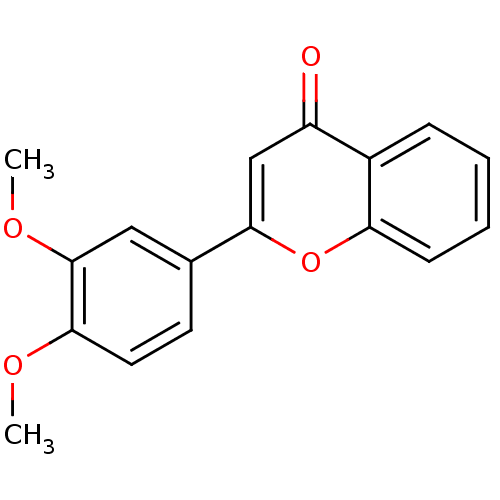
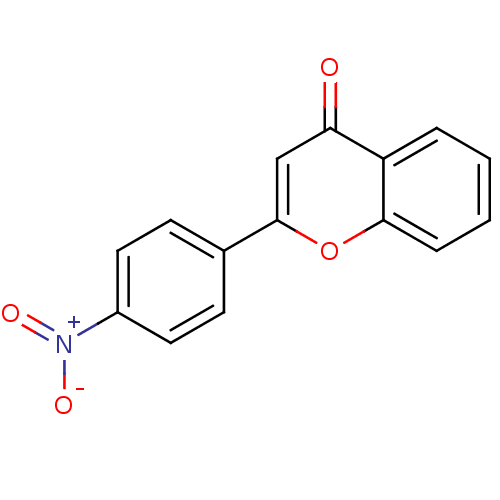
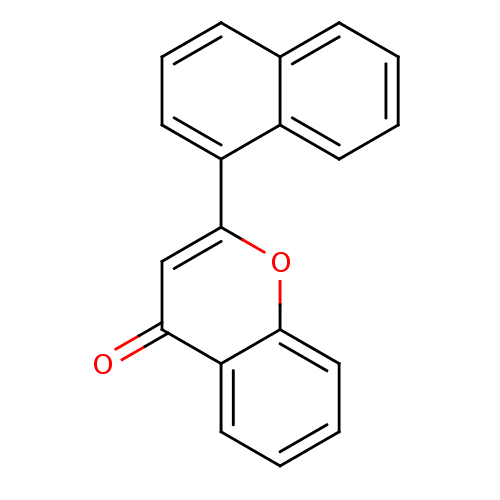
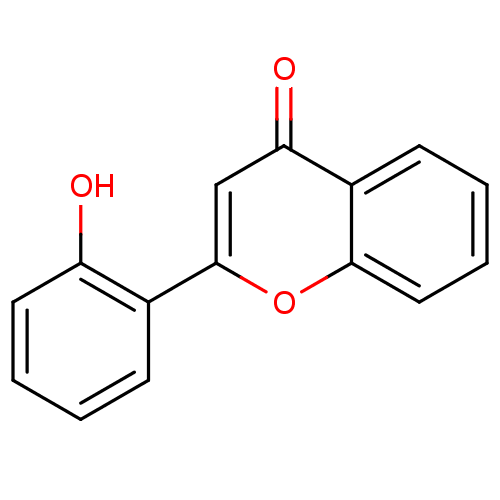
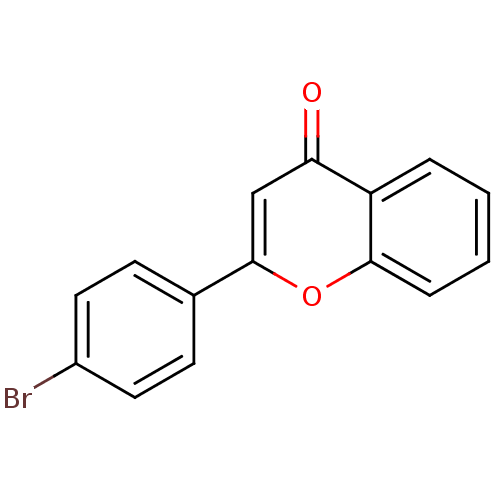
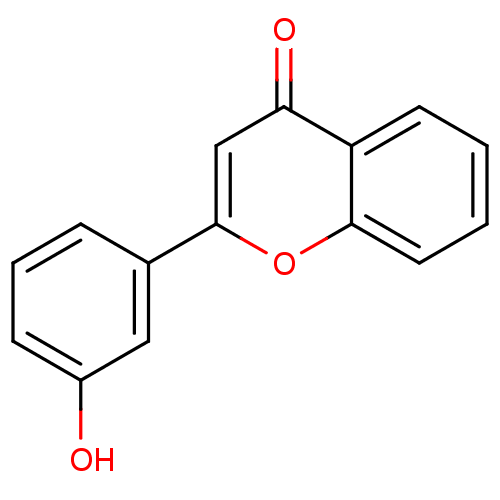
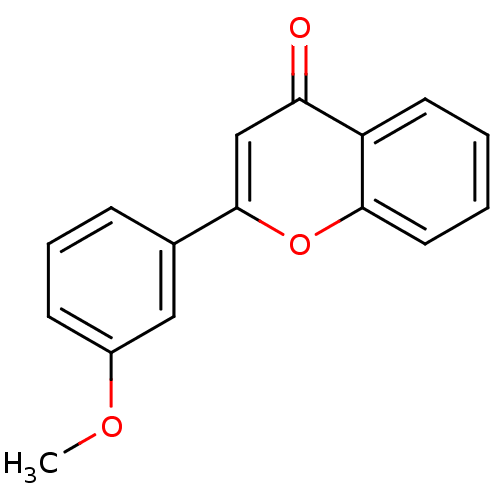
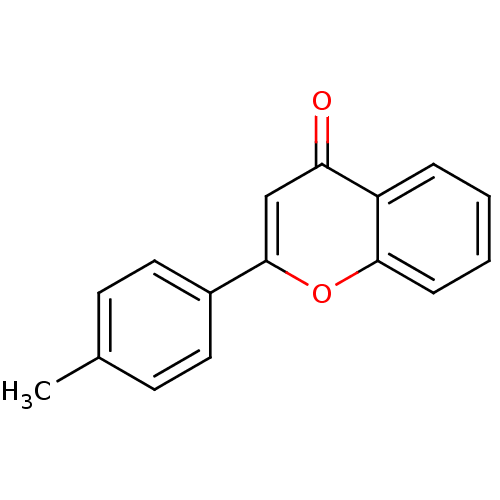
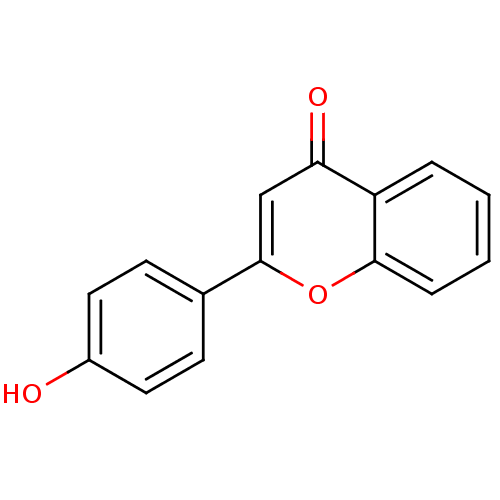
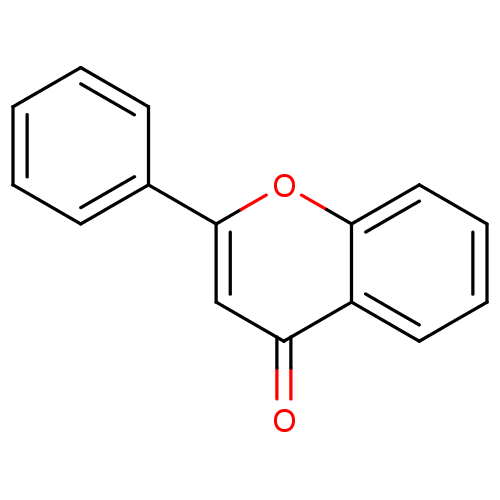
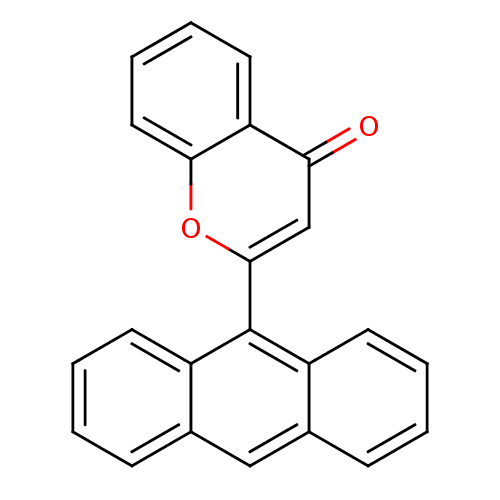
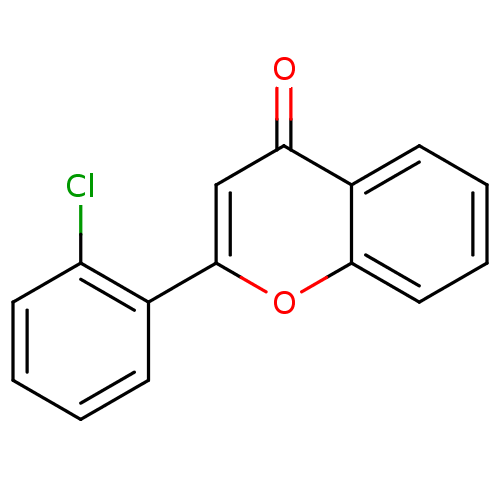
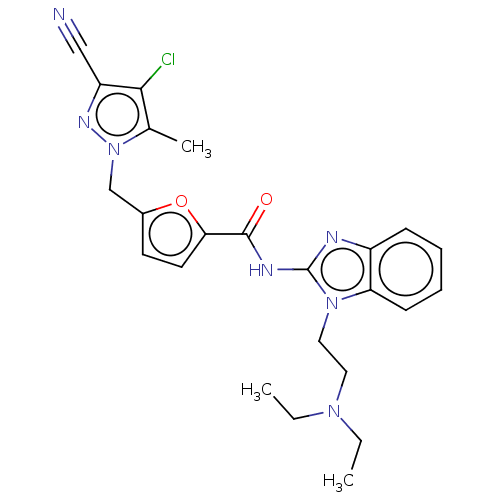
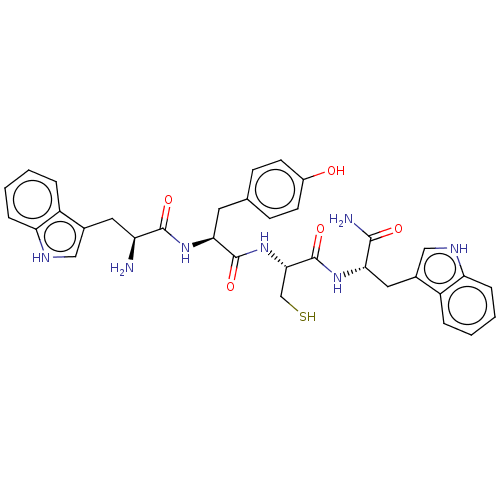
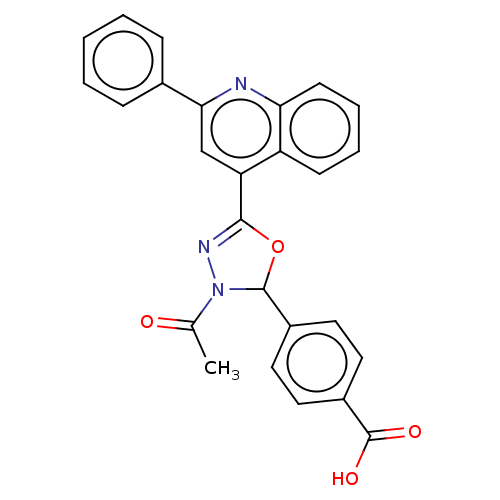
Affinity DataKi: 30nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 130nMAssay Description:Inhibition of West Nile virus NS2B-NS3 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 160nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 160nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 200nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 390nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 520nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 520nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 600nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 800nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 800nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 840nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 860nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 900nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 990nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 1.05E+3nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 1.06E+3nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 1.15E+3nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 1.20E+3nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 1.30E+3nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 1.40E+3nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 1.65E+3nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 1.80E+3nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 1.88E+3nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 1.90E+3nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 1.90E+3nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 1.99E+3nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 2.00E+3nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 2.00E+3nMAssay Description:Inhibition of West Nile virus NS2B-NS3 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 2.00E+3nMAssay Description:Inhibition of dengue virus-2 NS2B-NS3 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 2.50E+3nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 2.55E+3nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 2.64E+3nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 2.91E+3nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 3.02E+3nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 3.22E+3nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 3.32E+3nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 3.40E+3nMAssay Description:Inhibition of dengue virus NS2B-NS3 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 3.90E+3nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 4.00E+3nMAssay Description:Inhibition of dengue virus NS2B-NS3 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 4.20E+3nMAssay Description:Inhibition of dengue virus-1 NS2B-NS3 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 4.80E+3nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 4.80E+3nMAssay Description:Inhibition of dengue virus-2 NS2B-NS3 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 4.90E+3nMAssay Description:Inhibition of dengue virus NS2B-NS3 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 5.06E+3nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 5.70E+3nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 5.93E+3nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 5.98E+3nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 6.44E+3nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair
Affinity DataKi: 8.21E+3nMAssay Description:The activities of recombinant hMAO-A and hMAO-B were determined using p-tyramine as common substrate and calculated as 0.18 +/- 0.01 nmol/mg/min (n =...More data for this Ligand-Target Pair

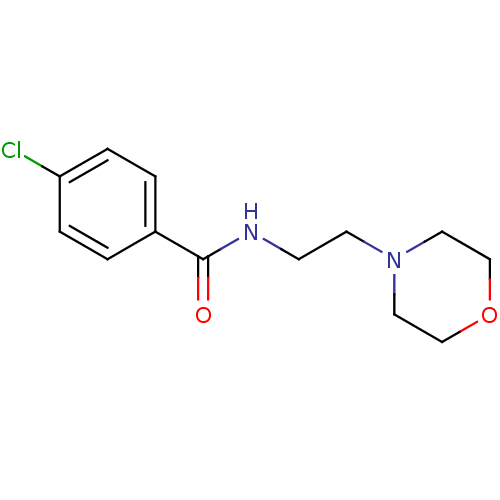
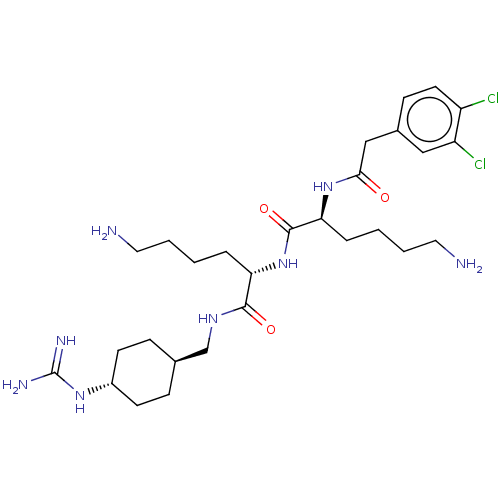
 3D Structure (crystal)
3D Structure (crystal)