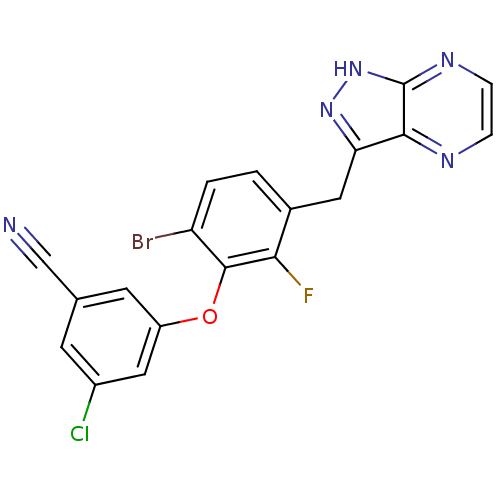
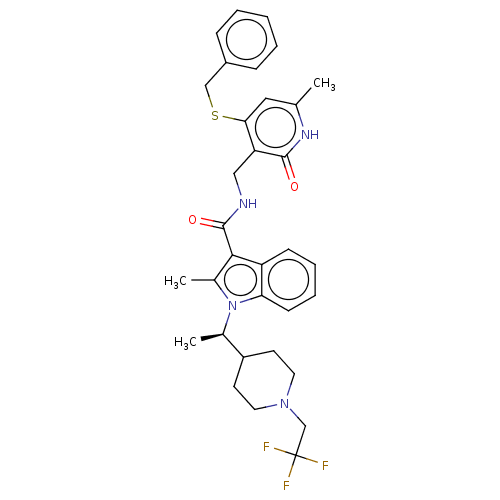
TargetAcetylcholinesterase(Electrophorus electricus (Electric eel))
Guru Nanak Dev University
Curated by ChEMBL
Guru Nanak Dev University
Curated by ChEMBL
Affinity DataKi: 0.0417nMAssay Description:Mixed type inhibition of electric eel AChE assessed as inhibition constant using varying levels of acetylthiocholine as substrate by reciprocal Linew...More data for this Ligand-Target Pair
TargetDelta-type opioid receptor(MOUSE)
University Of Medicine And Dentistry Of New Jersey-Robert Wood Johnson Medical School And The Informatics Institute Of Umdnj
Curated by ChEMBL
University Of Medicine And Dentistry Of New Jersey-Robert Wood Johnson Medical School And The Informatics Institute Of Umdnj
Curated by ChEMBL
Affinity DataKi: 460nMAssay Description:Displacement of [3H]bremazocine from mouse delta opioid receptor expressed in HEK293 cells by liquid scintillation countingMore data for this Ligand-Target Pair
TargetDelta-type opioid receptor(MOUSE)
University Of Medicine And Dentistry Of New Jersey-Robert Wood Johnson Medical School And The Informatics Institute Of Umdnj
Curated by ChEMBL
University Of Medicine And Dentistry Of New Jersey-Robert Wood Johnson Medical School And The Informatics Institute Of Umdnj
Curated by ChEMBL
Affinity DataKi: 2.60E+3nMAssay Description:Displacement of [3H]bremazocine from mouse delta opioid receptor expressed in HEK293 cells by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 3.53E+3nMAssay Description:Binding affinity to PKM2 (unknown origin) assessed as oxidation of beta-NADH by spectrophotometry based LDH coupled assayMore data for this Ligand-Target Pair
Affinity DataKi: 3.53E+3nMAssay Description:Binding affinity to PKM2 (unknown origin) assessed as oxidation of beta-NADH by spectrophotometry based LDH coupled assayMore data for this Ligand-Target Pair
Affinity DataKi: 3.53E+3nMAssay Description:Binding affinity to PKM2 (unknown origin) assessed as oxidation of beta-NADH by spectrophotometry based LDH coupled assayMore data for this Ligand-Target Pair
Affinity DataKi: 3.53E+3nMAssay Description:Binding affinity to PKM2 (unknown origin) assessed as oxidation of beta-NADH by spectrophotometry based LDH coupled assayMore data for this Ligand-Target Pair
Affinity DataKi: 3.53E+3nMAssay Description:Binding affinity to PKM2 (unknown origin) assessed as oxidation of beta-NADH by spectrophotometry based LDH coupled assayMore data for this Ligand-Target Pair
TargetMu-type opioid receptor(MOUSE)
University Of Medicine And Dentistry Of New Jersey-Robert Wood Johnson Medical School And The Informatics Institute Of Umdnj
Curated by ChEMBL
University Of Medicine And Dentistry Of New Jersey-Robert Wood Johnson Medical School And The Informatics Institute Of Umdnj
Curated by ChEMBL
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [3H]bremazocine from mouse mu opioid receptor expressed in HEK293 cells by liquid scintillation countingMore data for this Ligand-Target Pair
TargetKappa-type opioid receptor(Rattus norvegicus (rat))
University Of Medicine And Dentistry Of New Jersey-Robert Wood Johnson Medical School And The Informatics Institute Of Umdnj
Curated by ChEMBL
University Of Medicine And Dentistry Of New Jersey-Robert Wood Johnson Medical School And The Informatics Institute Of Umdnj
Curated by ChEMBL
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [3H]bremazocine from rat kappa opioid receptor expressed in HEK293 cells by liquid scintillation countingMore data for this Ligand-Target Pair
TargetKappa-type opioid receptor(Rattus norvegicus (rat))
University Of Medicine And Dentistry Of New Jersey-Robert Wood Johnson Medical School And The Informatics Institute Of Umdnj
Curated by ChEMBL
University Of Medicine And Dentistry Of New Jersey-Robert Wood Johnson Medical School And The Informatics Institute Of Umdnj
Curated by ChEMBL
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [3H]bremazocine from rat kappa opioid receptor expressed in HEK293 cells by liquid scintillation countingMore data for this Ligand-Target Pair
TargetMu-type opioid receptor(MOUSE)
University Of Medicine And Dentistry Of New Jersey-Robert Wood Johnson Medical School And The Informatics Institute Of Umdnj
Curated by ChEMBL
University Of Medicine And Dentistry Of New Jersey-Robert Wood Johnson Medical School And The Informatics Institute Of Umdnj
Curated by ChEMBL
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [3H]bremazocine from mouse mu opioid receptor expressed in HEK293 cells by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 2.50E+4nMAssay Description:Inhibition of HIV1 protease by FRET-based assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.50E+4nMAssay Description:Inhibition of HIV1 protease by FRET-based assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.16E+5nMAssay Description:Binding affinity to Leishmania mexicana PKMore data for this Ligand-Target Pair
Affinity DataKi: 5.00E+5nMAssay Description:Inhibition of HIV1 protease by FRET-based assayMore data for this Ligand-Target Pair
Affinity DataKi: 5.00E+5nMAssay Description:Inhibition of HIV1 protease by FRET-based assayMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Homo sapiens (Human))
Constellation Pharmaceuticals
Curated by ChEMBL
Constellation Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 0.0570nMAssay Description:Inhibition of N-terminal FLAG-tagged EZH2 in PRC2 complex (unknown origin) expressed in baculovirus infected Sf9 cells using H2N-RKQLATKAAR(Kme0)SAPA...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Homo sapiens (Human))
Constellation Pharmaceuticals
Curated by ChEMBL
Constellation Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 0.0590nMAssay Description:Inhibition of N-terminal FLAG-tagged EZH2 in PRC2 complex (unknown origin) expressed in baculovirus infected Sf9 cells using H2N-RKQLATKAAR(Kme0)SAPA...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Homo sapiens (Human))
Constellation Pharmaceuticals
Curated by ChEMBL
Constellation Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 0.0600nMAssay Description:Inhibition of N-terminal FLAG-tagged EZH2 in PRC2 complex (unknown origin) expressed in baculovirus infected Sf9 cells using H2N-RKQLATKAAR(Kme0)SAPA...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Homo sapiens (Human))
Constellation Pharmaceuticals
Curated by ChEMBL
Constellation Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 0.0630nMAssay Description:Inhibition of N-terminal FLAG-tagged EZH2 in PRC2 complex (unknown origin) expressed in baculovirus infected Sf9 cells using H2N-RKQLATKAAR(Kme0)SAPA...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Homo sapiens (Human))
Constellation Pharmaceuticals
Curated by ChEMBL
Constellation Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 0.0670nMAssay Description:Inhibition of N-terminal FLAG-tagged EZH2 in PRC2 complex (unknown origin) expressed in baculovirus infected Sf9 cells using H2N-RKQLATKAAR(Kme0)SAPA...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Homo sapiens (Human))
Constellation Pharmaceuticals
Curated by ChEMBL
Constellation Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 0.0720nMAssay Description:Inhibition of N-terminal FLAG-tagged EZH2 in PRC2 complex (unknown origin) expressed in baculovirus infected Sf9 cells using H2N-RKQLATKAAR(Kme0)SAPA...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Homo sapiens (Human))
Constellation Pharmaceuticals
Curated by ChEMBL
Constellation Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 0.0920nMAssay Description:Inhibition of N-terminal FLAG-tagged EZH2 in PRC2 complex (unknown origin) expressed in baculovirus infected Sf9 cells using H2N-RKQLATKAAR(Kme0)SAPA...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Homo sapiens (Human))
Constellation Pharmaceuticals
Curated by ChEMBL
Constellation Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 0.100nMAssay Description:Inhibition of N-terminal FLAG-tagged EZH2 in PRC2 complex (unknown origin) expressed in baculovirus infected Sf9 cells using H2N-RKQLATKAAR(Kme0)SAPA...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Homo sapiens (Human))
Constellation Pharmaceuticals
Curated by ChEMBL
Constellation Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 0.100nMAssay Description:Inhibition of N-terminal FLAG-tagged EZH2 in PRC2 complex (unknown origin) expressed in baculovirus infected Sf9 cells using H2N-RKQLATKAAR(Kme0)SAPA...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Homo sapiens (Human))
Constellation Pharmaceuticals
Curated by ChEMBL
Constellation Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 0.120nMAssay Description:Inhibition of N-terminal FLAG-tagged EZH2 in PRC2 complex (unknown origin) expressed in baculovirus infected Sf9 cells using H2N-RKQLATKAAR(Kme0)SAPA...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Homo sapiens (Human))
Constellation Pharmaceuticals
Curated by ChEMBL
Constellation Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 0.130nMAssay Description:Inhibition of N-terminal FLAG-tagged EZH2 in PRC2 complex (unknown origin) expressed in baculovirus infected Sf9 cells using H2N-RKQLATKAAR(Kme0)SAPA...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Homo sapiens (Human))
Constellation Pharmaceuticals
Curated by ChEMBL
Constellation Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 0.140nMAssay Description:Inhibition of N-terminal FLAG-tagged EZH2 in PRC2 complex (unknown origin) expressed in baculovirus infected Sf9 cells using H2N-RKQLATKAAR(Kme0)SAPA...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Homo sapiens (Human))
Constellation Pharmaceuticals
Curated by ChEMBL
Constellation Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 0.160nMAssay Description:Inhibition of N-terminal FLAG-tagged EZH2 in PRC2 complex (unknown origin) expressed in baculovirus infected Sf9 cells using H2N-RKQLATKAAR(Kme0)SAPA...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Homo sapiens (Human))
Constellation Pharmaceuticals
Curated by ChEMBL
Constellation Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 0.180nMAssay Description:Inhibition of N-terminal FLAG-tagged EZH2 in PRC2 complex (unknown origin) expressed in baculovirus infected Sf9 cells using H2N-RKQLATKAAR(Kme0)SAPA...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Homo sapiens (Human))
Constellation Pharmaceuticals
Curated by ChEMBL
Constellation Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 0.180nMAssay Description:Inhibition of N-terminal FLAG-tagged EZH2 in PRC2 complex (unknown origin) expressed in baculovirus infected Sf9 cells using H2N-RKQLATKAAR(Kme0)SAPA...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Homo sapiens (Human))
Constellation Pharmaceuticals
Curated by ChEMBL
Constellation Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 0.200nMAssay Description:Inhibition of N-terminal FLAG-tagged EZH2 in PRC2 complex (unknown origin) expressed in baculovirus infected Sf9 cells using H2N-RKQLATKAAR(Kme0)SAPA...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Homo sapiens (Human))
Constellation Pharmaceuticals
Curated by ChEMBL
Constellation Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 0.200nMAssay Description:Inhibition of N-terminal FLAG-tagged EZH2 in PRC2 complex (unknown origin) expressed in baculovirus infected Sf9 cells using H2N-RKQLATKAAR(Kme0)SAPA...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Homo sapiens (Human))
Constellation Pharmaceuticals
Curated by ChEMBL
Constellation Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 0.200nMAssay Description:Inhibition of N-terminal FLAG-tagged EZH2 in PRC2 complex (unknown origin) expressed in baculovirus infected Sf9 cells using H2N-RKQLATKAAR(Kme0)SAPA...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Homo sapiens (Human))
Constellation Pharmaceuticals
Curated by ChEMBL
Constellation Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 0.200nMAssay Description:Inhibition of N-terminal FLAG-tagged EZH2 in PRC2 complex (unknown origin) expressed in baculovirus infected Sf9 cells using H2N-RKQLATKAAR(Kme0)SAPA...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Homo sapiens (Human))
Constellation Pharmaceuticals
Curated by ChEMBL
Constellation Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 0.490nMAssay Description:Inhibition of N-terminal FLAG-tagged EZH2 in PRC2 complex (unknown origin) expressed in baculovirus infected Sf9 cells using H2N-RKQLATKAAR(Kme0)SAPA...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Homo sapiens (Human))
Constellation Pharmaceuticals
Curated by ChEMBL
Constellation Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 0.620nMAssay Description:Inhibition of N-terminal FLAG-tagged EZH2 in PRC2 complex (unknown origin) expressed in baculovirus infected Sf9 cells using H2N-RKQLATKAAR(Kme0)SAPA...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Homo sapiens (Human))
Constellation Pharmaceuticals
Curated by ChEMBL
Constellation Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 0.730nMAssay Description:Inhibition of N-terminal FLAG-tagged EZH2 in PRC2 complex (unknown origin) expressed in baculovirus infected Sf9 cells using H2N-RKQLATKAAR(Kme0)SAPA...More data for this Ligand-Target Pair
TargetDelta-type opioid receptor(MOUSE)
University Of Medicine And Dentistry Of New Jersey-Robert Wood Johnson Medical School And The Informatics Institute Of Umdnj
Curated by ChEMBL
University Of Medicine And Dentistry Of New Jersey-Robert Wood Johnson Medical School And The Informatics Institute Of Umdnj
Curated by ChEMBL
Affinity DataIC50: 0.800nMAssay Description:Displacement of [3H]bremazocine from mouse delta opioid receptor expressed in HEK293 cells by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataIC50: 1.60nMAssay Description:Binding affinity to recombinant His-tagged ERalpha LBD (307 to 554 residue) (unknown origin) expressed in Escherichia coli preincubated for 15 mins f...More data for this Ligand-Target Pair
Affinity DataIC50: 2.10nMAssay Description:Binding affinity to ERalpha S463P mutant (unknown origin) expressed in Escherichia coli preincubated for 15 mins followed by ligand addition and meas...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [588-1027,K691N,Y769C]/[588-1147,K691N,Y769C](Human immunodeficiency virus type 1)
Roche Palo Alto
Roche Palo Alto
Affinity DataIC50: 3nM EC50: 4nMpH: 8.0 T: 2°CAssay Description:IC50s were obtained from the inhibition of the RNA-dependent DNA polymerase activity of the HIV-1 reverse transcriptase enzyme using a primer extensi...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 5B(Homo sapiens (Human))
Constellation Pharmaceuticals
Curated by ChEMBL
Constellation Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 3nMAssay Description:Inhibition of KDM5B (unknown origin) using ART(Kme3)QTARKSTGGKAPRKQLA-NovaTagPEG-biotin/2-OG as substrate/co-factor by TR-FRET assayMore data for this Ligand-Target Pair
TargetLysine-specific demethylase 5B(Homo sapiens (Human))
Constellation Pharmaceuticals
Curated by ChEMBL
Constellation Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 3nMAssay Description:Inhibition of KDM5B (unknown origin) using ART(Kme3)QTARKSTGGKAPRKQLA-NovaTagPEG-biotin/2-OG as substrate/co-factor by TR-FRET assayMore data for this Ligand-Target Pair
TargetLysine-specific demethylase 5B(Homo sapiens (Human))
Constellation Pharmaceuticals
Curated by ChEMBL
Constellation Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 3nMpH: 7.0Assay Description:Time-resolved fluorescence resonance energy transfer (TR-FRET) assays were carried out using full-length KDM5 enzymes in 384-well black ProxiPlates (...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [588-1027](Human immunodeficiency virus type 1 group M subtyp...)
Roche Palo Alto
Roche Palo Alto
Affinity DataIC50: 3nM EC50: >25nMpH: 8.0 T: 2°CAssay Description:IC50s were obtained from the inhibition of the RNA-dependent DNA polymerase activity of the HIV-1 reverse transcriptase enzyme using a primer extensi...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [588-1027](Human immunodeficiency virus type 1 group M subtyp...)
Roche Palo Alto
Roche Palo Alto
Affinity DataIC50: 3nM EC50: 1nMpH: 8.0 T: 2°CAssay Description:IC50s were obtained from the inhibition of the RNA-dependent DNA polymerase activity of the HIV-1 reverse transcriptase enzyme using a primer extensi...More data for this Ligand-Target Pair
Affinity DataIC50: 3.20nMAssay Description:Binding affinity to ERalpha Y537S mutant (unknown origin) expressed in Escherichia coli preincubated for 15 mins followed by ligand addition and meas...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Homo sapiens (Human))
Constellation Pharmaceuticals
Curated by ChEMBL
Constellation Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 3.80nMAssay Description:Inhibition of N-terminal FLAG-tagged EZH2 in PRC2 complex (unknown origin) expressed in baculovirus infected Sf9 cells using H2N-RKQLATKAAR(Kme0)SAPA...More data for this Ligand-Target Pair

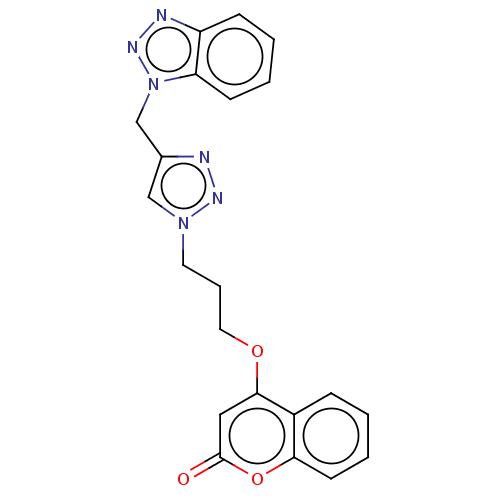

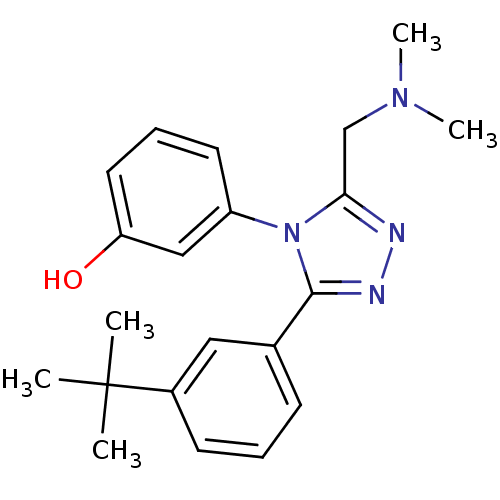
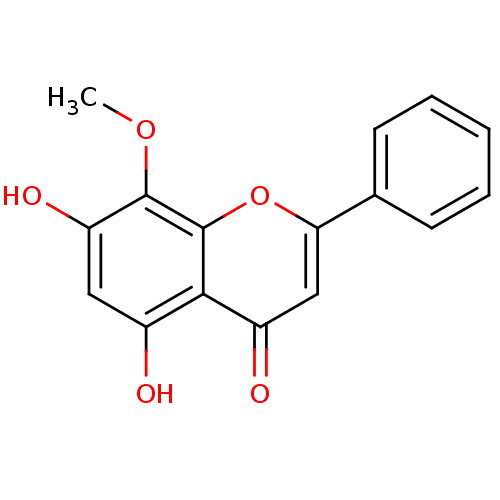

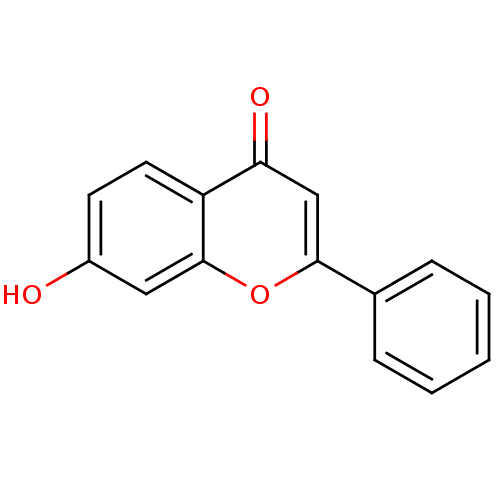
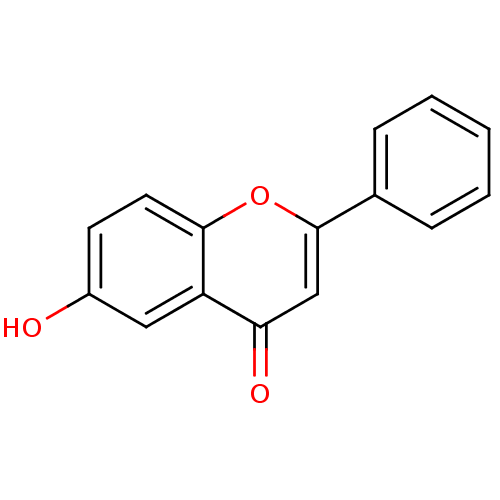
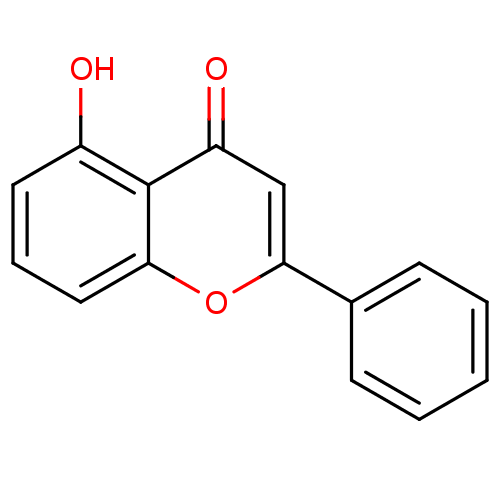
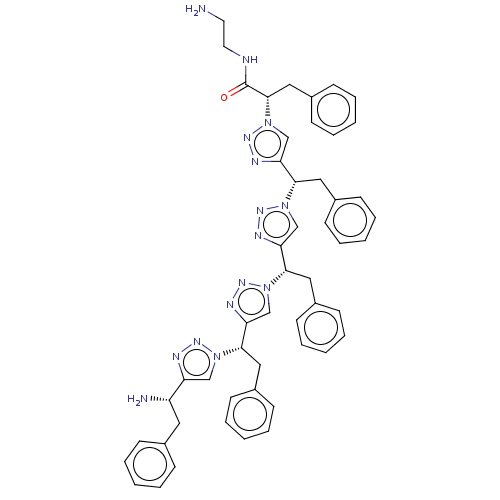
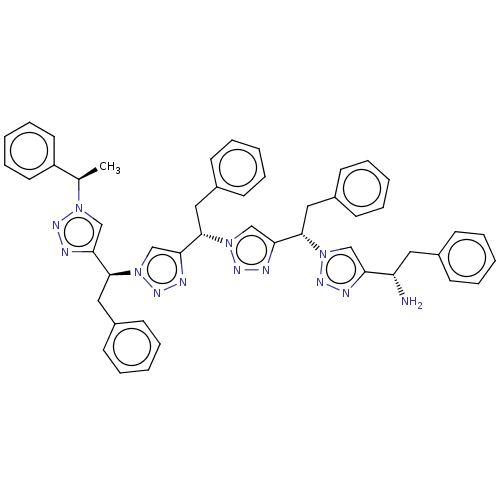
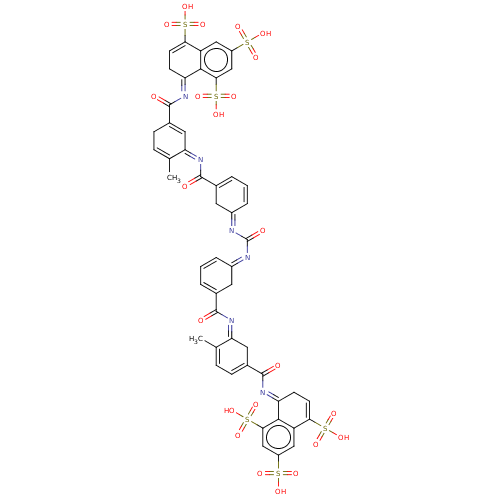
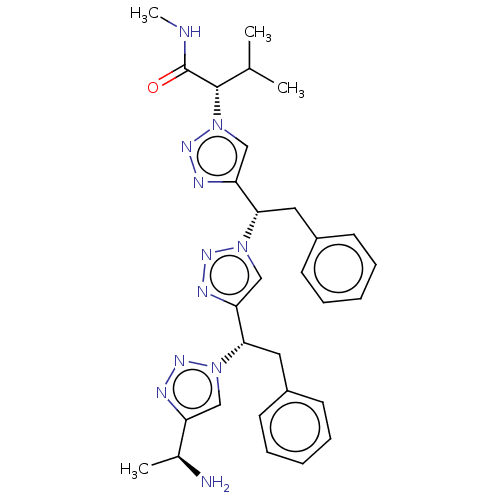
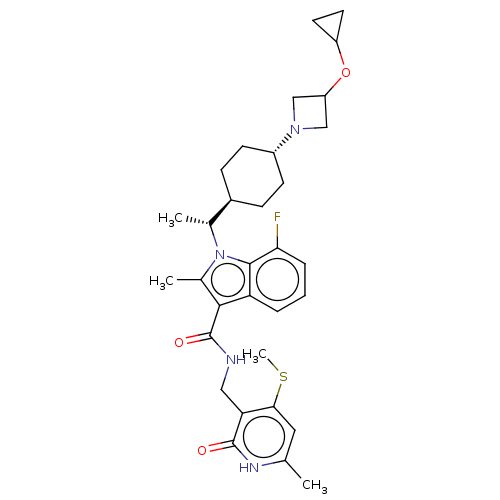
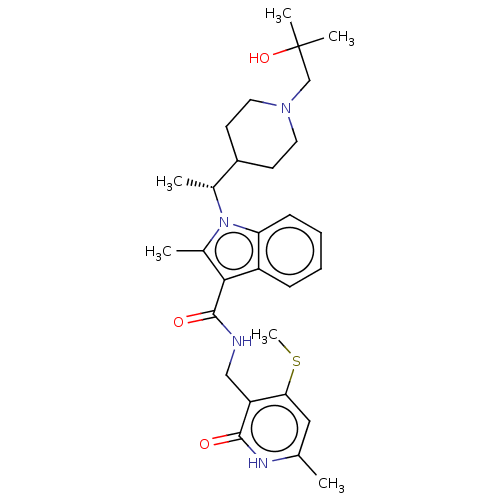
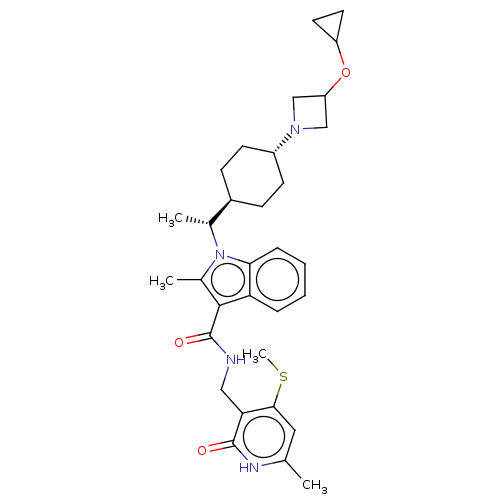

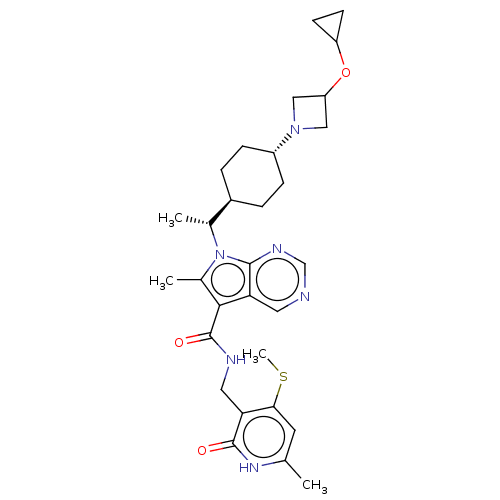
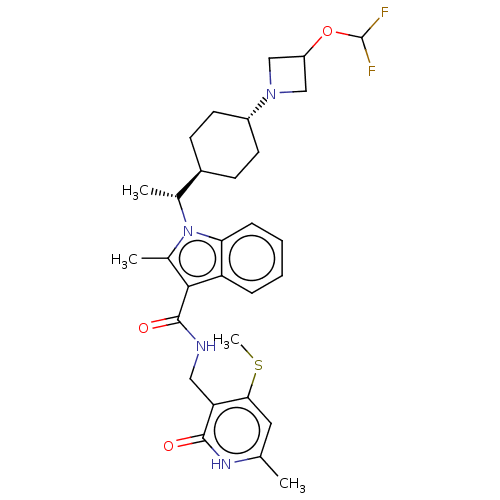
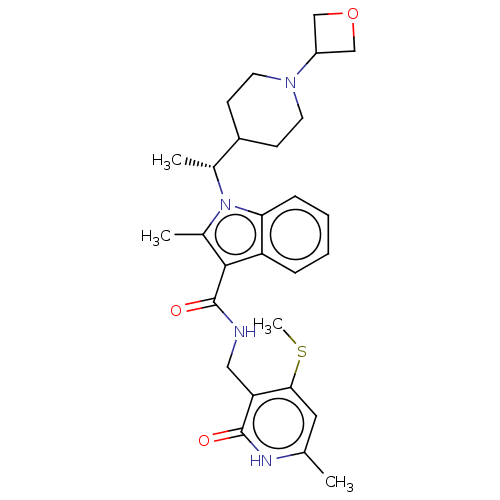
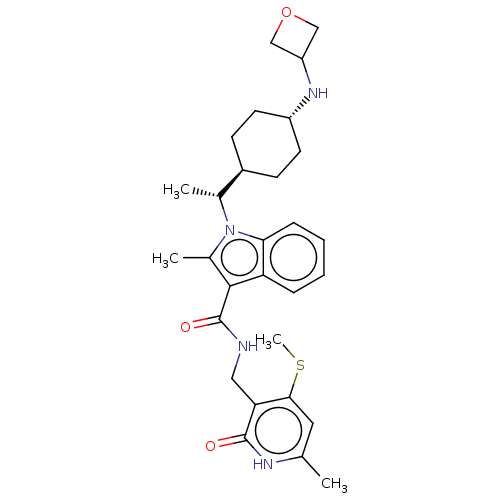
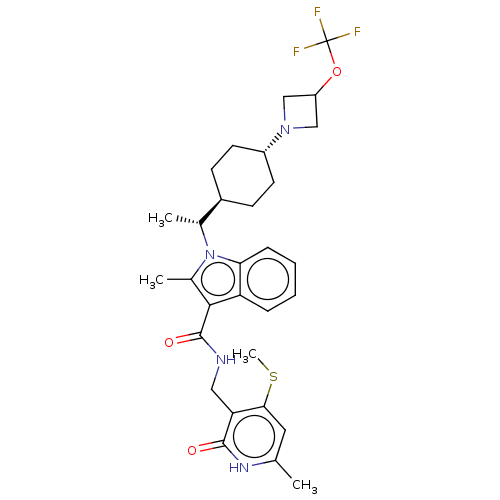
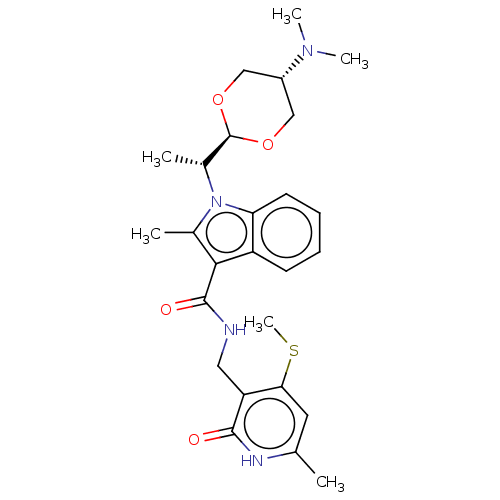
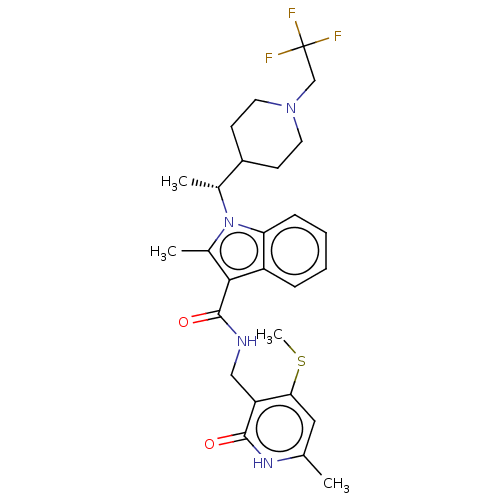
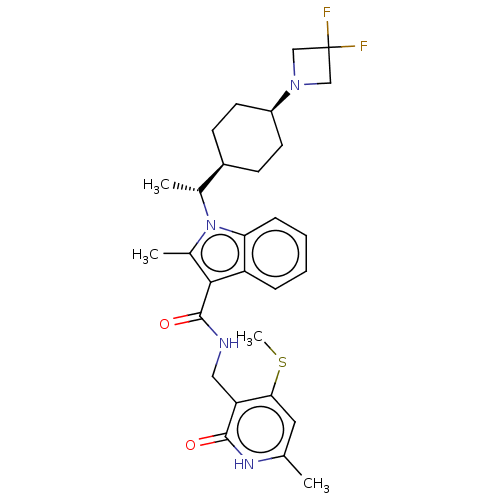
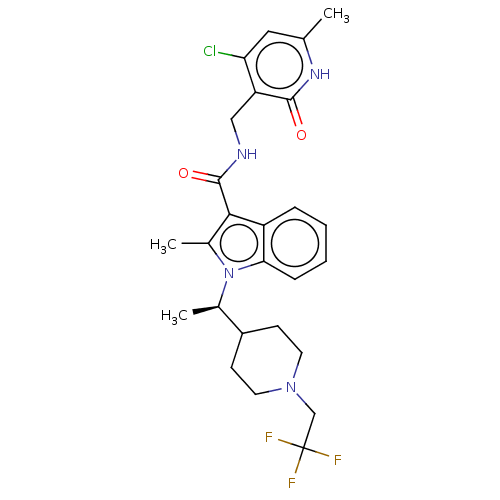
 3D Structure (crystal)
3D Structure (crystal)