Report error Found 66 with Last Name = 'mcpherson' and Initial = 'dw'
Affinity DataKi: 0.0300nMAssay Description:Ability to displace [3H](-)-quinuclidinyl bezilate(QNB) from M2 receptor in rat heart homogenateMore data for this Ligand-Target Pair
Affinity DataKi: 0.120nMAssay Description:Ability to displace [3H]N-methylscopolamine (NMS) from M3 receptor in rat submaxillary gland homogenateMore data for this Ligand-Target Pair
Affinity DataKi: 0.150nMAssay Description:Ability to displace [3H]pirenzepine (PZ) from M1 receptor in rat cortex homogenateMore data for this Ligand-Target Pair
Affinity DataKi: 0.160nMAssay Description:Ability to displace [3H]N-methylscopolamine (NMS) from M3 receptor in rat submaxillary gland homogenateMore data for this Ligand-Target Pair
Affinity DataKi: 0.190nMAssay Description:Binding affinity against Muscarinic acetylcholine receptor M1 by displacement of [3H]pirenzepine in bovine striatumMore data for this Ligand-Target Pair
Affinity DataKi: 0.200nMAssay Description:Binding affinity against Muscarinic acetylcholine receptor M1 by displacement of [3H]pirenzepine in bovine striatumMore data for this Ligand-Target Pair
Affinity DataKi: 0.200nMAssay Description:Binding affinity against Muscarinic acetylcholine receptor M1 by displacement of [3H]pirenzepine in bovine striatumMore data for this Ligand-Target Pair
Affinity DataKi: 0.230nMAssay Description:Evaluated for the inhibition of adenylate cyclase at M2 receptor in rat heartMore data for this Ligand-Target Pair
Affinity DataKi: 0.260nMAssay Description:Ability to displace [3H]pirenzepine (PZ) from M1 receptor in rat cortex homogenateMore data for this Ligand-Target Pair
Affinity DataKi: 0.270nMAssay Description:Ability to displace [3H]N-methylscopolamine (NMS) from M3 receptor in rat submaxillary gland homogenateMore data for this Ligand-Target Pair
Affinity DataKi: 0.290nMAssay Description:Binding affinity against Muscarinic acetylcholine receptor M1 by displacement of [3H]pirenzepine in bovine striatumMore data for this Ligand-Target Pair
Affinity DataKi: 0.390nMAssay Description:Binding affinity against Muscarinic acetylcholine receptor M2 by displacement of [3H]QNB in rat myocardiumMore data for this Ligand-Target Pair
Affinity DataKi: 0.450nMAssay Description:Displacement of [3H]QNB from rat ileum Muscarinic acetylcholine receptorMore data for this Ligand-Target Pair
Affinity DataKi: 0.5nMAssay Description:Evaluated for the phosphatidyl inositol turnover at Muscarinic acetylcholine receptor M1 in rat cortexMore data for this Ligand-Target Pair
Affinity DataKi: 0.570nMAssay Description:Binding affinity against Muscarinic acetylcholine receptor M1 by displacement of [3H]pirenzepine in bovine striatumMore data for this Ligand-Target Pair
Affinity DataKi: 0.700nMAssay Description:Binding affinity against Muscarinic acetylcholine receptor M2 by displacement of [3H]QNB in rat myocardiumMore data for this Ligand-Target Pair
Affinity DataKi: 0.760nMAssay Description:Ability to displace [3H](-)-quinuclidinyl bezilate(QNB) from M2 receptor in rat heart homogenateMore data for this Ligand-Target Pair
Affinity DataKi: 0.780nMAssay Description:Ability to displace [3H]pirenzepine (PZ) from M1 receptor in rat cortex homogenateMore data for this Ligand-Target Pair
Affinity DataKi: 1.10nMAssay Description:Ability to displace [3H](-)-quinuclidinyl bezilate(QNB) from M2 receptor in rat heart homogenateMore data for this Ligand-Target Pair
Affinity DataKi: 2.20nMAssay Description:Evaluated for the inhibition of adenylate cyclase at M2 receptor in rat heartMore data for this Ligand-Target Pair
Affinity DataKi: 2.90nMAssay Description:Pseudo hill coefficient at Muscarinic acetylcholine receptor M1More data for this Ligand-Target Pair
Affinity DataKi: 3.60nMAssay Description:Binding affinity against Muscarinic acetylcholine receptor M1 by displacement of [3H]pirenzepine in bovine striatumMore data for this Ligand-Target Pair
Affinity DataKi: 5.20nMAssay Description:Ability to displace [3H]pirenzepine (PZ) from M1 receptor in rat cortex homogenateMore data for this Ligand-Target Pair
Affinity DataKi: 5.20nMAssay Description:Binding affinity against Muscarinic acetylcholine receptor M1 by displacement of [3H]pirenzepine in bovine striatumMore data for this Ligand-Target Pair
Affinity DataKi: 5.80nMAssay Description:Binding affinity against Muscarinic acetylcholine receptor M2 by displacement of [3H]QNB in rat myocardiumMore data for this Ligand-Target Pair
Affinity DataKi: 6nMAssay Description:Displacement of [3H]QNB from rat ileum Muscarinic acetylcholine receptorMore data for this Ligand-Target Pair
Affinity DataKi: 6.90nMAssay Description:Binding affinity against Muscarinic acetylcholine receptor M2 by displacement of [3H]QNB in rat myocardiumMore data for this Ligand-Target Pair
Affinity DataKi: 6.90nMAssay Description:Evaluated for the inhibition of adenylate cyclase at M2 receptor in rat heartMore data for this Ligand-Target Pair
Affinity DataKi: 7.5nMAssay Description:Evaluated for the inhibition of adenylate cyclase at M2 receptor in rat heartMore data for this Ligand-Target Pair
Affinity DataKi: 19nMAssay Description:Binding affinity against Muscarinic acetylcholine receptor M1 by displacement of [3H]pirenzepine in bovine striatumMore data for this Ligand-Target Pair
Affinity DataKi: 27nMAssay Description:Evaluated for the phosphatidyl inositol turnover at Muscarinic acetylcholine receptor M1 in rat cortexMore data for this Ligand-Target Pair
Affinity DataKi: 28nMAssay Description:Ability to displace [3H]N-methylscopolamine (NMS) from M3 receptor in rat submaxillary gland homogenateMore data for this Ligand-Target Pair
Affinity DataKi: 37nMAssay Description:Binding affinity against Muscarinic acetylcholine receptor M2 by displacement of [3H]QNB in rat myocardiumMore data for this Ligand-Target Pair
Affinity DataKi: 74nMAssay Description:Binding affinity against Muscarinic acetylcholine receptor M2 by displacement of [3H]QNB in rat myocardiumMore data for this Ligand-Target Pair
Affinity DataKi: 123nMAssay Description:Evaluated for the inhibition of adenylate cyclase at M2 receptor in rat heartMore data for this Ligand-Target Pair
Affinity DataKi: 153nMAssay Description:Binding affinity against Muscarinic acetylcholine receptor M2 by displacement of [3H]QNB in rat myocardiumMore data for this Ligand-Target Pair
Affinity DataKi: 156nMAssay Description:Binding affinity against Muscarinic acetylcholine receptor M1 by displacement of [3H]pirenzepine in bovine striatumMore data for this Ligand-Target Pair
Affinity DataKi: 158nMAssay Description:Displacement of [3H]QNB from rat ileum Muscarinic acetylcholine receptorMore data for this Ligand-Target Pair
Affinity DataKi: 195nMAssay Description:Displacement of [3H]QNB from rat ileum Muscarinic acetylcholine receptorMore data for this Ligand-Target Pair
Affinity DataKi: 267nMAssay Description:Ability to displace [3H](-)-quinuclidinyl bezilate(QNB) from M2 receptor in rat heart homogenateMore data for this Ligand-Target Pair
Affinity DataKi: 377nMAssay Description:Evaluated for the inhibition of adenylate cyclase at M2 receptor in rat heartMore data for this Ligand-Target Pair
Affinity DataKi: 934nMAssay Description:Binding affinity against Muscarinic acetylcholine receptor M2 by displacement of [3H]QNB in rat myocardiumMore data for this Ligand-Target Pair
Affinity DataKi: 1.37E+3nMAssay Description:Binding affinity against Muscarinic acetylcholine receptor M2 by displacement of [3H]QNB in rat myocardiumMore data for this Ligand-Target Pair
Affinity DataKi: 1.46E+3nMAssay Description:Evaluated for the inhibition of adenylate cyclase at M2 receptor in rat heartMore data for this Ligand-Target Pair
Affinity DataKi: 1.62E+3nMAssay Description:Displacement of [3H]QNB from rat ileum Muscarinic acetylcholine receptorMore data for this Ligand-Target Pair
Affinity DataKi: 1.66E+3nMAssay Description:Evaluated for the inhibition of adenylate cyclase at M2 receptor in rat heartMore data for this Ligand-Target Pair
Affinity DataKi: 2.38E+3nMAssay Description:Displacement of [3H]QNB from rat ileum Muscarinic acetylcholine receptorMore data for this Ligand-Target Pair
Affinity DataKi: 3.98E+3nMAssay Description:Displacement of [3H]QNB from rat ileum Muscarinic acetylcholine receptorMore data for this Ligand-Target Pair
Affinity DataKi: 5.24E+3nMAssay Description:Displacement of [3H]QNB from rat ileum Muscarinic acetylcholine receptorMore data for this Ligand-Target Pair
Affinity DataKi: 8.28E+3nMAssay Description:Displacement of [3H]QNB from rat ileum Muscarinic acetylcholine receptorMore data for this Ligand-Target Pair
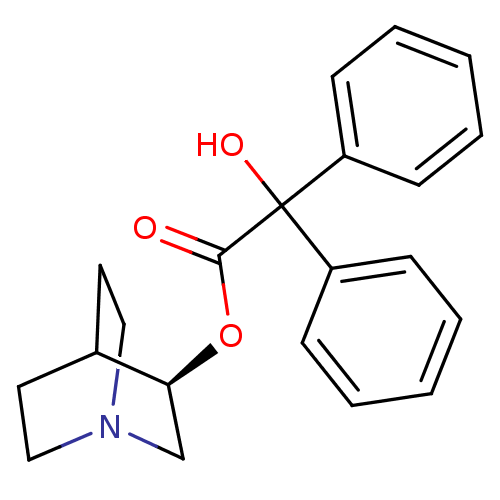
 3D Structure (crystal)
3D Structure (crystal)