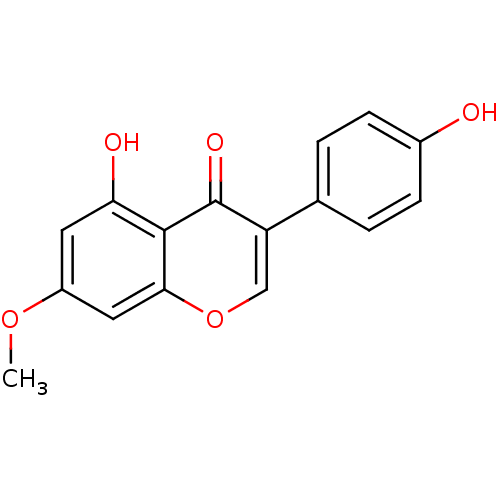
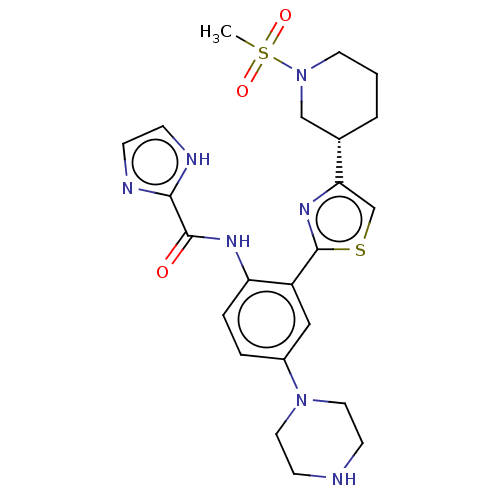
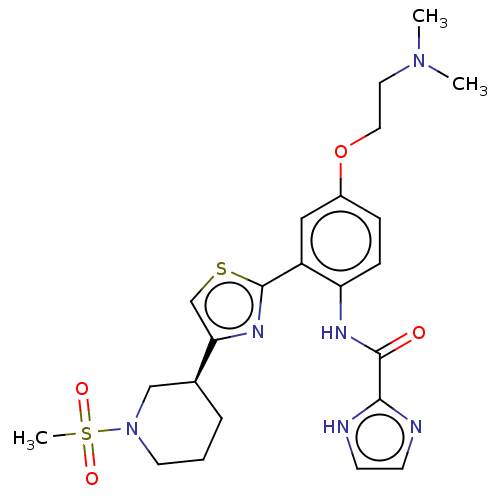
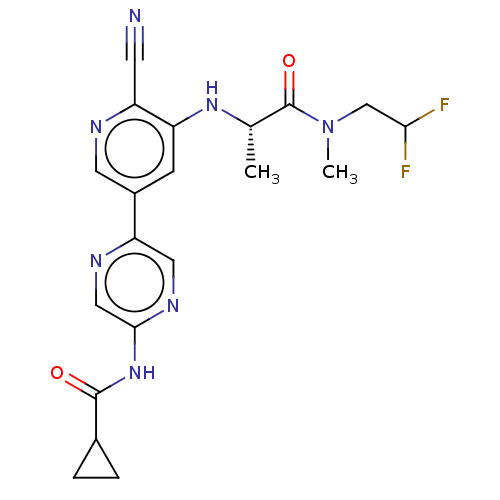
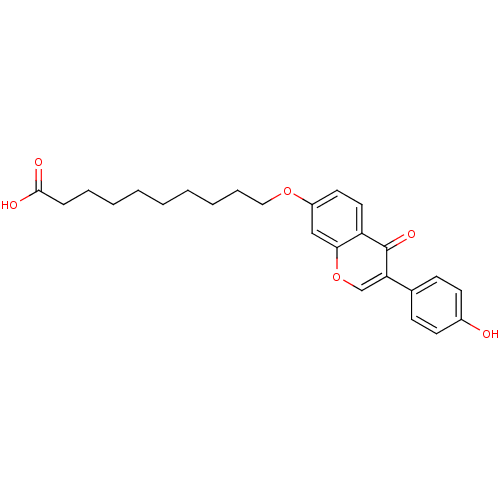
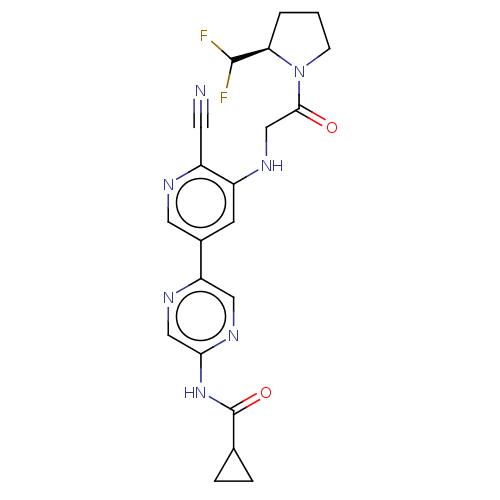
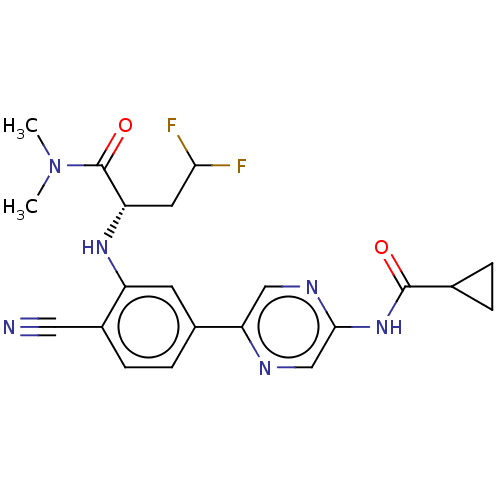
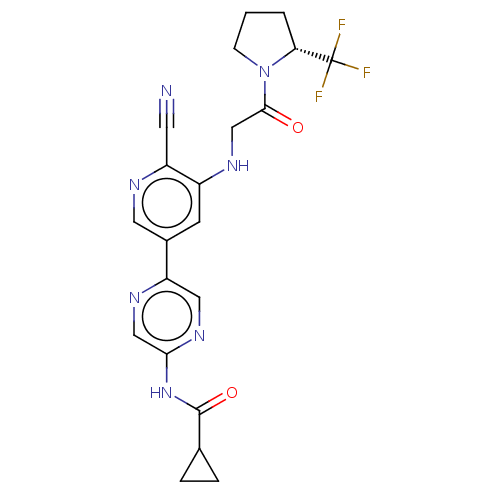
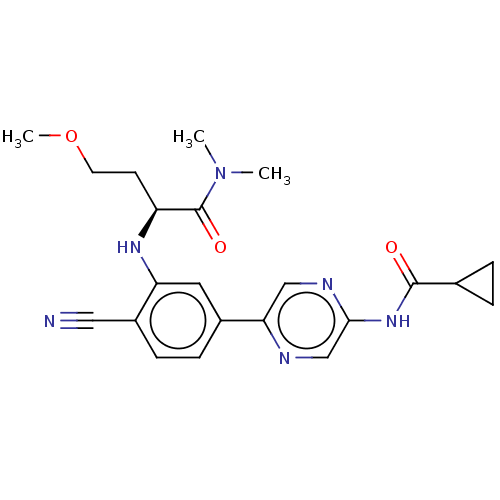
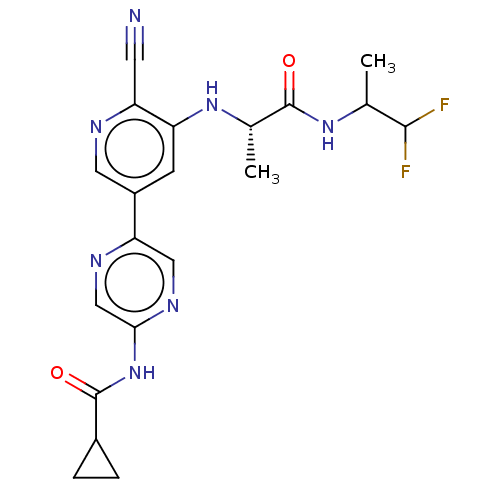
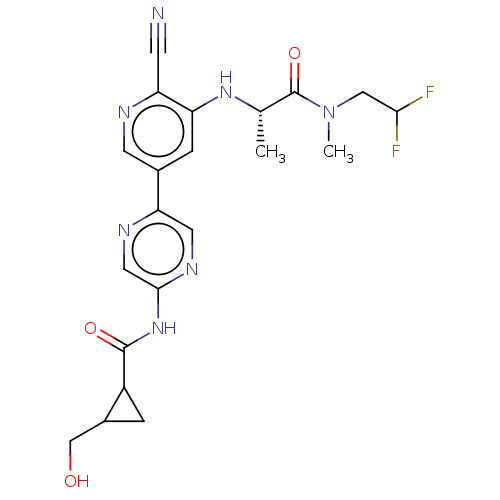
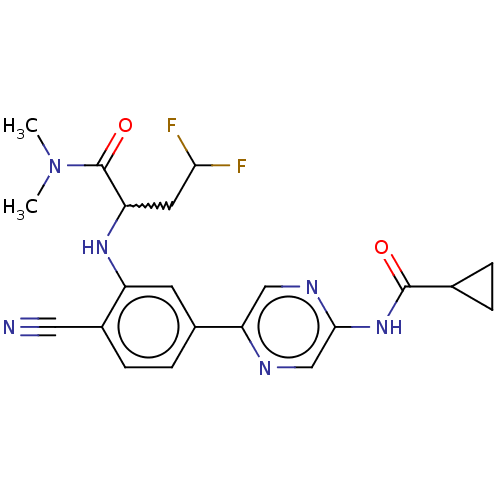
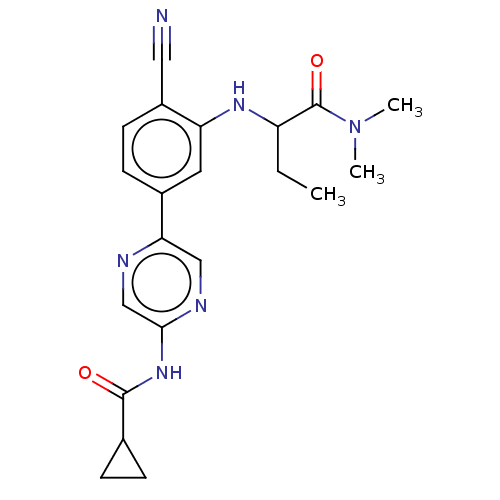
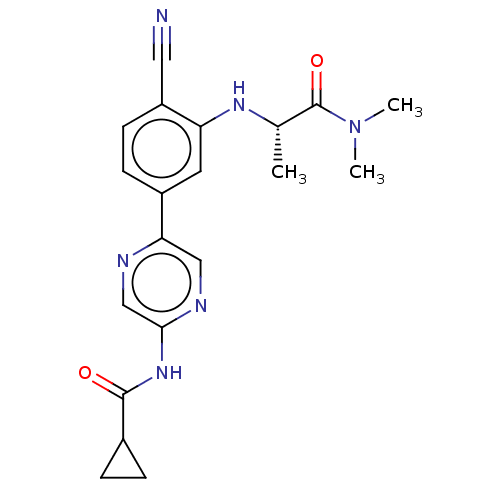
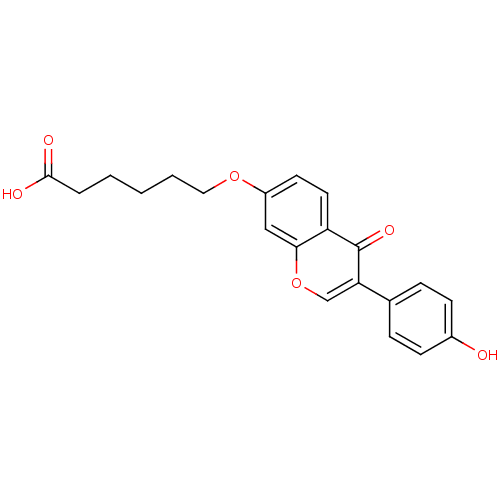
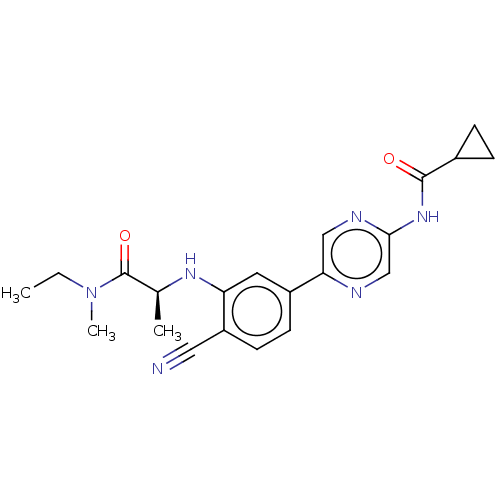
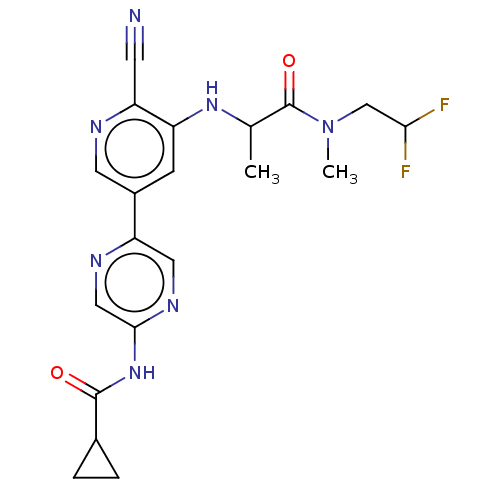
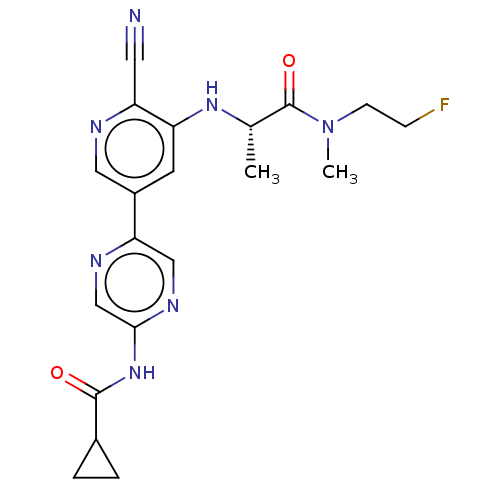
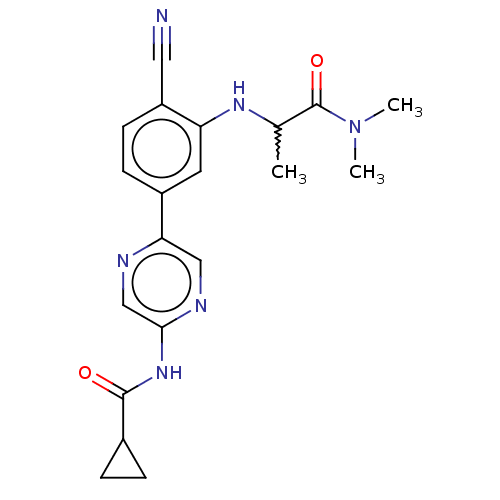
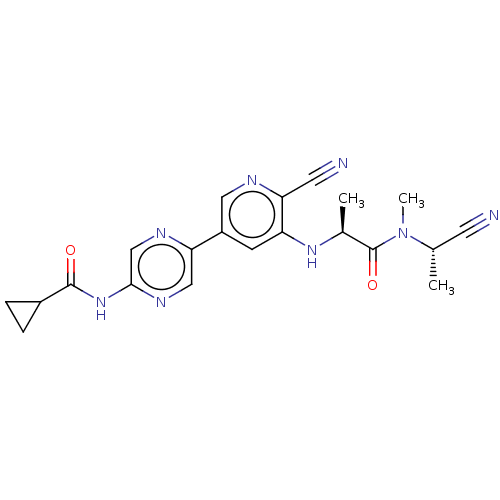
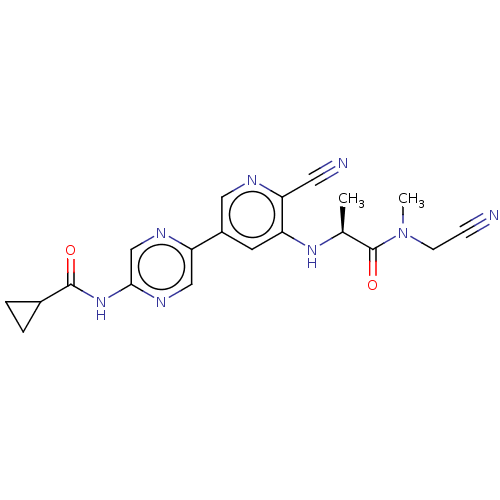
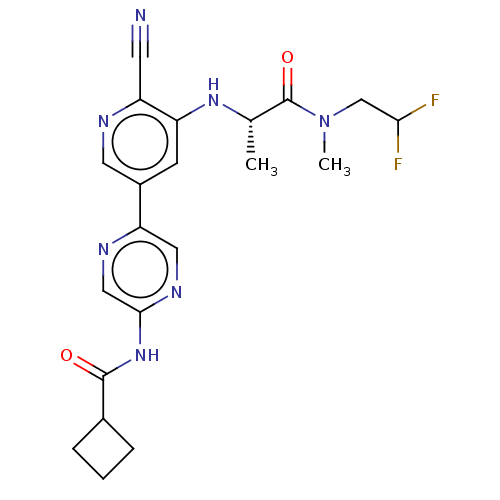
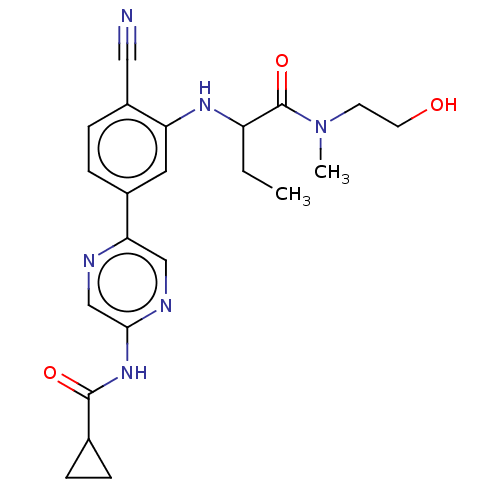
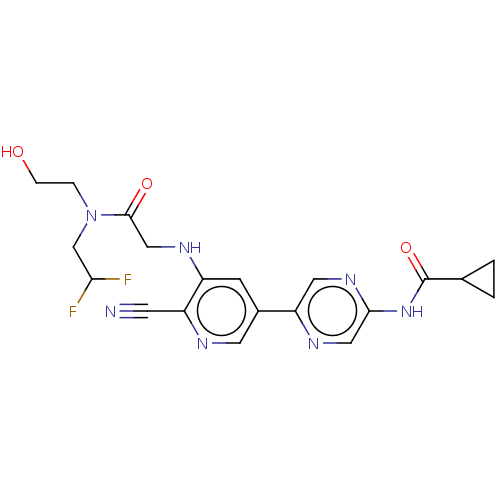
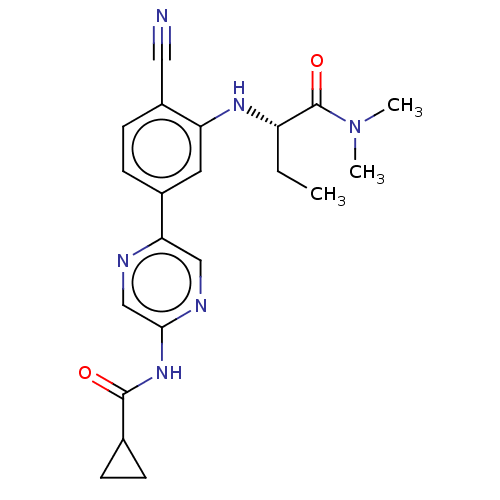
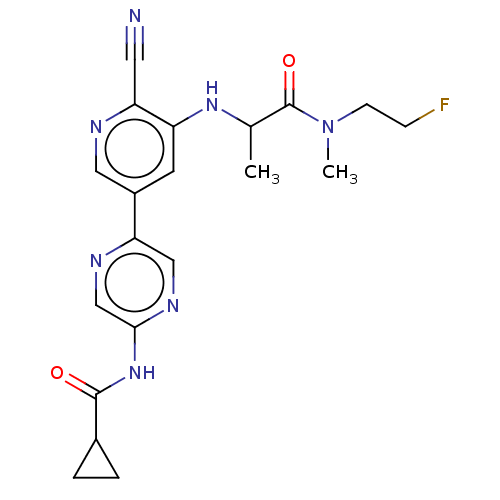
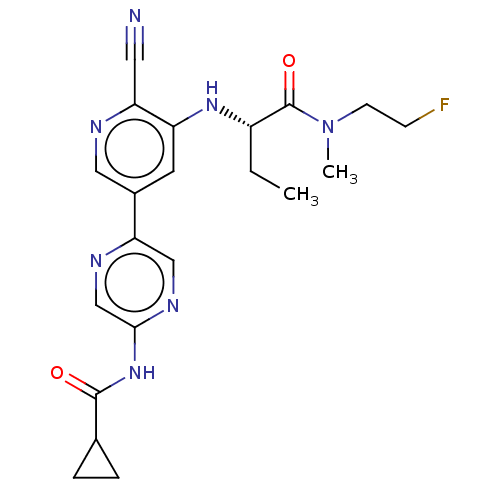
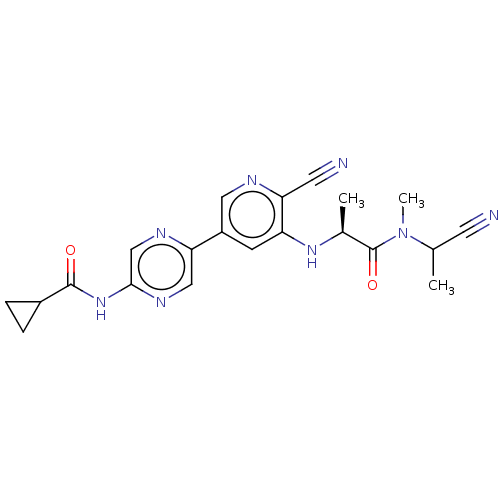
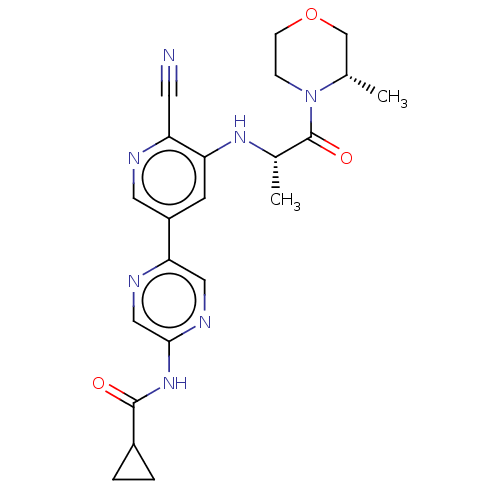
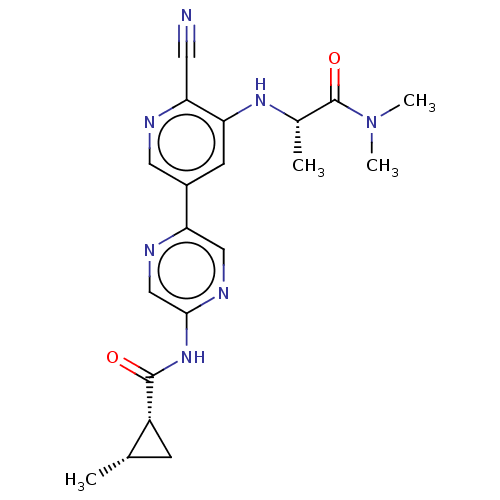
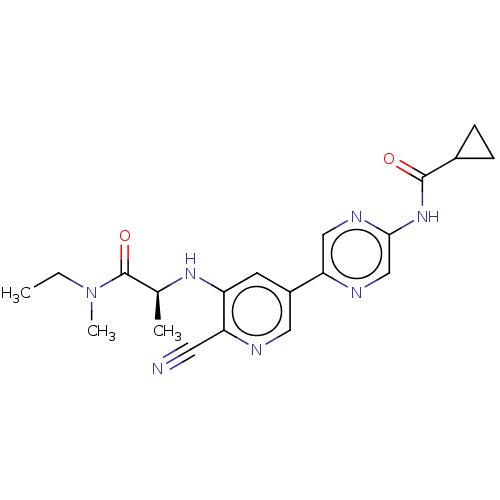
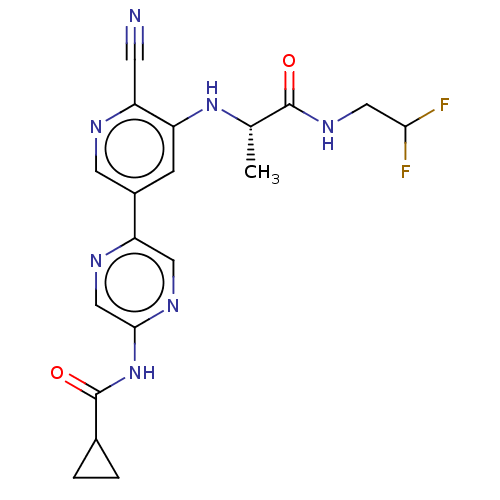
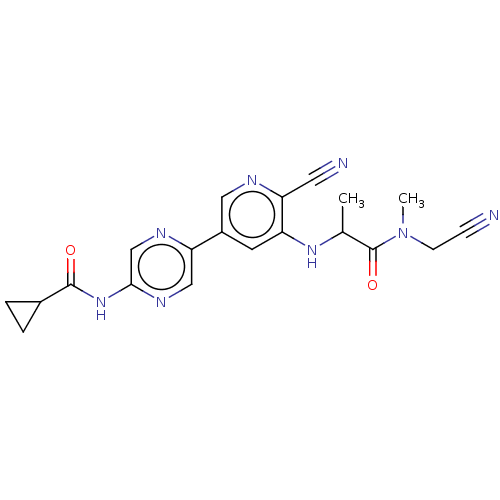
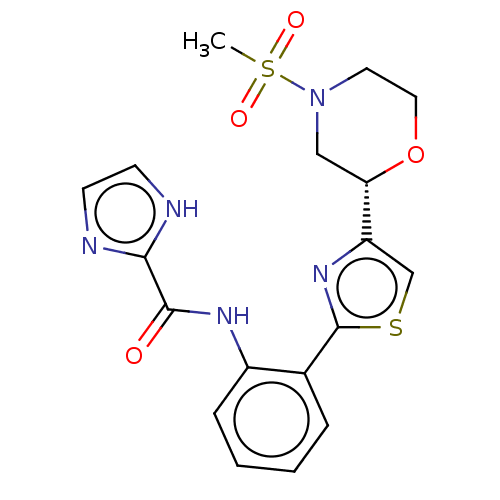
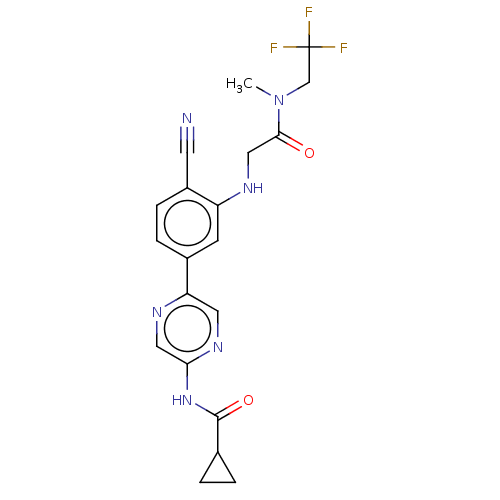
TargetAldehyde dehydrogenase, mitochondrial(Homo sapiens (Human))
University Of Oxford
Curated by ChEMBL
University Of Oxford
Curated by ChEMBL
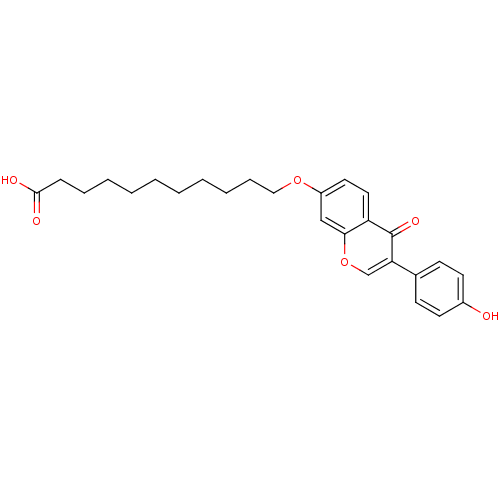
Affinity DataKi: 450nMAssay Description:Inhibition of human recombinant ALDH2More data for this Ligand-Target Pair
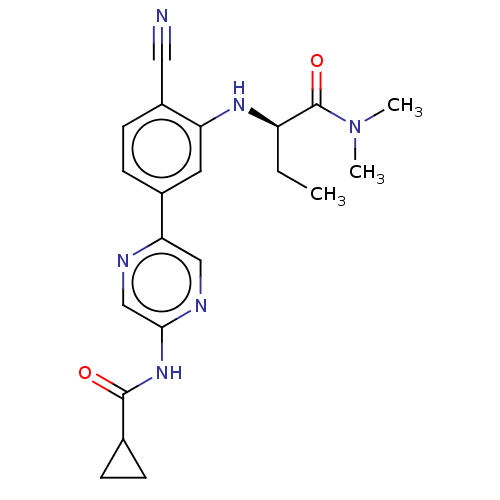
Affinity DataIC50: 1nMAssay Description:Inhibition of Axl (unknown origin)More data for this Ligand-Target Pair
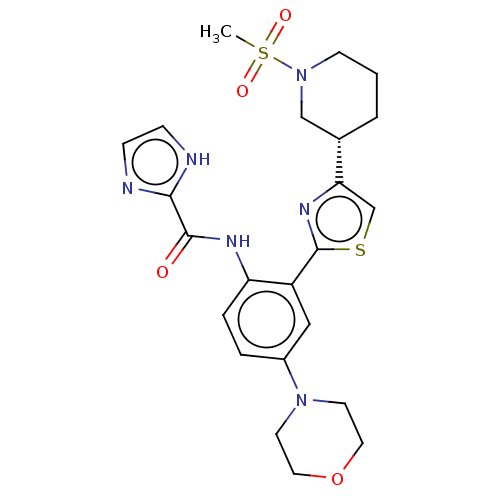
Affinity DataIC50: 1.80nMAssay Description:Inhibition of Axl (unknown origin)More data for this Ligand-Target Pair
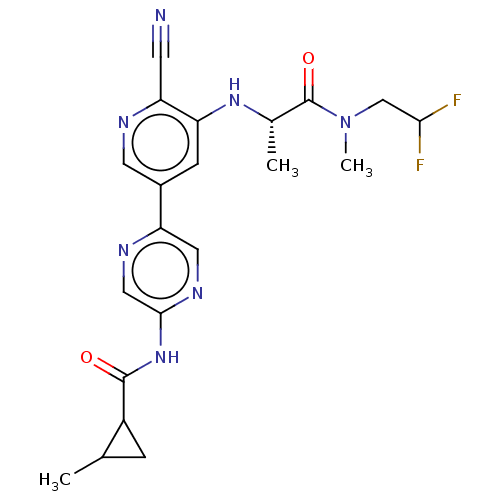
Affinity DataIC50: 2.20nMAssay Description:Inhibition of Axl (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 2.51nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
TargetAldehyde dehydrogenase, mitochondrial(Homo sapiens (Human))
University Of Oxford
Curated by ChEMBL
University Of Oxford
Curated by ChEMBL
Affinity DataIC50: 3nMAssay Description:Inhibition of Hamster Liver mitochondrial ALDH-2More data for this Ligand-Target Pair
Affinity DataIC50: 3.16nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.40nMAssay Description:Inhibition of Axl (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 3.98nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
TargetAldehyde dehydrogenase, mitochondrial(Homo sapiens (Human))
University Of Oxford
Curated by ChEMBL
University Of Oxford
Curated by ChEMBL
Affinity DataIC50: 4nMAssay Description:Inhibition of Hamster Liver mitochondrial ALDH-2More data for this Ligand-Target Pair
Affinity DataIC50: 5.01nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
Affinity DataIC50: 6.31nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
Affinity DataIC50: 6.31nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
Affinity DataIC50: 6.31nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
Affinity DataIC50: 7.94nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
Affinity DataIC50: 7.94nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
Affinity DataIC50: 7.94nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
Affinity DataIC50: 7.94nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
Affinity DataIC50: 7.94nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
TargetAldehyde dehydrogenase, mitochondrial(Homo sapiens (Human))
University Of Oxford
Curated by ChEMBL
University Of Oxford
Curated by ChEMBL
Affinity DataIC50: 9nMAssay Description:Inhibition of Hamster Liver mitochondrial ALDH-2More data for this Ligand-Target Pair
TargetAldehyde dehydrogenase, mitochondrial(Homo sapiens (Human))
University Of Oxford
Curated by ChEMBL
University Of Oxford
Curated by ChEMBL
Affinity DataIC50: 9nMAssay Description:Inhibition of Hamster Liver mitochondrial ALDH-2More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
Affinity DataIC50: 12.6nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
Affinity DataIC50: 12.6nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
Affinity DataIC50: 12.6nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
Affinity DataIC50: 15.8nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
Affinity DataIC50: 15.8nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
Affinity DataIC50: 15.8nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
Affinity DataIC50: 15.8nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
Affinity DataIC50: 15.8nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
Affinity DataIC50: 15.8nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
Affinity DataIC50: 15.8nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
Affinity DataIC50: 15.8nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
Affinity DataIC50: 25nMAssay Description:Inhibition of Axl (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 25.1nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair
Affinity DataIC50: 25.1nMAssay Description:TBK1 inhibition is determined using a 384 well μlate format in buffer containing 20 mM Hepes, pH 7.4, 10 mM MgCl2, 1 mM EDTA, 0.01% Brij-L23, 1 ...More data for this Ligand-Target Pair