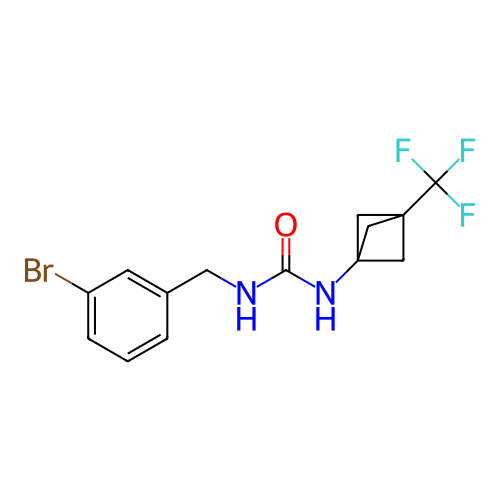Report error Found 203 Enz. Inhib. hit(s) with Target = 'Potassium voltage-gated channel subfamily KQT member 1'
TargetPotassium voltage-gated channel subfamily KQT member 1(Guinea pig)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
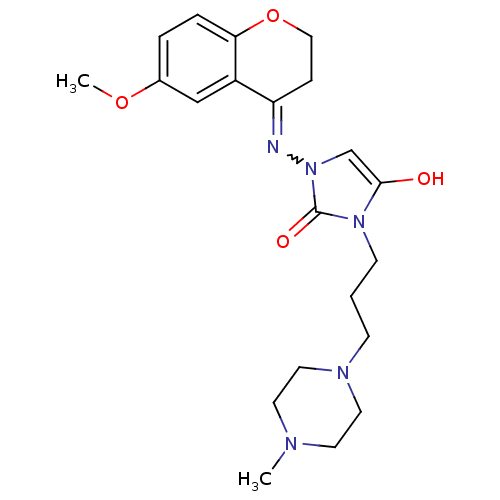
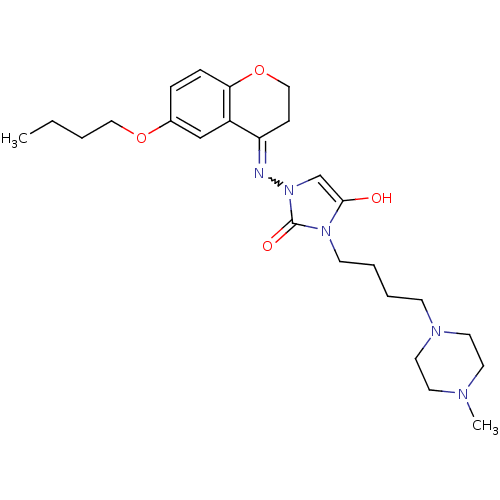
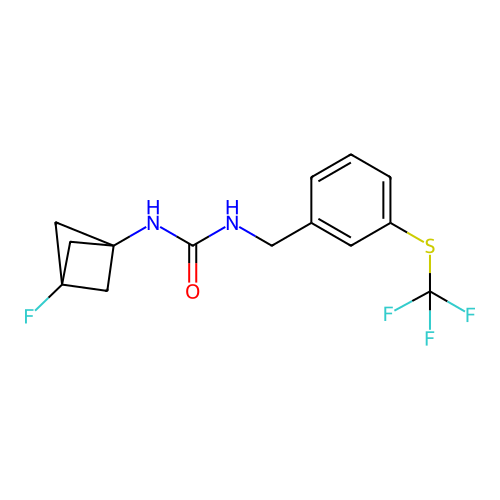
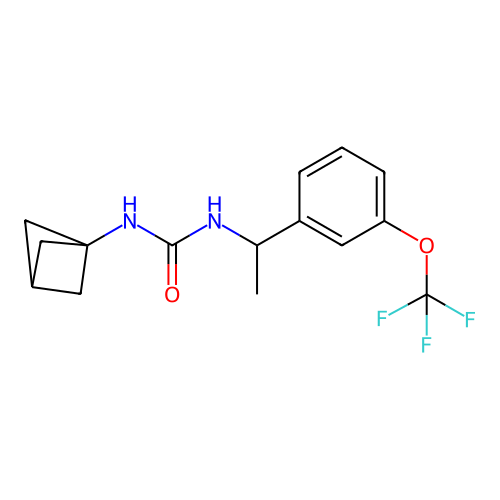
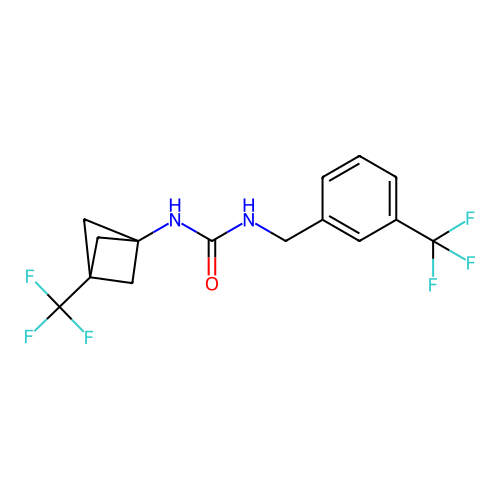
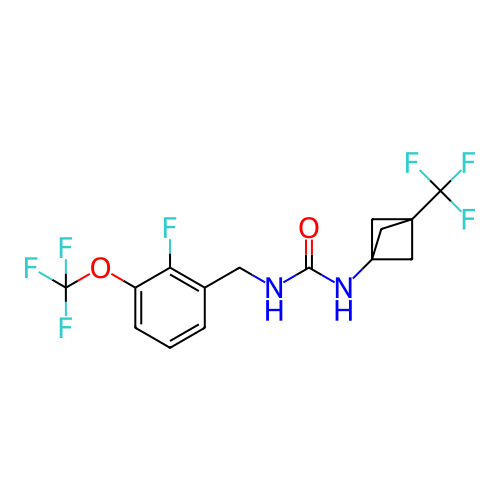
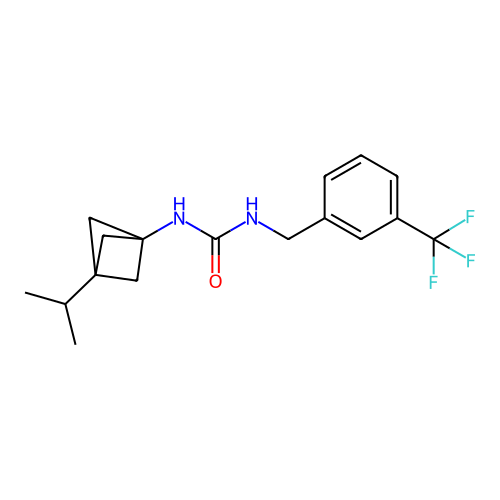
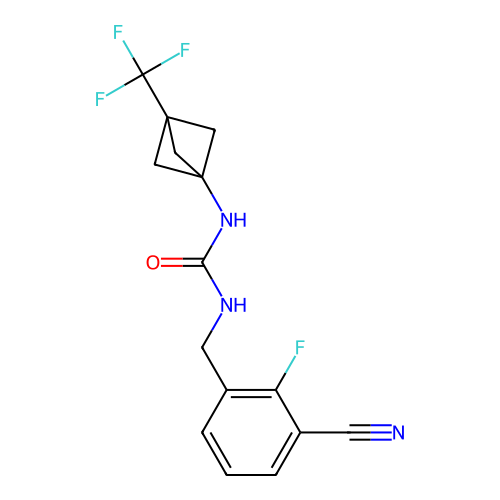
Affinity DataIC50: 1.90nMAssay Description:Inhibition of KVLQT1/minK in guinea pig ventricular myocytes assessed as blockade of slow delayed rectifier potassium current by whole cell patch-cla...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily KQT member 1(Guinea pig)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
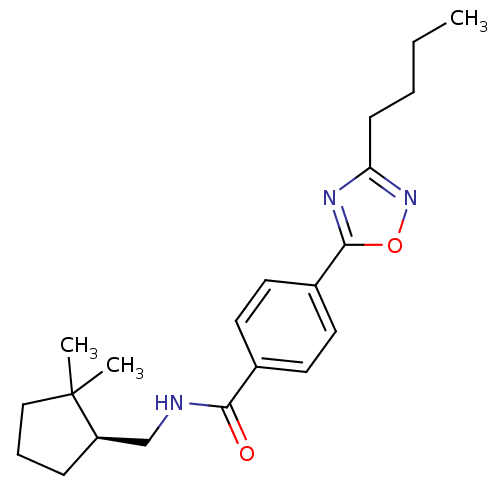
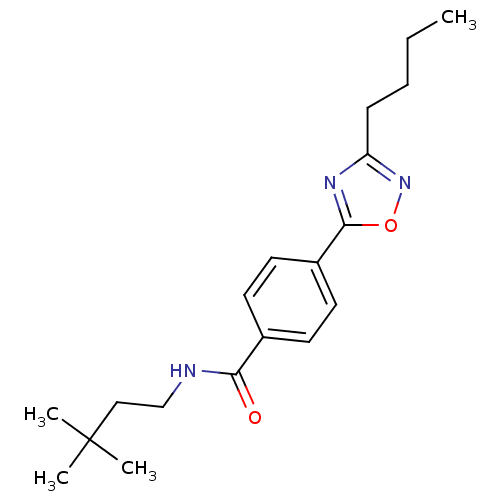
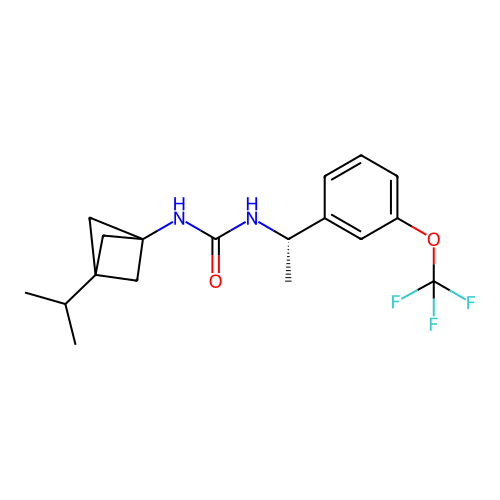
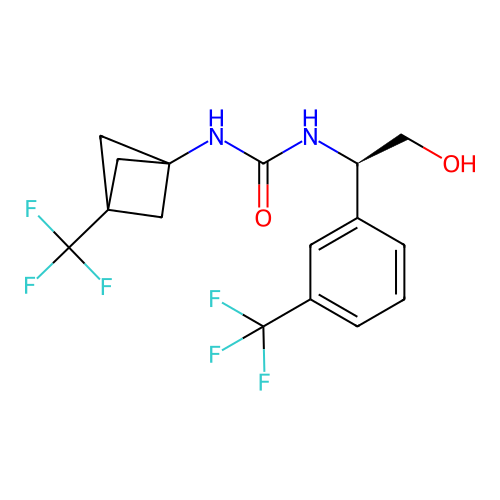
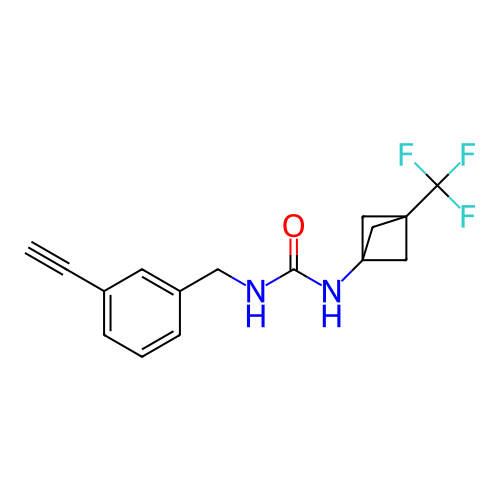
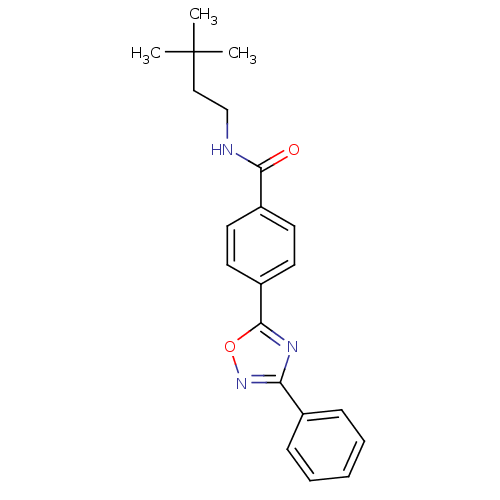
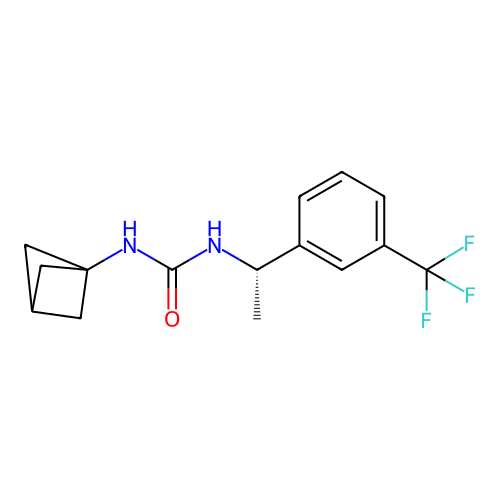
Affinity DataIC50: 2nMAssay Description:Molar concentration required to inhibit 50% of the activating delayed-rectifier K+ current in isolated guinea pig ventricular myocytesMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily KQT member 1(Guinea pig)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
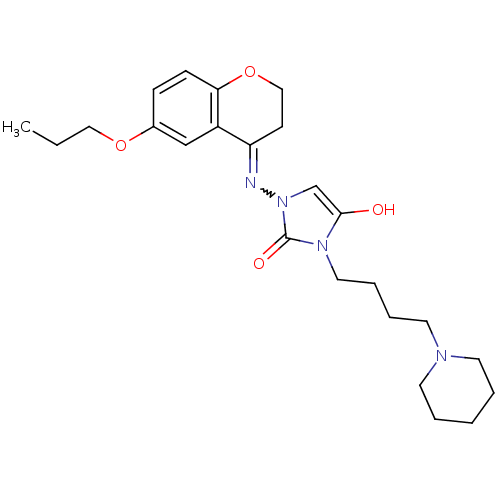
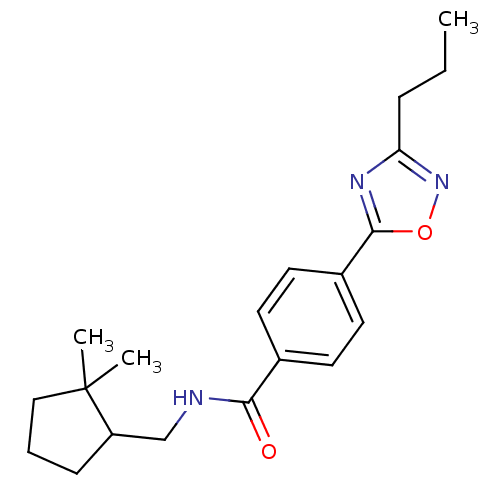
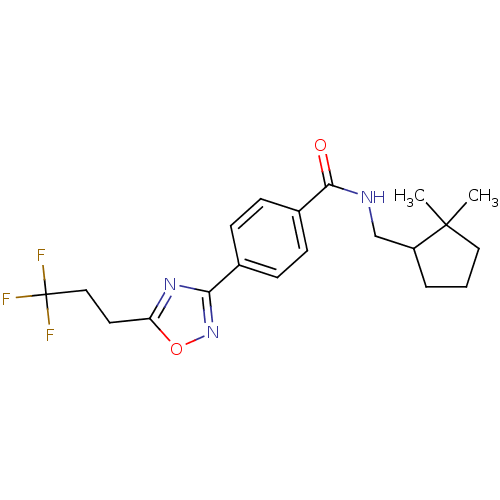
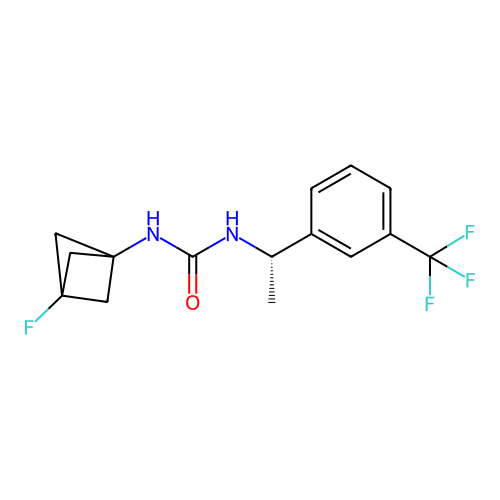
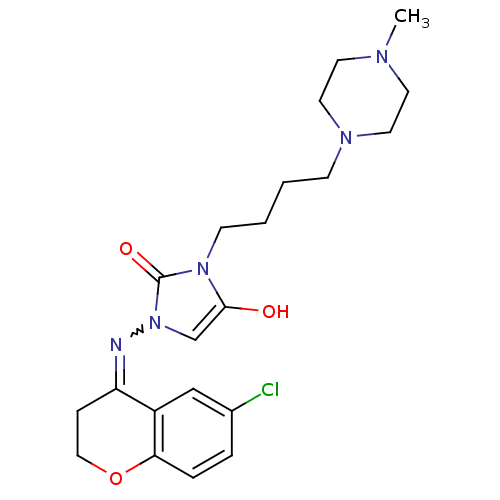
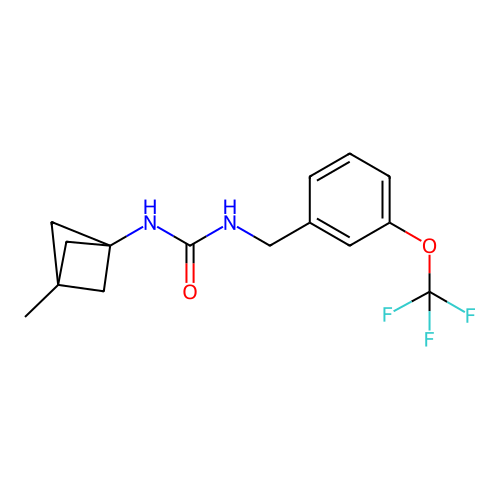
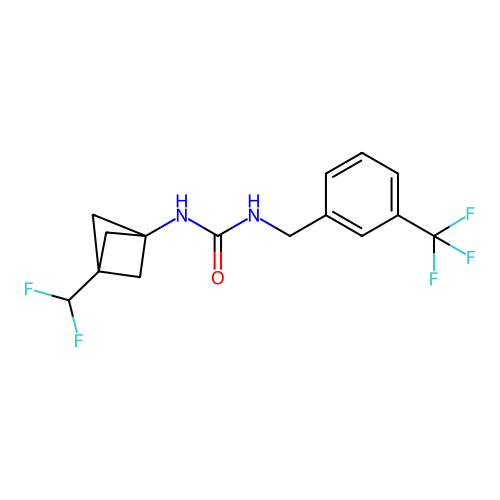
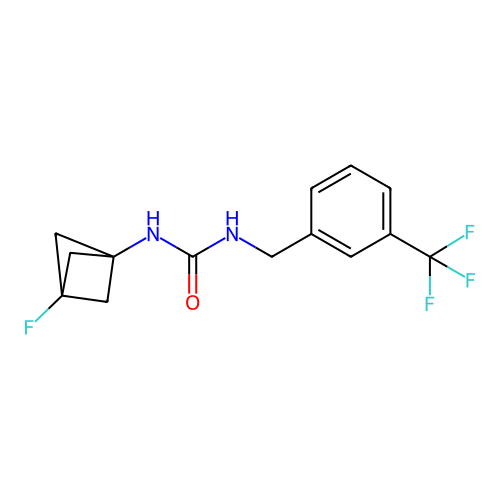
Affinity DataIC50: 3.20nMAssay Description:Inhibition of KVLQT1/minK in guinea pig ventricular myocytes assessed as blockade of slow delayed rectifier potassium current by whole cell patch-cla...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily KQT member 1(Guinea pig)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
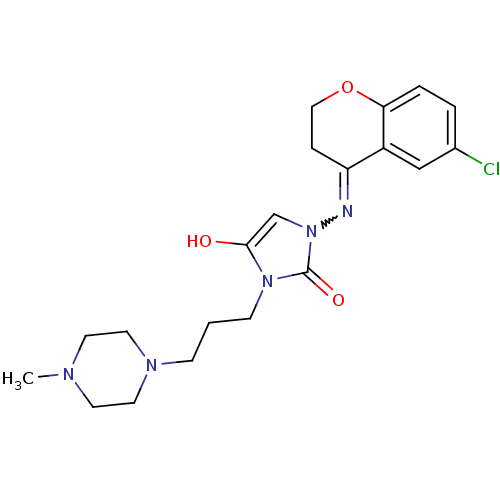
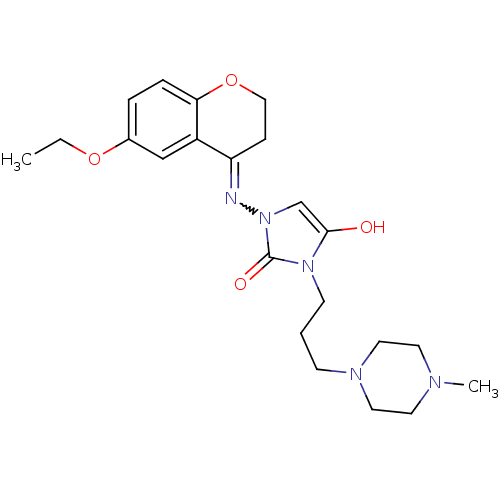
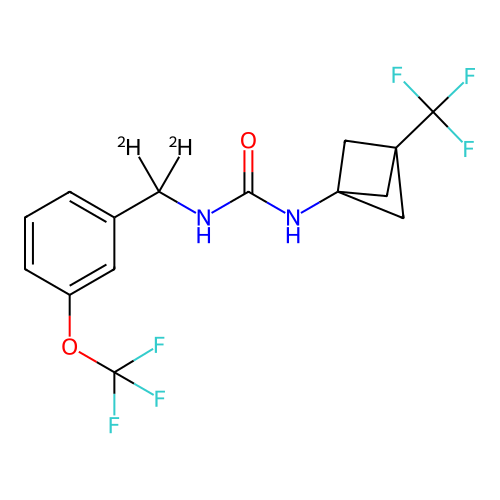
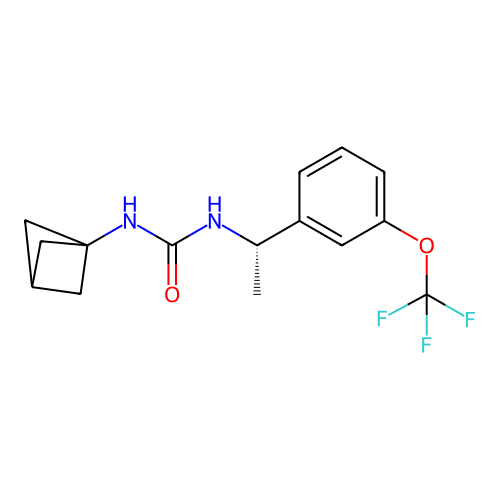
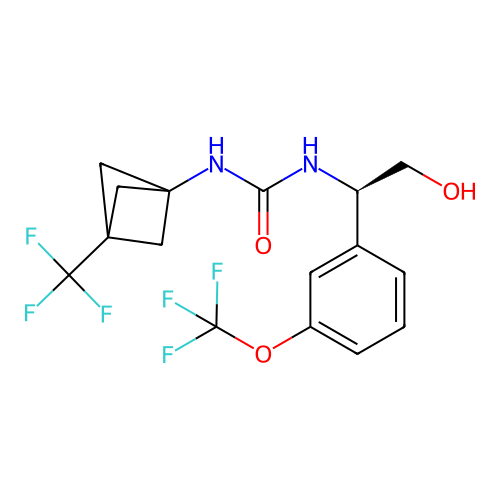
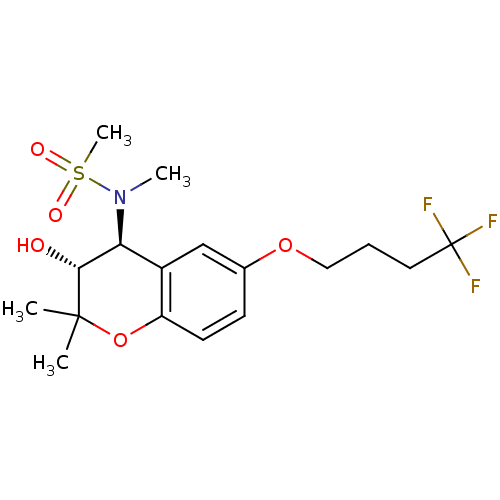
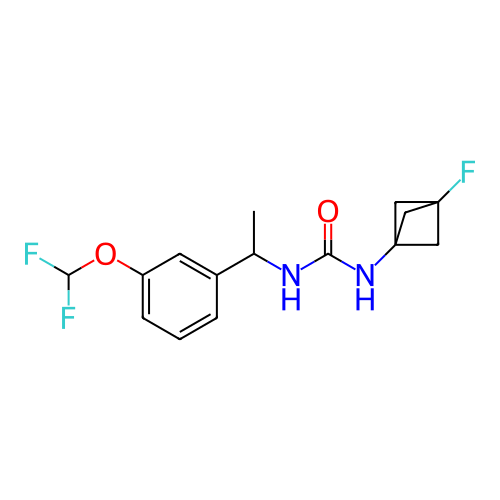
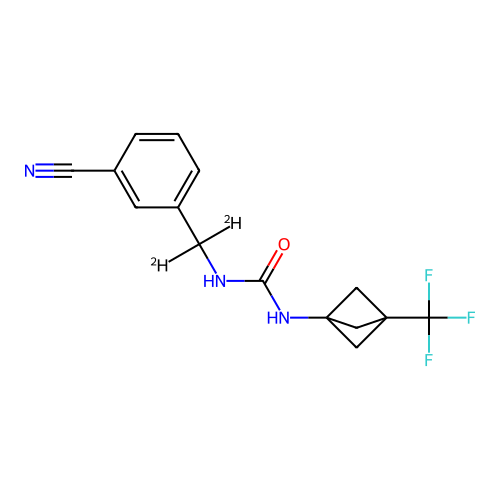
Affinity DataIC50: 5.80nMAssay Description:Inhibition of KVLQT1/minK in guinea pig ventricular myocytes assessed as blockade of slow delayed rectifier potassium current by whole cell patch-cla...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily KQT member 1(Guinea pig)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 9nMAssay Description:Molar concentration required to inhibit 50% of the activating delayed-rectifier K+ current in isolated guinea pig ventricular myocytesMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily KQT member 1(Guinea pig)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 9.5nMAssay Description:Inhibition of KVLQT1/minK in guinea pig ventricular myocytes assessed as blockade of slow delayed rectifier potassium current by whole cell patch-cla...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily KQT member 1(Guinea pig)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 18nMAssay Description:Molar concentration required to inhibit 50% of the activating delayed-rectifier K+ current in isolated guinea pig ventricular myocytesMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily KQT member 1(Guinea pig)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 18nMAssay Description:Molar concentration required to inhibit 50% of the activating delayed-rectifier K+ current in isolated guinea pig ventricular myocytesMore data for this Ligand-Target Pair
Affinity DataEC50: 20nMAssay Description:Solutions were of the following composition:Earle's balanced salt solution (in mM): 135 NaCl, 5.4 KCl, 5 Glucose, 2 CaCl2), 1 MgCl2, 5 HEPES, pH ...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily KQT member 1(Guinea pig)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 22nMAssay Description:Inhibition of KVLQT1/minK in guinea pig ventricular myocytes assessed as blockade of slow delayed rectifier potassium current by whole cell patch-cla...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily KQT member 1(Guinea pig)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 23nMAssay Description:Molar concentration required to inhibit 50% of the activating delayed-rectifier K+ current in isolated guinea pig ventricular myocytesMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily KQT member 1(Guinea pig)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 27nMAssay Description:Molar concentration required to inhibit 50% of the activating delayed-rectifier K+ current in isolated guinea pig ventricular myocytesMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily KQT member 1(Guinea pig)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 27nMAssay Description:Molar concentration required to inhibit 50% of the activating delayed-rectifier K+ current in isolated guinea pig ventricular myocytesMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily KQT member 1(Guinea pig)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 27nMAssay Description:Inhibition of KVLQT1/minK in guinea pig ventricular myocytes assessed as blockade of slow delayed rectifier potassium current by whole cell patch-cla...More data for this Ligand-Target Pair
Affinity DataEC50: 30nMAssay Description:Solutions were of the following composition:Earle's balanced salt solution (in mM): 135 NaCl, 5.4 KCl, 5 Glucose, 2 CaCl2), 1 MgCl2, 5 HEPES, pH ...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily KQT member 1(Guinea pig)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 31nMAssay Description:Molar concentration required to inhibit 50% of the activating delayed-rectifier K+ current in isolated guinea pig ventricular myocytesMore data for this Ligand-Target Pair
Affinity DataEC50: 40nMAssay Description:Solutions were of the following composition:Earle's balanced salt solution (in mM): 135 NaCl, 5.4 KCl, 5 Glucose, 2 CaCl2), 1 MgCl2, 5 HEPES, pH ...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily KQT member 1(Guinea pig)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 40nMAssay Description:Molar concentration required to inhibit 50% of the activating delayed-rectifier K+ current in isolated guinea pig ventricular myocytesMore data for this Ligand-Target Pair
Affinity DataEC50: 40nMAssay Description:Solutions were of the following composition:Earle's balanced salt solution (in mM): 135 NaCl, 5.4 KCl, 5 Glucose, 2 CaCl2), 1 MgCl2, 5 HEPES, pH ...More data for this Ligand-Target Pair
Affinity DataEC50: 40nMAssay Description:Solutions were of the following composition:Earle's balanced salt solution (in mM): 135 NaCl, 5.4 KCl, 5 Glucose, 2 CaCl2), 1 MgCl2, 5 HEPES, pH ...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily KQT member 1(Guinea pig)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 40nMAssay Description:Molar concentration required to inhibit 50% of the activating delayed-rectifier K+ current in isolated guinea pig ventricular myocytesMore data for this Ligand-Target Pair
Affinity DataEC50: 50nMAssay Description:Solutions were of the following composition:Earle's balanced salt solution (in mM): 135 NaCl, 5.4 KCl, 5 Glucose, 2 CaCl2), 1 MgCl2, 5 HEPES, pH ...More data for this Ligand-Target Pair
Affinity DataEC50: 50nMAssay Description:Solutions were of the following composition:Earle's balanced salt solution (in mM): 135 NaCl, 5.4 KCl, 5 Glucose, 2 CaCl2), 1 MgCl2, 5 HEPES, pH ...More data for this Ligand-Target Pair
Affinity DataEC50: 60nMAssay Description:Solutions were of the following composition:Earle's balanced salt solution (in mM): 135 NaCl, 5.4 KCl, 5 Glucose, 2 CaCl2), 1 MgCl2, 5 HEPES, pH ...More data for this Ligand-Target Pair
Affinity DataEC50: 60nMAssay Description:Solutions were of the following composition:Earle's balanced salt solution (in mM): 135 NaCl, 5.4 KCl, 5 Glucose, 2 CaCl2), 1 MgCl2, 5 HEPES, pH ...More data for this Ligand-Target Pair
Affinity DataEC50: 60nMAssay Description:Solutions were of the following composition:Earle's balanced salt solution (in mM): 135 NaCl, 5.4 KCl, 5 Glucose, 2 CaCl2), 1 MgCl2, 5 HEPES, pH ...More data for this Ligand-Target Pair
Affinity DataEC50: 60nMAssay Description:Solutions were of the following composition:Earle's balanced salt solution (in mM): 135 NaCl, 5.4 KCl, 5 Glucose, 2 CaCl2), 1 MgCl2, 5 HEPES, pH ...More data for this Ligand-Target Pair
Affinity DataEC50: 70nMAssay Description:Solutions were of the following composition:Earle's balanced salt solution (in mM): 135 NaCl, 5.4 KCl, 5 Glucose, 2 CaCl2), 1 MgCl2, 5 HEPES, pH ...More data for this Ligand-Target Pair
Affinity DataEC50: 70nMAssay Description:Solutions were of the following composition:Earle's balanced salt solution (in mM): 135 NaCl, 5.4 KCl, 5 Glucose, 2 CaCl2), 1 MgCl2, 5 HEPES, pH ...More data for this Ligand-Target Pair
Affinity DataEC50: 70nMAssay Description:Solutions were of the following composition:Earle's balanced salt solution (in mM): 135 NaCl, 5.4 KCl, 5 Glucose, 2 CaCl2), 1 MgCl2, 5 HEPES, pH ...More data for this Ligand-Target Pair
Affinity DataEC50: 70nMAssay Description:Solutions were of the following composition:Earle's balanced salt solution (in mM): 135 NaCl, 5.4 KCl, 5 Glucose, 2 CaCl2), 1 MgCl2, 5 HEPES, pH ...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily KQT member 1(Guinea pig)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 73nMAssay Description:Inhibition of KVLQT1/minK in guinea pig ventricular myocytes assessed as blockade of slow delayed rectifier potassium current by whole cell patch-cla...More data for this Ligand-Target Pair
Affinity DataEC50: 80nMAssay Description:Solutions were of the following composition:Earle's balanced salt solution (in mM): 135 NaCl, 5.4 KCl, 5 Glucose, 2 CaCl2), 1 MgCl2, 5 HEPES, pH ...More data for this Ligand-Target Pair
Affinity DataEC50: 80nMAssay Description:Solutions were of the following composition:Earle's balanced salt solution (in mM): 135 NaCl, 5.4 KCl, 5 Glucose, 2 CaCl2), 1 MgCl2, 5 HEPES, pH ...More data for this Ligand-Target Pair
Affinity DataEC50: 80nMAssay Description:Solutions were of the following composition:Earle's balanced salt solution (in mM): 135 NaCl, 5.4 KCl, 5 Glucose, 2 CaCl2), 1 MgCl2, 5 HEPES, pH ...More data for this Ligand-Target Pair
Affinity DataEC50: 80nMAssay Description:Solutions were of the following composition:Earle's balanced salt solution (in mM): 135 NaCl, 5.4 KCl, 5 Glucose, 2 CaCl2), 1 MgCl2, 5 HEPES, pH ...More data for this Ligand-Target Pair
Affinity DataEC50: 80nMAssay Description:Solutions were of the following composition:Earle's balanced salt solution (in mM): 135 NaCl, 5.4 KCl, 5 Glucose, 2 CaCl2), 1 MgCl2, 5 HEPES, pH ...More data for this Ligand-Target Pair
Affinity DataEC50: 90nMAssay Description:Solutions were of the following composition:Earle's balanced salt solution (in mM): 135 NaCl, 5.4 KCl, 5 Glucose, 2 CaCl2), 1 MgCl2, 5 HEPES, pH ...More data for this Ligand-Target Pair
Affinity DataEC50: 100nMAssay Description:Solutions were of the following composition:Earle's balanced salt solution (in mM): 135 NaCl, 5.4 KCl, 5 Glucose, 2 CaCl2), 1 MgCl2, 5 HEPES, pH ...More data for this Ligand-Target Pair
Affinity DataEC50: 100nMAssay Description:Solutions were of the following composition:Earle's balanced salt solution (in mM): 135 NaCl, 5.4 KCl, 5 Glucose, 2 CaCl2), 1 MgCl2, 5 HEPES, pH ...More data for this Ligand-Target Pair
Affinity DataEC50: 100nMAssay Description:Solutions were of the following composition:Earle's balanced salt solution (in mM): 135 NaCl, 5.4 KCl, 5 Glucose, 2 CaCl2), 1 MgCl2, 5 HEPES, pH ...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily KQT member 1(Guinea pig)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 120nMAssay Description:Inhibition of guinea pig Iks potassium channel expressed in Xenopus oocytesMore data for this Ligand-Target Pair
Affinity DataEC50: 120nMAssay Description:Solutions were of the following composition:Earle's balanced salt solution (in mM): 135 NaCl, 5.4 KCl, 5 Glucose, 2 CaCl2), 1 MgCl2, 5 HEPES, pH ...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily KQT member 1(Guinea pig)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 130nMAssay Description:Molar concentration required to inhibit 50% of the activating delayed-rectifier K+ current in isolated guinea pig ventricular myocytesMore data for this Ligand-Target Pair
Affinity DataEC50: 160nMAssay Description:Solutions were of the following composition:Earle's balanced salt solution (in mM): 135 NaCl, 5.4 KCl, 5 Glucose, 2 CaCl2), 1 MgCl2, 5 HEPES, pH ...More data for this Ligand-Target Pair
Affinity DataEC50: 180nMAssay Description:Solutions were of the following composition:Earle's balanced salt solution (in mM): 135 NaCl, 5.4 KCl, 5 Glucose, 2 CaCl2), 1 MgCl2, 5 HEPES, pH ...More data for this Ligand-Target Pair
Affinity DataEC50: 210nMAssay Description:Solutions were of the following composition:Earle's balanced salt solution (in mM): 135 NaCl, 5.4 KCl, 5 Glucose, 2 CaCl2), 1 MgCl2, 5 HEPES, pH ...More data for this Ligand-Target Pair
Affinity DataEC50: 230nMAssay Description:Solutions were of the following composition:Earle's balanced salt solution (in mM): 135 NaCl, 5.4 KCl, 5 Glucose, 2 CaCl2), 1 MgCl2, 5 HEPES, pH ...More data for this Ligand-Target Pair
Affinity DataEC50: 240nMAssay Description:Solutions were of the following composition:Earle's balanced salt solution (in mM): 135 NaCl, 5.4 KCl, 5 Glucose, 2 CaCl2), 1 MgCl2, 5 HEPES, pH ...More data for this Ligand-Target Pair
Affinity DataEC50: 240nMAssay Description:Solutions were of the following composition:Earle's balanced salt solution (in mM): 135 NaCl, 5.4 KCl, 5 Glucose, 2 CaCl2), 1 MgCl2, 5 HEPES, pH ...More data for this Ligand-Target Pair