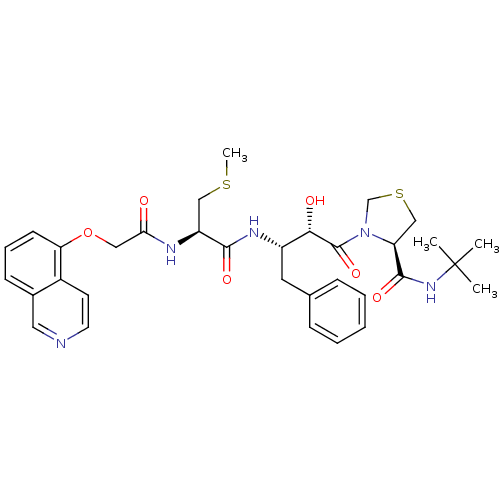
BDBM579 (4R)-N-tert-butyl-3-[(2S,3S)-2-hydroxy-3-[(2R)-2-[2-(isoquinolin-5-yloxy)acetamido]-3-(methylsulfanyl)propanamido]-4-phenylbutanoyl]-1,3-thiazolidine-4-carboxamide::CHEMBL414640::KNI-272::Kynostatin 272::iQoa-Mta-Apns-Thz-NH-tBu
SMILES CSC[C@H](NC(=O)COc1cccc2cnccc12)C(=O)N[C@@H](Cc1ccccc1)[C@H](O)C(=O)N1CSC[C@H]1C(=O)NC(C)(C)C
InChI Key InChIKey=NJBBLOIWMSYVCQ-UHFFFAOYSA-N
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 13 hits for monomerid = 579
Found 13 hits for monomerid = 579
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
National Cancer Institute-Bethesda
National Cancer Institute-Bethesda
Affinity DataKi: 0.00550nM ΔG°: -16.0kcal/molepH: 6.2 T: 2°CAssay Description:Inhibition constants were determined by a fluorometric assay with the fluorogenic substrate Arg-Glu(EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Lys(DABCYL)-Ar...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,A572V](Human immunodeficiency virus type 1)
University of Florida College of Medicine
University of Florida College of Medicine
Affinity DataKi: 0.370nM ΔG°: -13.4kcal/molepH: 4.7 T: 2°CAssay Description:The inhibition constant, Ki was determined by monitoring the competitive inhibition of the hydrolysis of the chromogenic substrate.More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,M537I,A572V](Human immunodeficiency virus type 1)
University of Florida College of Medicine
University of Florida College of Medicine
Affinity DataKi: 0.380nM ΔG°: -13.4kcal/molepH: 4.7 T: 2°CAssay Description:The inhibition constant, Ki was determined by monitoring the competitive inhibition of the hydrolysis of the chromogenic substrate.More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599](Human immunodeficiency virus type 1)
University of Florida College of Medicine
University of Florida College of Medicine
Affinity DataKi: 0.580nM ΔG°: -13.1kcal/molepH: 4.7 T: 2°CAssay Description:The inhibition constant, Ki was determined by monitoring the competitive inhibition of the hydrolysis of the chromogenic substrate.More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,M537I](Human immunodeficiency virus type 1)
University of Florida College of Medicine
University of Florida College of Medicine
Affinity DataKi: 0.600nM ΔG°: -13.1kcal/molepH: 4.7 T: 2°CAssay Description:The inhibition constant, Ki was determined by monitoring the competitive inhibition of the hydrolysis of the chromogenic substrate.More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
National Cancer Institute-Bethesda
National Cancer Institute-Bethesda
Affinity DataKi: 0.740nM ΔG°: -12.9kcal/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataKi: 0.740nMAssay Description:Inhibition of HIV1 proteaseMore data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,Q508K,L534I,L564I,C568A,C596A,D531N](Human immunodeficiency virus type 1)
University of Florida College of Medicine
University of Florida College of Medicine
Affinity DataKi: 0.900nM ΔG°: -12.8kcal/molepH: 4.7 T: 2°CAssay Description:The inhibition constant, Ki was determined by monitoring the competitive inhibition of the hydrolysis of the chromogenic substrate.More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,D531N,A572V](Human immunodeficiency virus type 1)
University of Florida College of Medicine
University of Florida College of Medicine
Affinity DataKi: 1.10nM ΔG°: -12.7kcal/molepH: 4.7 T: 2°CAssay Description:The inhibition constant, Ki was determined by monitoring the competitive inhibition of the hydrolysis of the chromogenic substrate.More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,D531N,M537I](Human immunodeficiency virus type 1)
University of Florida College of Medicine
University of Florida College of Medicine
Affinity DataKi: 1.70nM ΔG°: -12.4kcal/molepH: 4.7 T: 2°CAssay Description:The inhibition constant, Ki was determined by monitoring the competitive inhibition of the hydrolysis of the chromogenic substrate.More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,D531N,M537I,A572V](Human immunodeficiency virus type 1)
University of Florida College of Medicine
University of Florida College of Medicine
Affinity DataKi: 1.70nM ΔG°: -12.4kcal/molepH: 4.7 T: 2°CAssay Description:The inhibition constant, Ki was determined by monitoring the competitive inhibition of the hydrolysis of the chromogenic substrate.More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Bristol Myers Squibb
Curated by ChEMBL
Bristol Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 6.5nMAssay Description:Inhibition of HIV1 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 1.00E+3nMAssay Description:Inhibition of Plasmodium falciparum plasmepsin-2More data for this Ligand-Target Pair
Activity Spreadsheet -- ITC Data from BindingDB
 Found 6 hits for monomerid = 579
Found 6 hits for monomerid = 579
ITC DataΔG°: -13.3kcal/mole −TΔS°: -7.89kcal/mole ΔH°: -5.39kcal/mole logk: 6.00E+9
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
ITC DataΔG°: -9.09kcal/mole −TΔS°: -11.1kcal/mole ΔH°: 2.00kcal/mole logk: 4.60E+6
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
ITC DataΔG°: -10.9kcal/mole −TΔS°: -10.9kcal/mole ΔH°: 0kcal/mole logk: 9.00E+7
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
CellHIV-1 Protease Mutant (V82A/I84V)(Human immunodeficiency virus type 1)
The Johns Hopkins University
The Johns Hopkins University
ITC DataΔG°: -11.6kcal/mole −TΔS°: -8.19kcal/mole ΔH°: -3.39kcal/mole logk: 3.10E+8
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
CellHIV-1 Protease Mutant (M46I/I54V)(Human immunodeficiency virus type 1)
The Johns Hopkins University
The Johns Hopkins University
ITC DataΔG°: -13.0kcal/mole −TΔS°: -10.1kcal/mole ΔH°: -2.90kcal/mole logk: 3.60E+9
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
CellHIV-1 Protease Mutant (L10I/L90M)(Human immunodeficiency virus type 1)
The Johns Hopkins University
The Johns Hopkins University
ITC DataΔG°: -12.9kcal/mole −TΔS°: -7.69kcal/mole ΔH°: -5.19kcal/mole logk: 2.90E+9
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C