 hits for monomerid = 519
TargetDimer of Gag-Pol polyprotein [501-599,Q508K,L534I,L564I,C568A,C596A](Human immunodeficiency virus type 1)
Glaxosmithkline
Glaxosmithkline
Affinity DataKi: 0.0400nM ΔG°: -14.2kcal/molepH: 6.4 T: 2°CAssay Description:The Ki values were determined using fluorogenic substrate, 2-(aminobenzoyl)-Thr-Ile-Nle-Phe(p-NO2)-Gln-ArgNH2. A standard curve relating changes in f...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataKi: 0.0620nMAssay Description:Dissociation constant obtained by inhibition of Wild-type proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 0.0700nMAssay Description:Inhibition assay using HIV protease and Sulfonamide compounds.More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataKi: 0.120nMAssay Description:Binding affinity to HIV-1 proteaseMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataKi: 0.120nMAssay Description:Binding activity against HIV-1 ProteaseMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataKi: 0.120nMAssay Description:The compound was tested for binding activity against HIV-1 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 0.150nMpH: 5.5Assay Description:Protease inhibition fluorescence-based assay using cyclic ureas to inhibit HIV-protease.More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataKi: 0.190nMAssay Description:Inhibitory activity against HIV-1 proteaseMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 0.200nMAssay Description:In vitro inhibition HIV-1 IIIB protease.More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Merck Research Laboratories
Merck Research Laboratories
Affinity DataIC50: 0.230nMpH: 5.5 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. The products were analyzed by usi...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 0.230nMAssay Description:Inhibitory activity against HIV-1 protease.More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataKi: 0.230nMAssay Description:Inhibition constant for human immunodeficiency virus type 1 proteaseMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataKi: 0.230nM Kd: 0.315nM Koff: 8.17E+5s-1Assay Description:Association rate constant for the interaction between inhibitor and HIV-1 proteaseMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 0.230nMAssay Description:Inhibition of HIV-1 protease was determined in vitroMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataKi: 0.25nMAssay Description:Binding affinity to inhibit the purified wild-type HIV-1 ProteaseMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataKi: 0.25nMAssay Description:Affinity against HIV proteaseMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataKi: 0.25nMAssay Description:Inhibitory activity against HIV-1 proteaseMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataKd: 0.310nMAssay Description:Binding affinity for human immunodeficiency virus type 1 proteaseMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 0.340nMAssay Description:In vitro inhibition of HIV-Protease, using a peptide hydrolysis assayMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 0.400nMAssay Description:Binding activity against HIV-1 ProteaseMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 0.400nMAssay Description:Inhibition of wild type HIV1 recombinant aspartic proteaseMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 0.400nMAssay Description:Binding activity against HIV-1 ProteaseMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 0.400nMAssay Description:Inhibitory potency against HIV-1 proteaseMore data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [482-580,I502K,I528M,T553A,V564F](Human immunodeficiency virus type 1)
Johnson & Johnson Pharmaceutical
Johnson & Johnson Pharmaceutical
Affinity DataKi: 0.430nMAssay Description:The inhibition constants Ki were determined by monitoring the inhibition of hydrolysis of the chromogenic substrate using a Hewlett-Packard 8452A spe...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataKi: 0.452nMAssay Description:Inhibitory activity against P2 site in HIV protease.More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587,Q496K](Human immunodeficiency virus type 1)
The Johns Hopkins University
The Johns Hopkins University
Affinity DataKi: 0.463nM ΔG°: -12.7kcal/molepH: 5.0 T: 2°CAssay Description:Protease activity was determined by following the increase in fluorescence with hydrolysis of the fluorogenic substrate Arg-Glu(EDANS)-Ser-Gln-Asn-Ty...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,V583F](Human immunodeficiency virus type 1)
Johnson & Johnson Pharmaceutical
Johnson & Johnson Pharmaceutical
Affinity DataKi: 0.520nM ΔG°: -13.2kcal/molepH: 4.7 T: 2°CAssay Description:The inhibition constants Ki were determined by monitoring the inhibition of hydrolysis of the chromogenic substrate using a Hewlett-Packard 8452A spe...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Merck Research Laboratories
Merck Research Laboratories
Affinity DataKi: 0.900nM ΔG°: -12.8kcal/molepH: 5.6 T: 2°CAssay Description:Protease activity was measured by continuous chromogenic assay. The chromogenic peptide His-Lys-Ala-Arg-Val-Leu- (p-NO2-Phe)-Glu-Ala-Nleu-Ser (Bache...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Merck Research Laboratories
Merck Research Laboratories
Affinity DataKi: 0.900nM ΔG°: -12.8kcal/molepH: 5.6 T: 2°CAssay Description:Protease activity was measured by continuous chromogenic assay. The chromogenic peptide His-Lys-Ala-Arg-Val-Leu-(p-NO2-Phe)-Glu-Ala-Nleu-Ser was use...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 1nMAssay Description:Inhibitory concentration required against HIV-1 proteaseMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 1nMAssay Description:The assay involves the use of a HIV-1 protease peptide substrate which has been modified to contain a biotin moiety on one side and a fluorescent rep...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [491-589,Q498K,D521N,V555I,N579D](Human immunodeficiency virus type 1)
University of Massachusetts Medical School
University of Massachusetts Medical School
Affinity DataKi: 1.03nMAssay Description:HIV-1 protease inhibitor activities were determined by the fluorescence resonance energy transfer (FRET) method. The energy transfer donor (EDANS) an...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599](Human immunodeficiency virus type 1)
Johnson & Johnson Pharmaceutical
Johnson & Johnson Pharmaceutical
Affinity DataKi: 1.30nM ΔG°: -12.6kcal/molepH: 4.7 T: 2°CAssay Description:The inhibition constants Ki were determined by monitoring the inhibition of hydrolysis of the chromogenic substrate using a Hewlett-Packard 8452A spe...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataKi: 1.40nMAssay Description:Compound was evaluated for binding affinity against HIV proteaseMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 1.70nMAssay Description:Inhibitory activity against human immunodeficiency virus protease (HIVP) using protease inhibition assayMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 1.80nMAssay Description:Inhibitory activity against HIV-1 proteaseMore data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [482-580,I502K,I528M,T553A](Human immunodeficiency virus type 1)
Johnson & Johnson Pharmaceutical
Johnson & Johnson Pharmaceutical
Affinity DataKi: 1.80nM ΔG°: -12.4kcal/molepH: 4.7 T: 2°CAssay Description:The inhibition constants Ki were determined by monitoring the inhibition of hydrolysis of the chromogenic substrate using a Hewlett-Packard 8452A spe...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587,Q496K,I573V](Human immunodeficiency virus type 1)
The Johns Hopkins University
The Johns Hopkins University
Affinity DataKi: 2nM ΔG°: -11.9kcal/molepH: 5.0 T: 2°CAssay Description:Protease activity was determined by following the increase in fluorescence with hydrolysis of the fluorogenic substrate Arg-Glu(EDANS)-Ser-Gln-Asn-Ty...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataKi: 3.70nMAssay Description:Dissociation constant obtained by inhibition of mutant HIV-protease (K-60)More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Merck Research Laboratories
Merck Research Laboratories
Affinity DataIC50: 8.40nMpH: 5.5 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Cleavage products and substrate w...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 11nMAssay Description:In vitro inhibitory activity against HIV proteinaseMore data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Merck Research Laboratories
Merck Research Laboratories
Affinity DataIC50: 11.2nMT: 2°CAssay Description:IC50 values were obtained by assaying the enzyme against the synthetic substrate peptide KQGTVSFNFPQIT, which was tritiated at the proline residue, a...More data for this Ligand-Target Pair
Affinity DataIC50: 13.1nMpH: 6.5Assay Description:Enzyme inhibition assay using cyclic ureas to inhibit GAG polyprotein.More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataKi: 15nMAssay Description:Dissociation constant obtained by inhibition of mutant HIV-protease (A-44)More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,Q508K,L534I,L564I,C568A,C596A](Human immunodeficiency virus type 1)
Glaxosmithkline
Glaxosmithkline
Affinity DataIC50: 16nMpH: 6.4 T: 2°CAssay Description:The Ki values were determined by substrate cleavage assay using fluorogenic substrate, 2-(aminobenzoyl)-Thr-Ile-Nle-Phe(p-NO2)-Gln-Arg-NH2. A standar...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [490-588,L523I,E525D,M526I,I544V,L553H,H559K,V572F,L579M](Human immunodeficiency virus type 1)
Johnson & Johnson Pharmaceutical
Johnson & Johnson Pharmaceutical
Affinity DataKi: 24nMAssay Description:The inhibition constants Ki were determined by monitoring the inhibition of hydrolysis of the chromogenic substrate using a Hewlett-Packard 8452A spe...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 30nMAssay Description:Concentration required for inhibitory activity against HIV-1 ProteaseMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 45nMAssay Description:Inhibitory activity against HIV-1 protease using scintillation proximity assay (SPA assay)More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [490-588,L523I,E525D,M526I,I544V,L553H,H559K,L579M](Human immunodeficiency virus type 1)
Johnson & Johnson Pharmaceutical
Johnson & Johnson Pharmaceutical
Affinity DataKi: 48nMAssay Description:The inhibition constants Ki were determined by monitoring the inhibition of hydrolysis of the chromogenic substrate using a Hewlett-Packard 8452A spe...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [491-589,Q498K,L501I,V555I,A562V,G564S,I575V,L581M](Human immunodeficiency virus type 1)
University of Massachusetts Medical School
University of Massachusetts Medical School
Affinity DataKi: 78.4nMAssay Description:HIV-1 protease inhibitor activities were determined by the fluorescence resonance energy transfer (FRET) method. The energy transfer donor (EDANS) an...More data for this Ligand-Target Pair
Activity Spreadsheet -- ITC Data from BindingDB
 Found 18 hits for monomerid = 519
Found 18 hits for monomerid = 519
ITC DataΔG°: -11.8kcal/mole −TΔS°: -14.0kcal/mole ΔH°: 2.20kcal/mole
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
CellHIV-1 Protease Mutant (V82F/I84V)(Human immunodeficiency virus type 1)
The Johns Hopkins University
The Johns Hopkins University
ITC DataΔG°: -10.4kcal/mole −TΔS°: -13.9kcal/mole ΔH°: 3.49kcal/mole
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
ITC DataΔG°: -13.0kcal/mole −TΔS°: -14.2kcal/mole ΔH°: 1.20kcal/mole logk: 3.60E+9
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
ITC DataΔG°: -8.49kcal/mole −TΔS°: -18.9kcal/mole ΔH°: 10.4kcal/mole logk: 1.80E+6
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
ITC DataΔG°: -9.98kcal/mole −TΔS°: -18.5kcal/mole ΔH°: 8.49kcal/mole logk: 2.30E+7
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
CellHIV-1 Protease Mutant (V82A/I84V)(Human immunodeficiency virus type 1)
The Johns Hopkins University
The Johns Hopkins University
ITC DataΔG°: -11.8kcal/mole −TΔS°: -15.5kcal/mole ΔH°: 3.69kcal/mole logk: 4.60E+8
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
CellHIV-1 Protease Mutant (M46I/I54V)(Human immunodeficiency virus type 1)
The Johns Hopkins University
The Johns Hopkins University
ITC DataΔG°: -11.9kcal/mole −TΔS°: -17.0kcal/mole ΔH°: 5.09kcal/mole logk: 5.10E+8
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
CellHIV-1 Protease Mutant (L10I/L90M)(Human immunodeficiency virus type 1)
The Johns Hopkins University
The Johns Hopkins University
ITC DataΔG°: -12.0kcal/mole −TΔS°: -15.6kcal/mole ΔH°: 3.59kcal/mole logk: 5.90E+8
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
ITC DataΔG°: -12.8kcal/mole −TΔS°: -14.7kcal/mole ΔH°: 1.90kcal/mole logk: 2.50E+9
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
ITC DataΔG°: -12.4kcal/mole −TΔS°: -14.5kcal/mole ΔH°: 2.10kcal/mole logk: 1.29E+9
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
ITC DataΔG°: -12.2kcal/mole −TΔS°: -14.4kcal/mole ΔH°: 2.20kcal/mole logk: 9.01E+8
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
CellHIV-1 Protease B Subtype Mutant (V82F/I84V)(Human immunodeficiency virus type 1)
The Johns Hopkins University
The Johns Hopkins University
ITC DataΔG°: -11.0kcal/mole −TΔS°: -14.3kcal/mole ΔH°: 3.29kcal/mole logk: 1.20E+8
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
CellHIV-1 Protease C Subtype Mutant (V82F/I84V)(Human immunodeficiency virus type 1)
The Johns Hopkins University
The Johns Hopkins University
ITC DataΔG°: -10.6kcal/mole −TΔS°: -13.6kcal/mole ΔH°: 3.00kcal/mole logk: 6.21E+7
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
CellHIV-1 Protease A Subtype Mutant (V82F/I84V)(Human immunodeficiency virus type 1)
The Johns Hopkins University
The Johns Hopkins University
ITC DataΔG°: -10.4kcal/mole −TΔS°: -13.3kcal/mole ΔH°: 2.90kcal/mole logk: 4.17E+7
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
ITC DataΔG°: -7.67kcal/mole −TΔS°: -17.3kcal/mole ΔH°: 9.58kcal/mole logk: 4.29E+5
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
ITC DataΔG°: -12.0kcal/mole −TΔS°: -18.0kcal/mole ΔH°: 5.99kcal/mole logk: 6.25E+8
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
ITC DataΔG°: -8.69kcal/mole −TΔS°: -15.1kcal/mole ΔH°: 6.39kcal/mole logk: 2.38E+6
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
ITC DataΔG°: -12.8kcal/mole −TΔS°: -14.7kcal/mole ΔH°: 1.90kcal/mole logk: 2.50E+9
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C