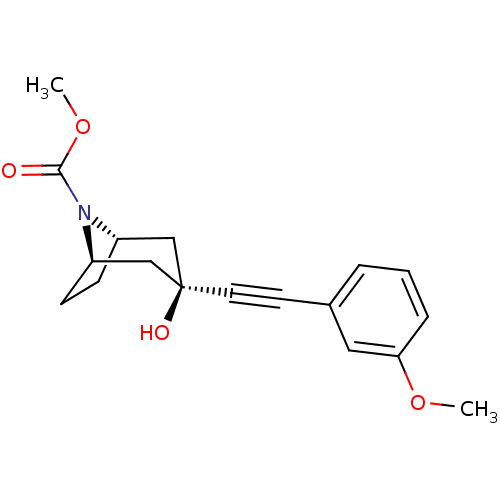
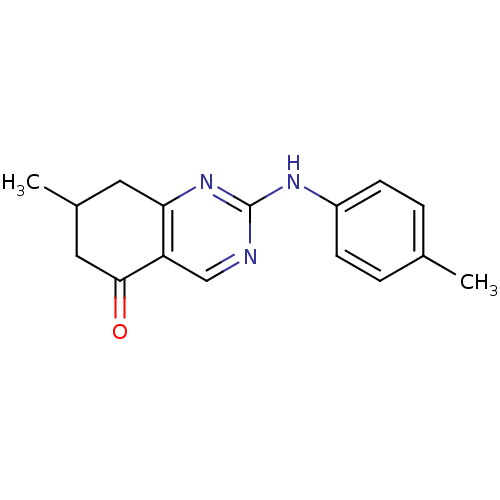
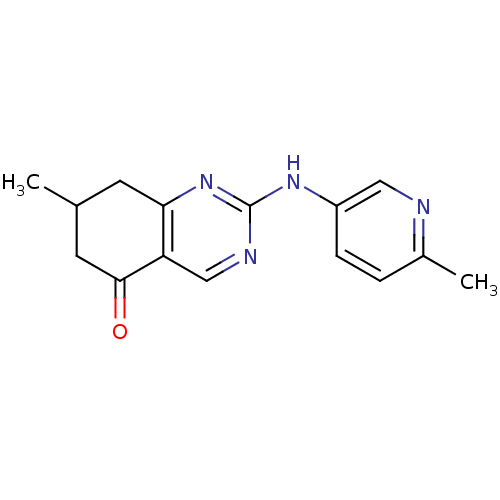
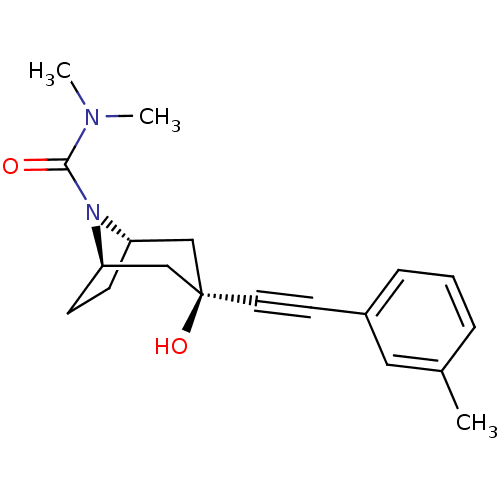
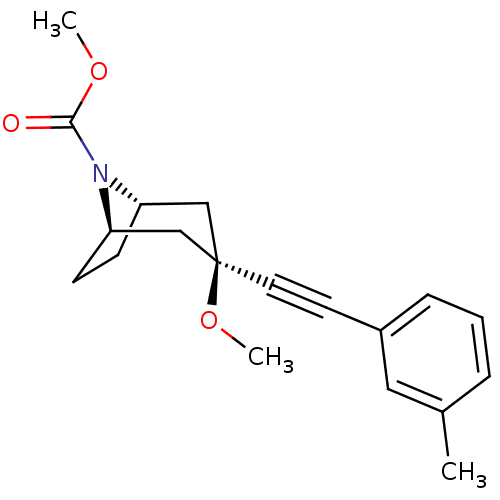
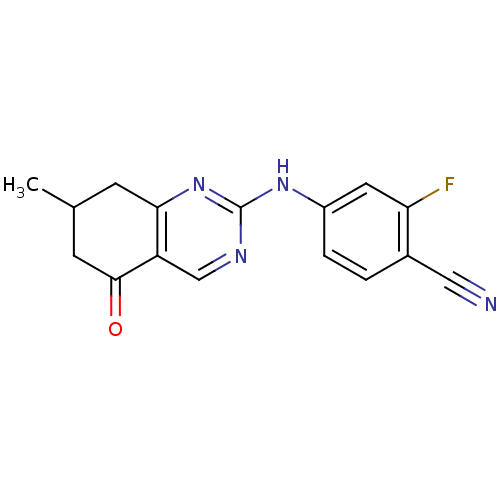
TargetMetabotropic glutamate receptor 5(Rattus norvegicus (Rat))
Institute Of Organic Synthesis
Curated by ChEMBL
Institute Of Organic Synthesis
Curated by ChEMBL
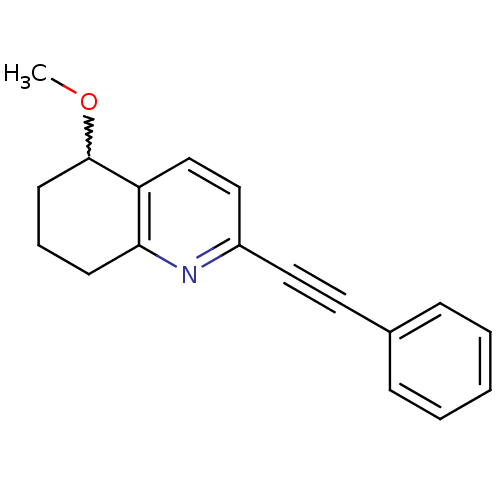
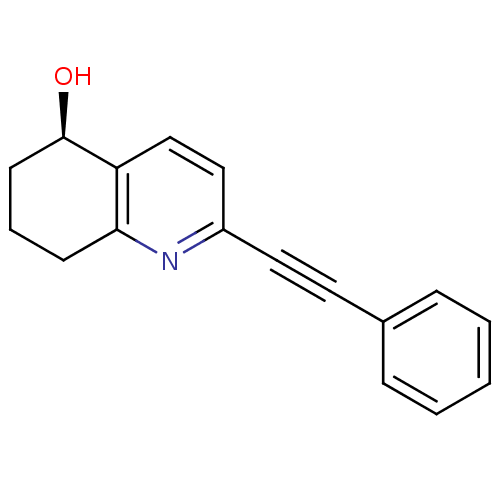
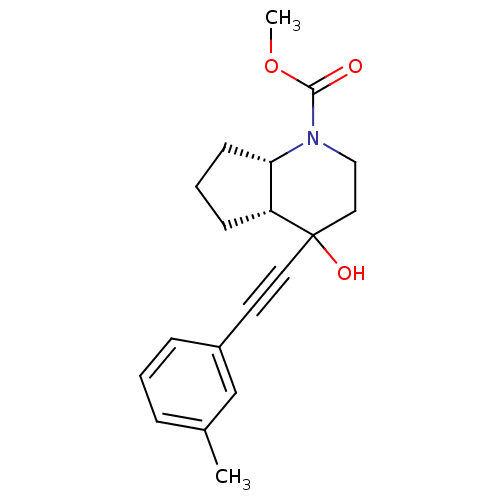
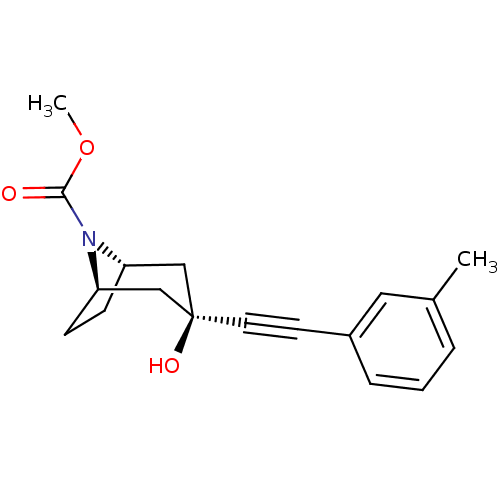
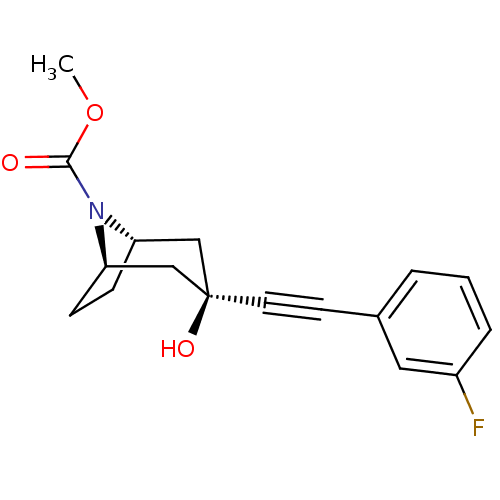
Affinity DataKi: 12nMAssay Description:Displacement of [3H]MPEP from rat mGluR5More data for this Ligand-Target Pair
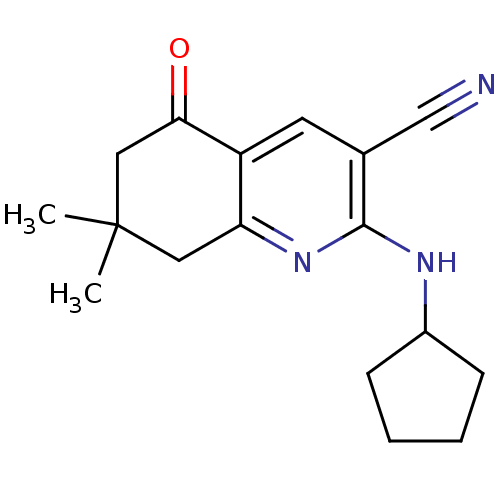
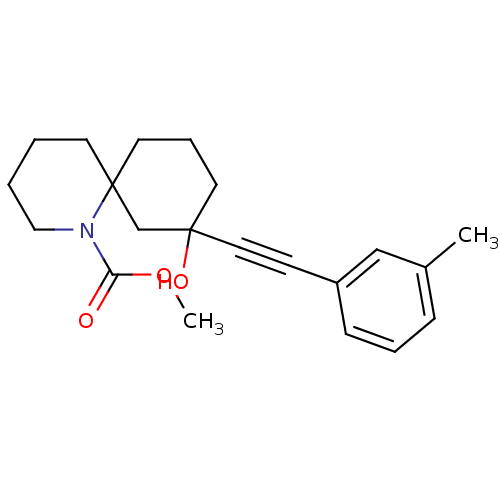
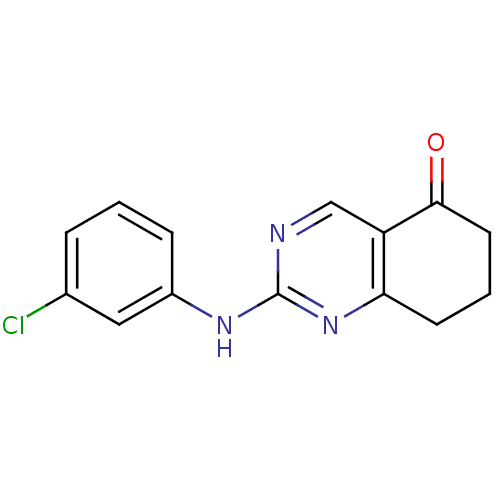
TargetMetabotropic glutamate receptor 5(Rattus norvegicus (Rat))
Institute Of Organic Synthesis
Curated by ChEMBL
Institute Of Organic Synthesis
Curated by ChEMBL
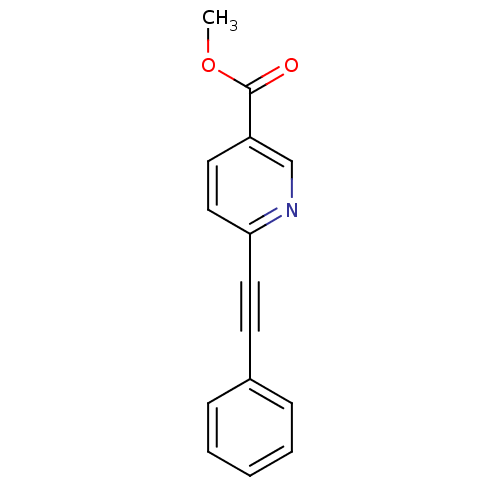
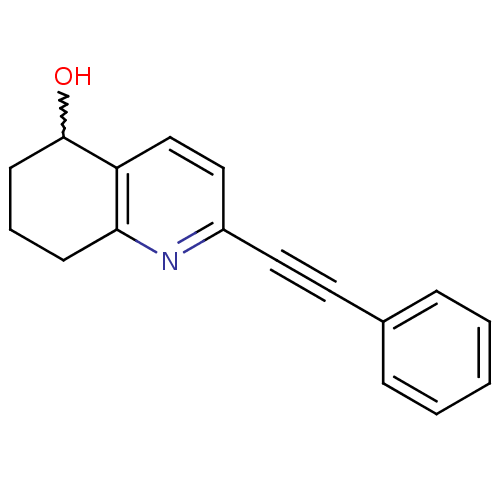
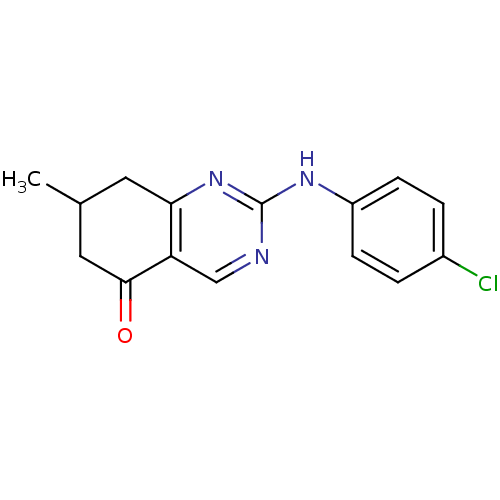
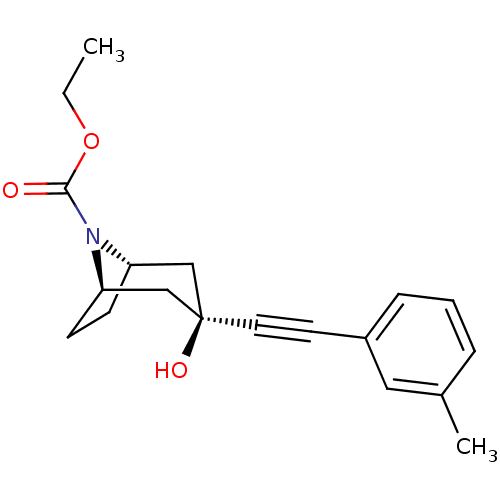
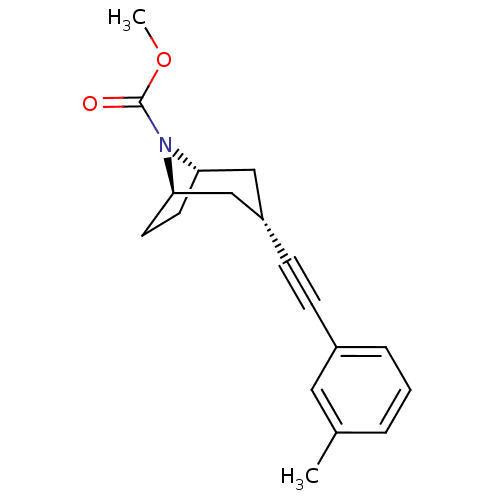
Affinity DataKi: 12nMAssay Description:Displacement of [3H]MPEP from rat mGluR5More data for this Ligand-Target Pair
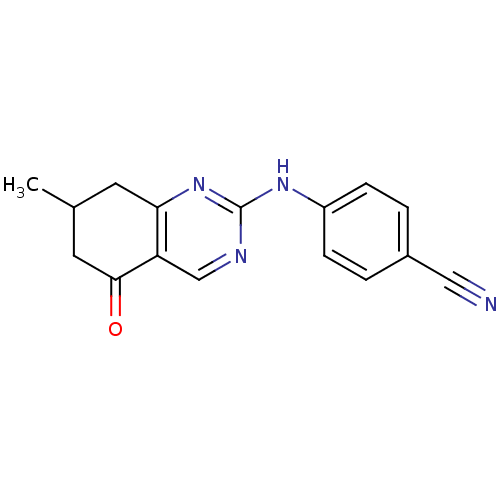
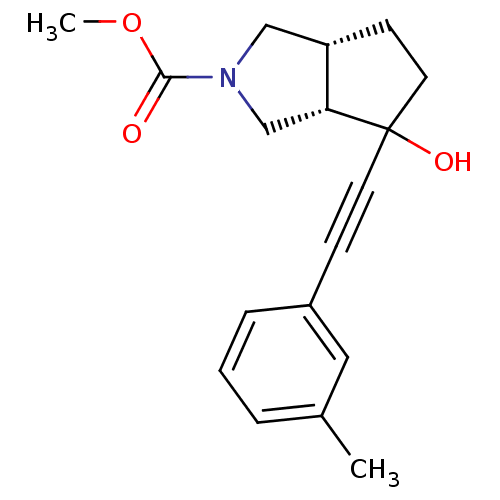
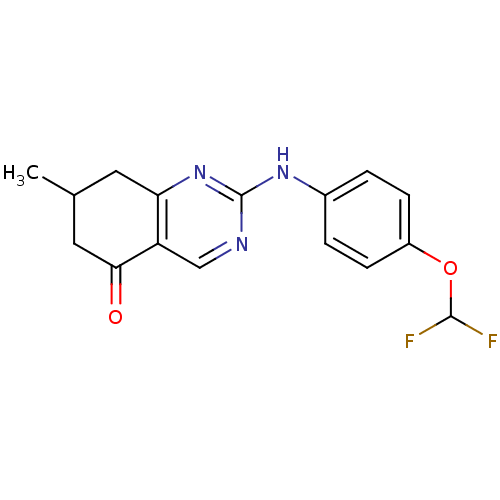
TargetMetabotropic glutamate receptor 5(Rattus norvegicus (Rat))
Institute Of Organic Synthesis
Curated by ChEMBL
Institute Of Organic Synthesis
Curated by ChEMBL
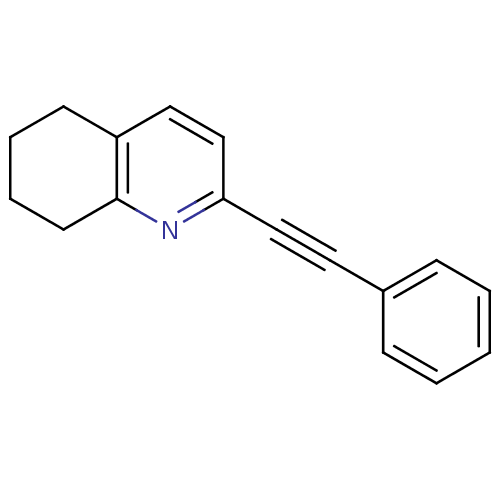
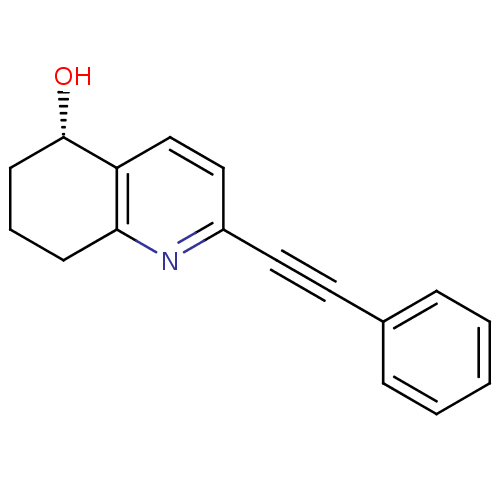
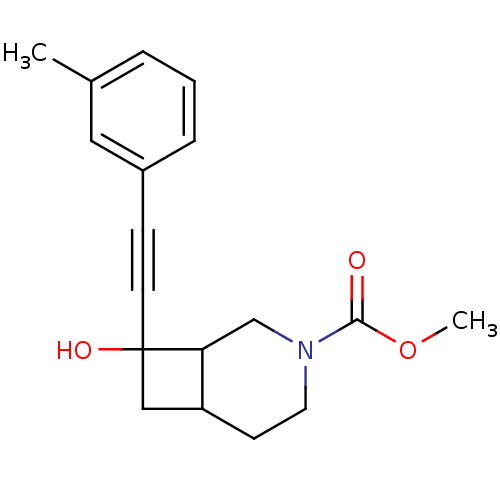
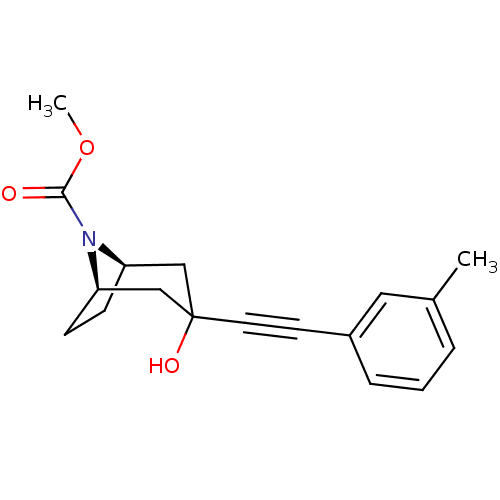
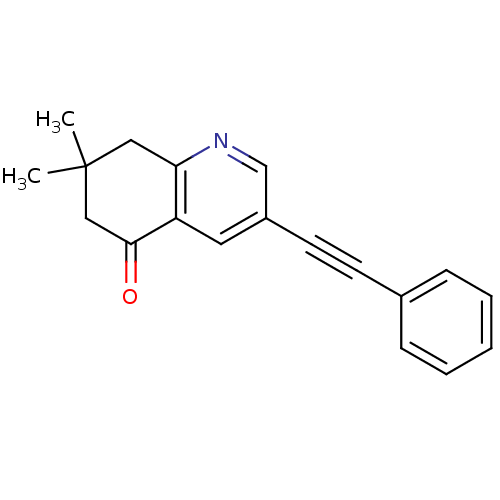
Affinity DataKi: 51nMAssay Description:Displacement of [3H]MPEP from rat mGluR5More data for this Ligand-Target Pair
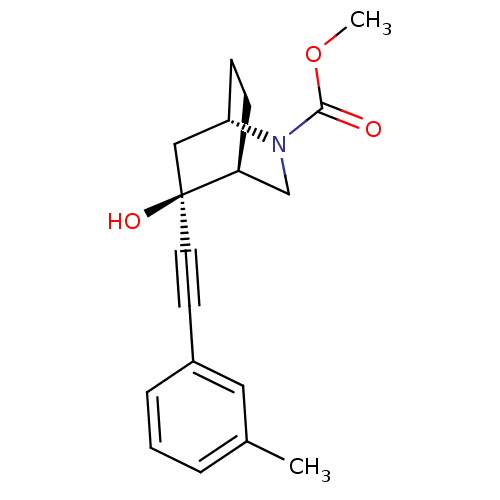
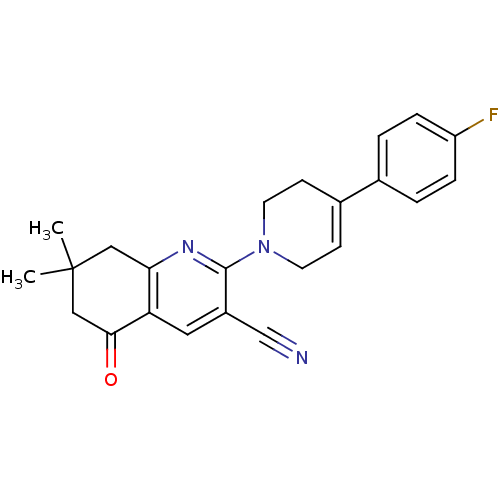
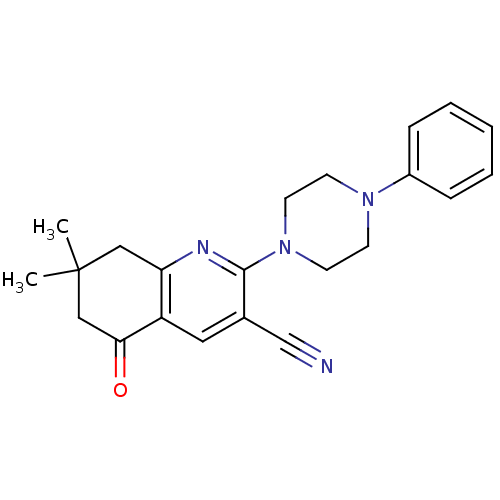
TargetMetabotropic glutamate receptor 5(Rattus norvegicus (Rat))
Institute Of Organic Synthesis
Curated by ChEMBL
Institute Of Organic Synthesis
Curated by ChEMBL
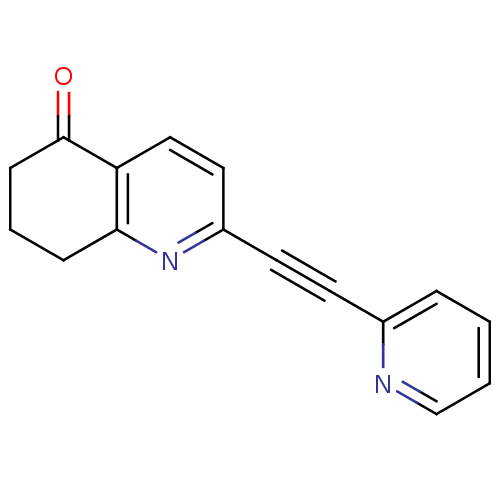
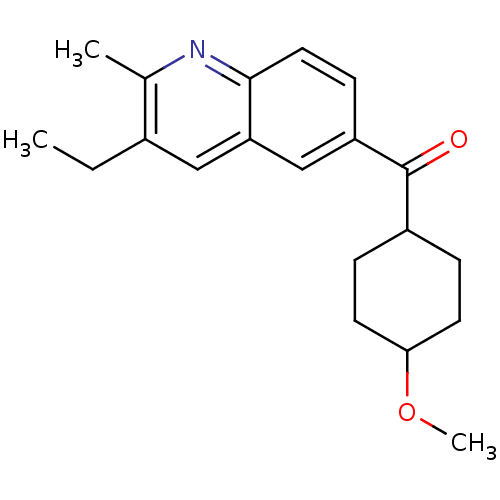
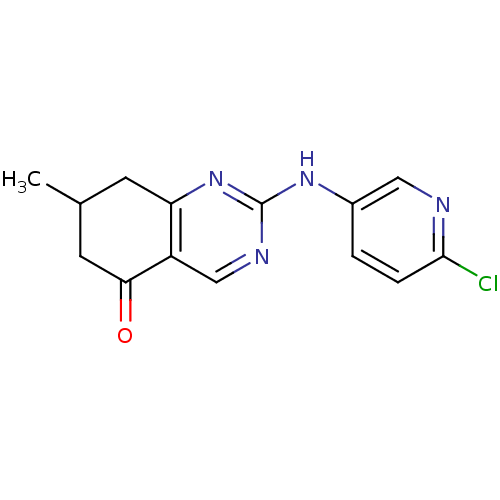
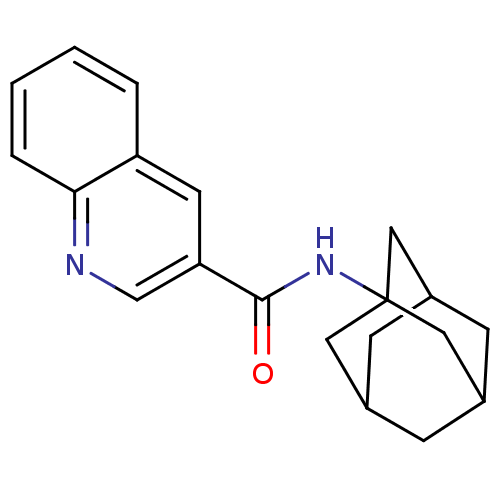
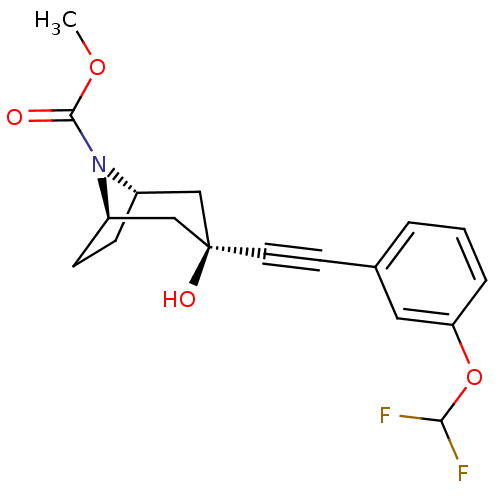
Affinity DataKi: 139nMAssay Description:Displacement of [3H]MPEP from rat mGluR5More data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 5(Rattus norvegicus (Rat))
Institute Of Organic Synthesis
Curated by ChEMBL
Institute Of Organic Synthesis
Curated by ChEMBL
Affinity DataKi: 210nMAssay Description:Displacement of [3H]MPEP from rat mGluR5More data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 5(Rattus norvegicus (Rat))
Institute Of Organic Synthesis
Curated by ChEMBL
Institute Of Organic Synthesis
Curated by ChEMBL
Affinity DataKi: 800nMAssay Description:Displacement of [3H]MPEP from rat mGluR5More data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 5(Rattus norvegicus (Rat))
Institute Of Organic Synthesis
Curated by ChEMBL
Institute Of Organic Synthesis
Curated by ChEMBL
Affinity DataKi: 1.40E+3nMAssay Description:Displacement of [3H]MPEP from rat mGluR5More data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 5(Rattus norvegicus (Rat))
Institute Of Organic Synthesis
Curated by ChEMBL
Institute Of Organic Synthesis
Curated by ChEMBL
Affinity DataKi: 1.50E+3nMAssay Description:Displacement of [3H]MPEP from rat mGluR5More data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 5(Rattus norvegicus (Rat))
Institute Of Organic Synthesis
Curated by ChEMBL
Institute Of Organic Synthesis
Curated by ChEMBL
Affinity DataKi: 2.36E+3nMAssay Description:Displacement of [3H]MPEP from rat mGluR5More data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 5(Rattus norvegicus (Rat))
Institute Of Organic Synthesis
Curated by ChEMBL
Institute Of Organic Synthesis
Curated by ChEMBL
Affinity DataKi: 4.71E+3nMAssay Description:Displacement of [3H]MPEP from rat mGluR5More data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 5(Rattus norvegicus (Rat))
Institute Of Organic Synthesis
Curated by ChEMBL
Institute Of Organic Synthesis
Curated by ChEMBL
Affinity DataKi: 7.60E+3nMAssay Description:Displacement of [3H]MPEP from rat mGluR5More data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 5(Rattus norvegicus (Rat))
Institute Of Organic Synthesis
Curated by ChEMBL
Institute Of Organic Synthesis
Curated by ChEMBL
Affinity DataKi: 9.28E+3nMAssay Description:Displacement of [3H]MPEP from rat mGluR5More data for this Ligand-Target Pair
Affinity DataIC50: 3nMAssay Description:Antagonist activity at rat mGluR1 expressed in cerebellar granule cells assessed as accumulation of [3H]inositol phosphateMore data for this Ligand-Target Pair
Affinity DataIC50: 5nMAssay Description:Antagonist activity at rat mGluR1 expressed in cerebellar granule cells assessed as accumulation of [3H]inositol phosphateMore data for this Ligand-Target Pair
Affinity DataIC50: 6nMAssay Description:Antagonist activity at rat mGluR1 expressed in cerebellar granule cells assessed as accumulation of [3H]inositol phosphateMore data for this Ligand-Target Pair
Affinity DataIC50: 23nMAssay Description:Negative allosteric modulation of human mGluR5 expressed in CHO cells assessed as inhibition of L-quisqualate-induced intracellular cAMP accumulation...More data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 5(Rattus norvegicus (Rat))
Institute Of Organic Synthesis
Curated by ChEMBL
Institute Of Organic Synthesis
Curated by ChEMBL
Affinity DataIC50: 23nMAssay Description:Displacement of [3H]MPEP from rat mGluR5More data for this Ligand-Target Pair
Affinity DataIC50: 27nMAssay Description:Negative allosteric modulation of human mGlu5 receptor expressed in CHO cells assessed as inhibition of L-quisqualate-induced intracellular calcium m...More data for this Ligand-Target Pair
Affinity DataIC50: 28nMAssay Description:Negative allosteric modulation of human mGluR5 expressed in CHO cells assessed as inhibition of L-quisqualate-induced intracellular cAMP accumulation...More data for this Ligand-Target Pair
Affinity DataIC50: 28nMAssay Description:Negative allosteric modulation of human mGlu5 receptor expressed in CHO cells assessed as inhibition of L-quisqualate-induced intracellular calcium m...More data for this Ligand-Target Pair
Affinity DataIC50: 28.4nMAssay Description:Negative allosteric modulation of human mGlu5 receptor expressed in CHO cells assessed as inhibition of L-quisqualate-induced intracellular calcium m...More data for this Ligand-Target Pair
Affinity DataIC50: 34nMAssay Description:Negative allosteric modulation of human mGluR5 expressed in CHO cells assessed as inhibition of L-quisqualate-induced intracellular cAMP accumulation...More data for this Ligand-Target Pair
Affinity DataIC50: 37nMAssay Description:Negative allosteric modulation of human mGlu5 receptor expressed in CHO cells assessed as inhibition of L-quisqualate-induced intracellular calcium m...More data for this Ligand-Target Pair
Affinity DataIC50: 43nMAssay Description:Negative allosteric modulation of human mGluR5 expressed in CHO cells assessed as inhibition of L-quisqualate-induced intracellular cAMP accumulation...More data for this Ligand-Target Pair
Affinity DataIC50: 47nMAssay Description:Negative allosteric modulation of human mGlu5 receptor expressed in CHO cells assessed as inhibition of L-quisqualate-induced intracellular calcium m...More data for this Ligand-Target Pair
Affinity DataIC50: 48nMAssay Description:Negative allosteric modulation of human mGlu5 receptor expressed in CHO cells assessed as inhibition of L-quisqualate-induced intracellular calcium m...More data for this Ligand-Target Pair
Affinity DataIC50: 51nMAssay Description:Negative allosteric modulation of human mGlu5 receptor expressed in CHO cells assessed as inhibition of L-quisqualate-induced intracellular calcium m...More data for this Ligand-Target Pair
Affinity DataIC50: 52nMAssay Description:Antagonist activity at rat mGluR1 expressed in cerebellar granule cells assessed as accumulation of [3H]inositol phosphateMore data for this Ligand-Target Pair
Affinity DataIC50: 61nMAssay Description:Negative allosteric modulation of human mGlu5 receptor expressed in CHO cells assessed as inhibition of L-quisqualate-induced intracellular calcium m...More data for this Ligand-Target Pair
Affinity DataIC50: 65nMAssay Description:Negative allosteric modulation of human mGlu5 receptor expressed in CHO cells assessed as inhibition of L-quisqualate-induced intracellular calcium m...More data for this Ligand-Target Pair
Affinity DataIC50: 68nMAssay Description:Negative allosteric modulation of human mGluR5 expressed in CHO cells assessed as inhibition of L-quisqualate-induced intracellular cAMP accumulation...More data for this Ligand-Target Pair
Affinity DataIC50: 72nMAssay Description:Negative allosteric modulation of human mGlu5 receptor expressed in CHO cells assessed as inhibition of L-quisqualate-induced intracellular calcium m...More data for this Ligand-Target Pair
Affinity DataIC50: 72nMAssay Description:Negative allosteric modulation of human mGlu5 receptor expressed in CHO cells assessed as inhibition of L-quisqualate-induced intracellular calcium m...More data for this Ligand-Target Pair
Affinity DataIC50: 76nMAssay Description:Negative allosteric modulation of human mGlu5 receptor expressed in CHO cells assessed as inhibition of L-quisqualate-induced intracellular calcium m...More data for this Ligand-Target Pair
Affinity DataIC50: 78nMAssay Description:Antagonist activity at rat mGluR1 expressed in cerebellar granule cells assessed as accumulation of [3H]inositol phosphateMore data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 5(Rattus norvegicus (Rat))
Institute Of Organic Synthesis
Curated by ChEMBL
Institute Of Organic Synthesis
Curated by ChEMBL
Affinity DataIC50: 80nMAssay Description:Displacement of [3H]MPEP from rat mGluR5More data for this Ligand-Target Pair
Affinity DataIC50: 83nMAssay Description:Negative allosteric modulation of human mGluR5 expressed in CHO cells assessed as inhibition of L-quisqualate-induced intracellular cAMP accumulation...More data for this Ligand-Target Pair
Affinity DataIC50: 85nMAssay Description:Negative allosteric modulation of human mGlu5 receptor expressed in CHO cells assessed as inhibition of L-quisqualate-induced intracellular calcium m...More data for this Ligand-Target Pair
Affinity DataIC50: 85nMAssay Description:Negative allosteric modulation of human mGluR5 expressed in CHO cells assessed as inhibition of L-quisqualate-induced intracellular cAMP accumulation...More data for this Ligand-Target Pair
Affinity DataIC50: 86nMAssay Description:Negative allosteric modulation of human mGlu5 receptor expressed in CHO cells assessed as inhibition of L-quisqualate-induced intracellular calcium m...More data for this Ligand-Target Pair
Affinity DataIC50: 103nMAssay Description:Negative allosteric modulation of human mGlu5 receptor expressed in CHO cells assessed as inhibition of L-quisqualate-induced intracellular calcium m...More data for this Ligand-Target Pair
Affinity DataIC50: 103nMAssay Description:Negative allosteric modulation of human mGlu5 receptor expressed in CHO cells assessed as inhibition of L-quisqualate-induced intracellular calcium m...More data for this Ligand-Target Pair
Affinity DataIC50: 110nMAssay Description:Antagonist activity at rat mGluR1 expressed in cerebellar granule cells assessed as accumulation of [3H]inositol phosphateMore data for this Ligand-Target Pair
Affinity DataIC50: 111nMAssay Description:Negative allosteric modulation of human mGluR5 expressed in CHO cells assessed as inhibition of L-quisqualate-induced intracellular cAMP accumulation...More data for this Ligand-Target Pair
Affinity DataIC50: 120nMAssay Description:Negative allosteric modulation of human mGluR5 expressed in CHO cells assessed as inhibition of L-quisqualate-induced intracellular cAMP accumulation...More data for this Ligand-Target Pair
Affinity DataIC50: 125nMAssay Description:Negative allosteric modulation of human mGluR5 expressed in CHO cells assessed as inhibition of L-quisqualate-induced intracellular cAMP accumulation...More data for this Ligand-Target Pair
Affinity DataIC50: 130nMAssay Description:Antagonist activity at rat mGluR1 expressed in cerebellar granule cells assessed as accumulation of [3H]inositol phosphateMore data for this Ligand-Target Pair
Affinity DataIC50: 132nMAssay Description:Negative allosteric modulation of human mGlu5 receptor expressed in CHO cells assessed as inhibition of L-quisqualate-induced intracellular calcium m...More data for this Ligand-Target Pair
Affinity DataIC50: 137nMAssay Description:Negative allosteric modulation of human mGlu5 receptor expressed in CHO cells assessed as inhibition of L-quisqualate-induced intracellular calcium m...More data for this Ligand-Target Pair
Affinity DataIC50: 157nMAssay Description:Negative allosteric modulation of human mGluR5 expressed in CHO cells assessed as inhibition of L-quisqualate-induced intracellular cAMP accumulation...More data for this Ligand-Target Pair

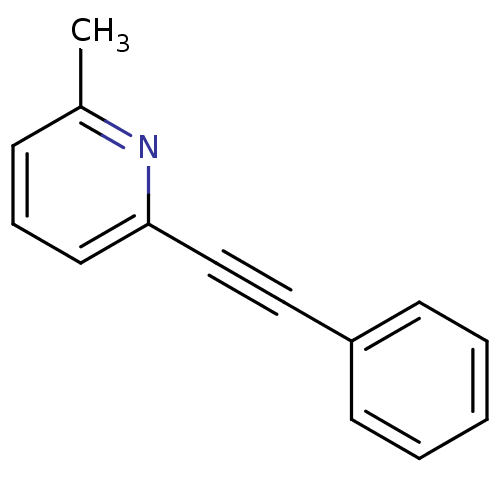



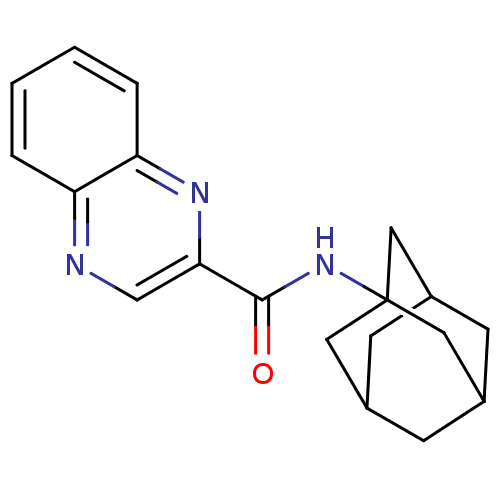
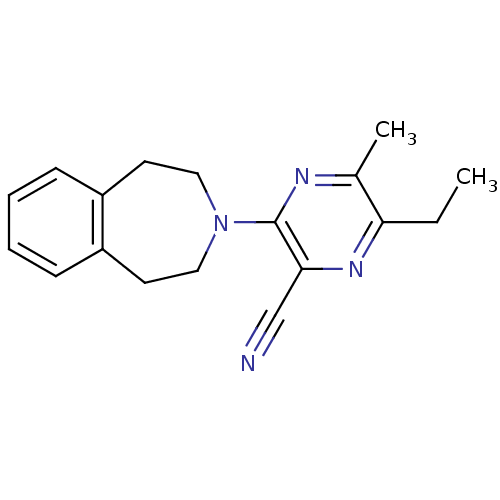
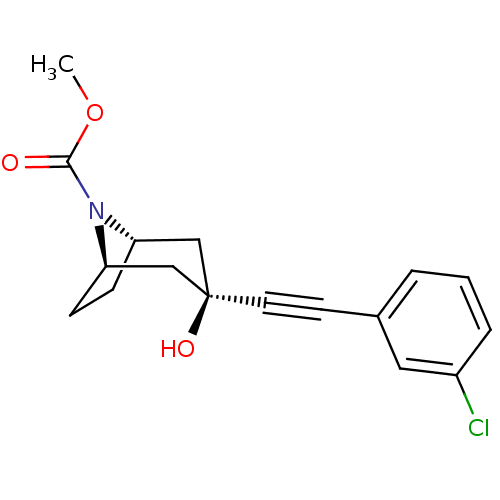
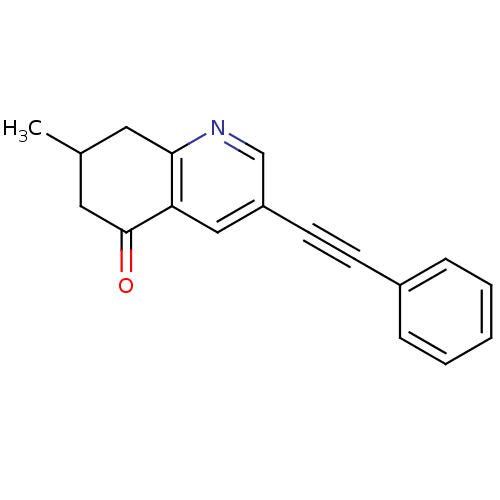
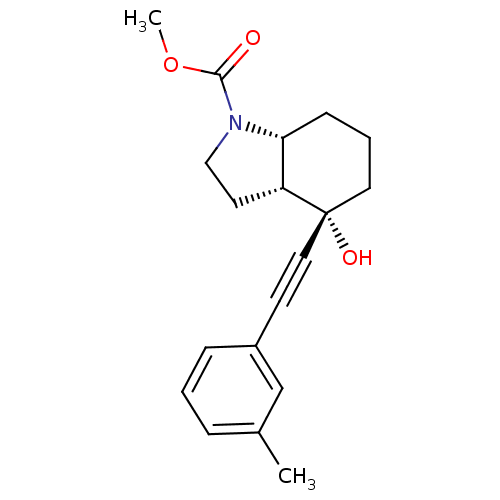
 3D Structure (crystal)
3D Structure (crystal)