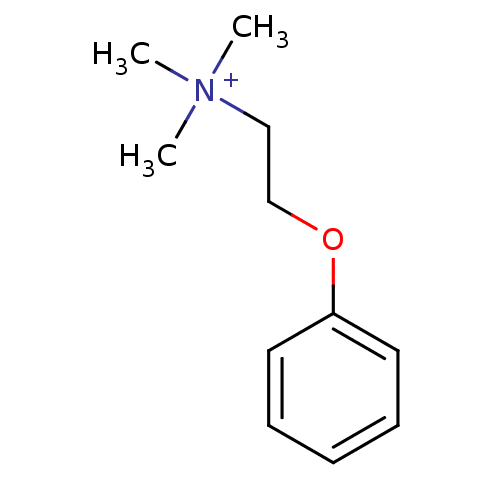
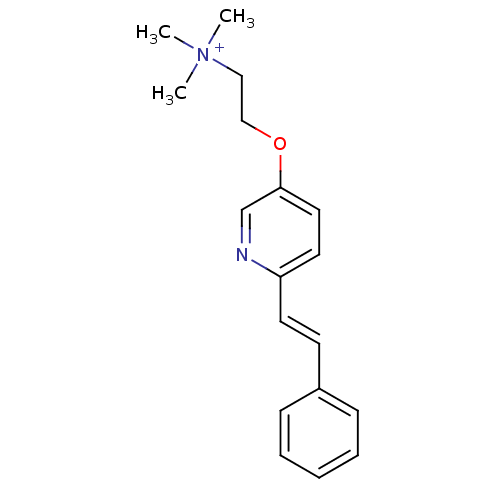
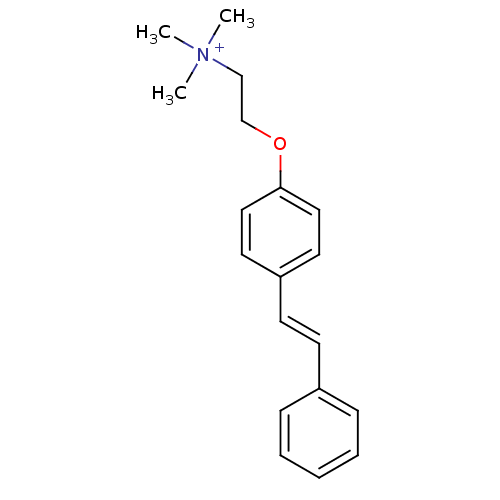
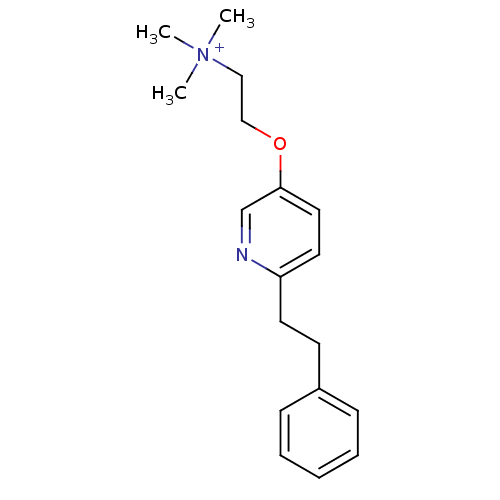
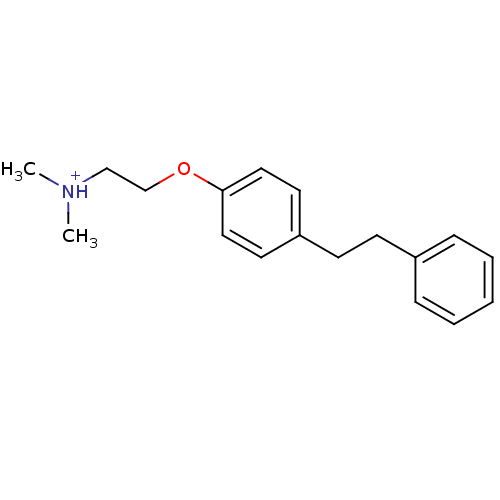
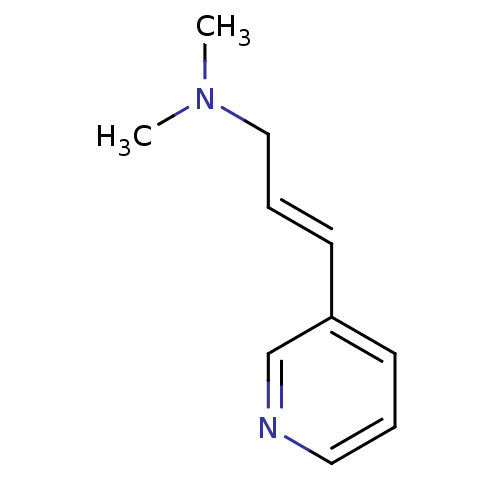
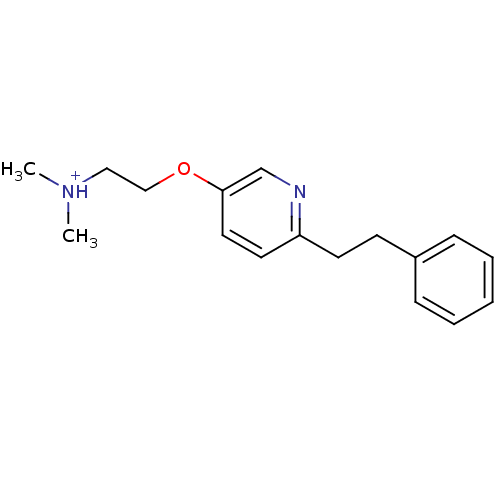
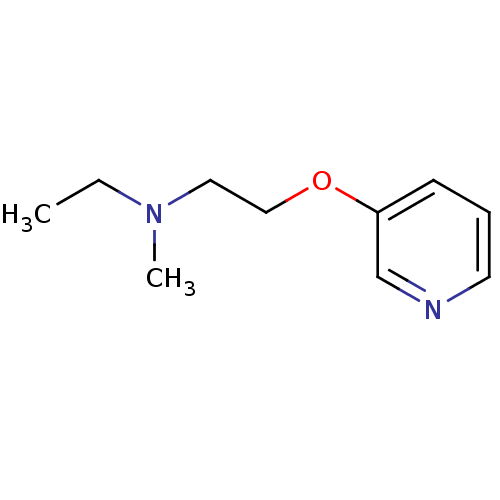
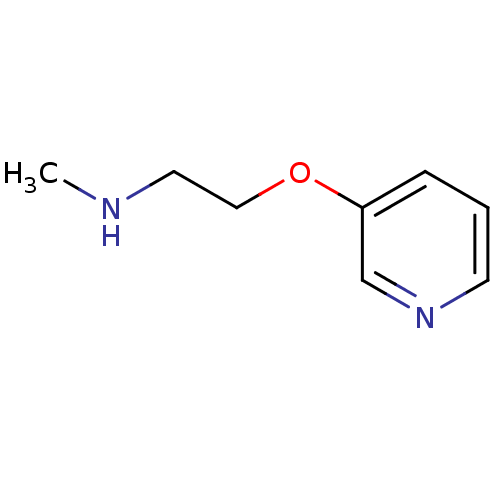
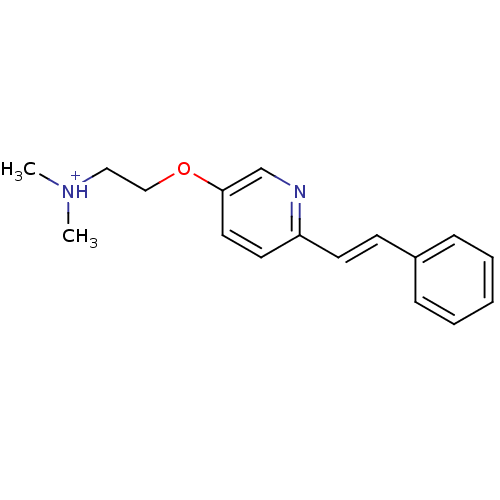
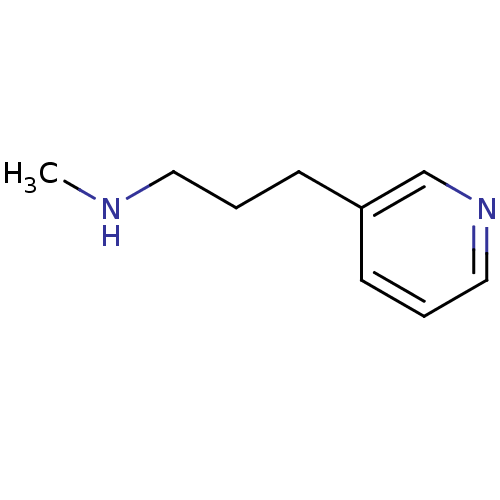
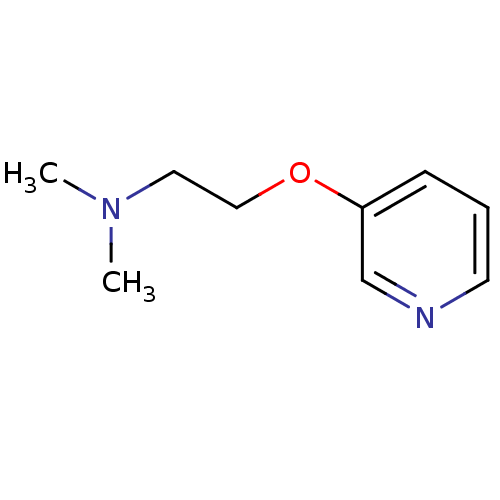
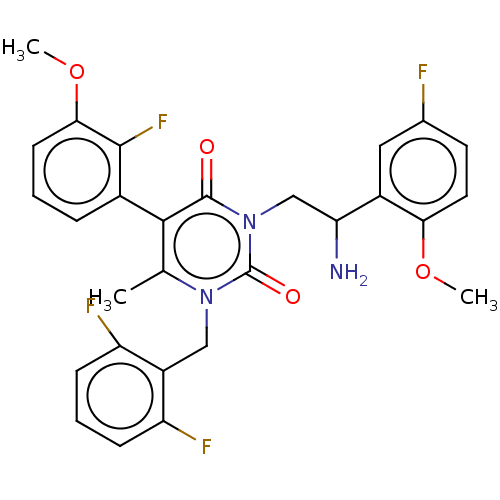
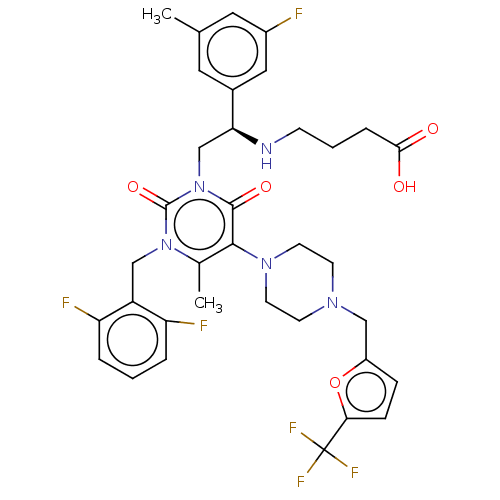
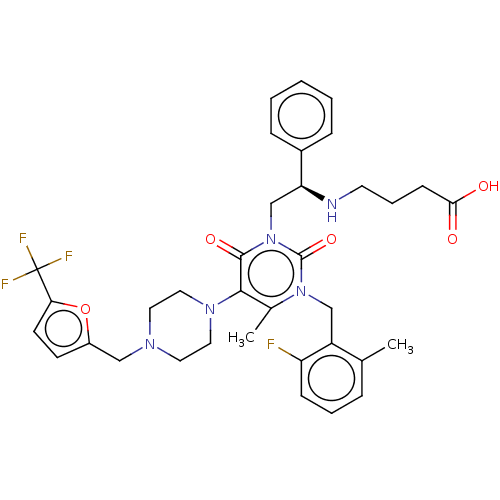
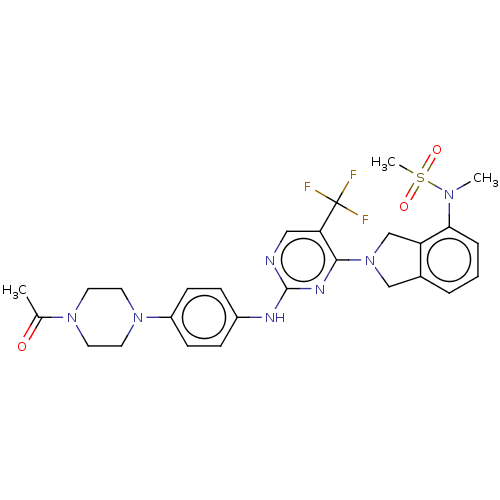
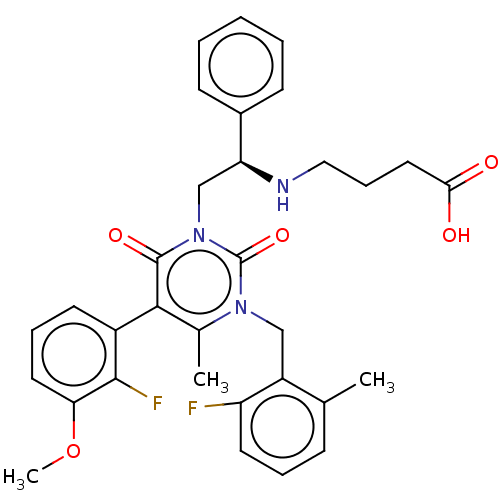
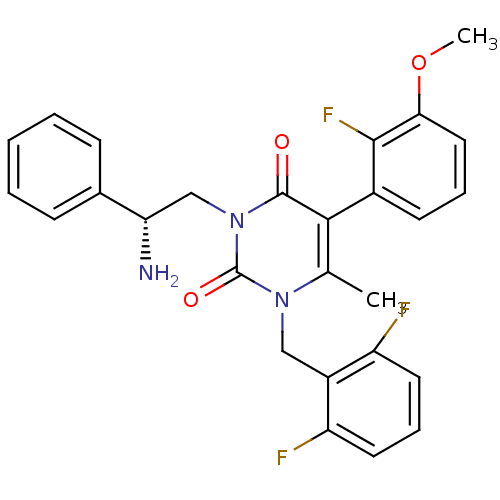
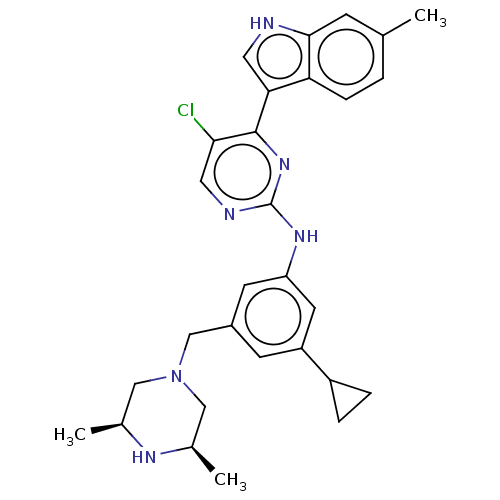
TargetAmyloid-beta precursor protein(Homo sapiens (Human))
Seoul National University Bundang Hospital
Curated by ChEMBL
Seoul National University Bundang Hospital
Curated by ChEMBL
Affinity DataKi: 0.200nMAssay Description:Displacement of [3H]PIB from amyloid beta (1 to 42) aggregates in Alzheimer's disease-human brain homogenate after 3 hrs by gamma countingMore data for this Ligand-Target Pair
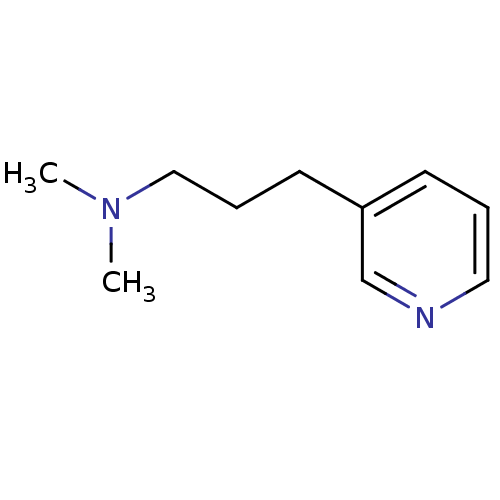
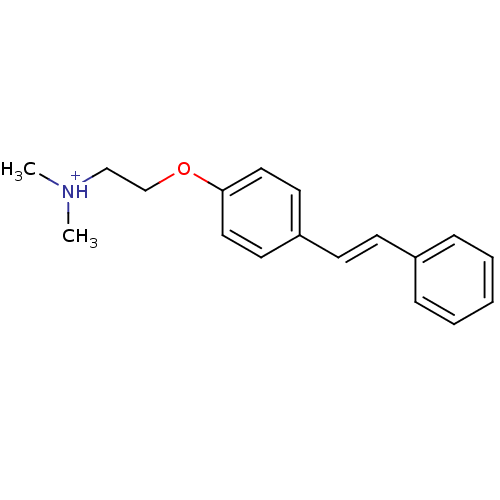
TargetAmyloid-beta precursor protein(Homo sapiens (Human))
Seoul National University Bundang Hospital
Curated by ChEMBL
Seoul National University Bundang Hospital
Curated by ChEMBL
Affinity DataKi: 0.300nMAssay Description:Displacement of [3H]PIB from amyloid beta (1 to 42) aggregates in Alzheimer's disease-human brain homogenate after 3 hrs by gamma countingMore data for this Ligand-Target Pair
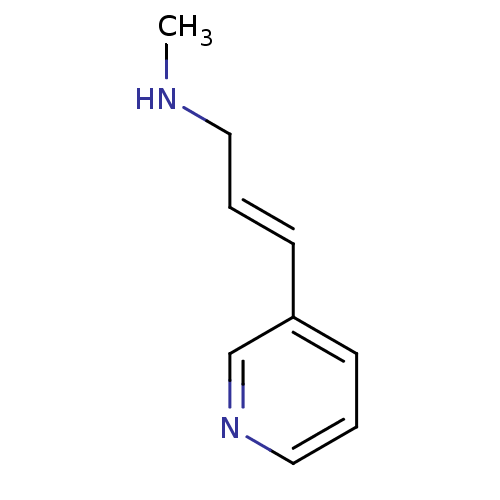
TargetNeuronal acetylcholine receptor subunit alpha-4/beta-2(Homo sapiens (Human))
Virginia Commonwealth University
Curated by ChEMBL
Virginia Commonwealth University
Curated by ChEMBL
Affinity DataKi: 0.5nMAssay Description:Binding affinity at alpha4beta2More data for this Ligand-Target Pair
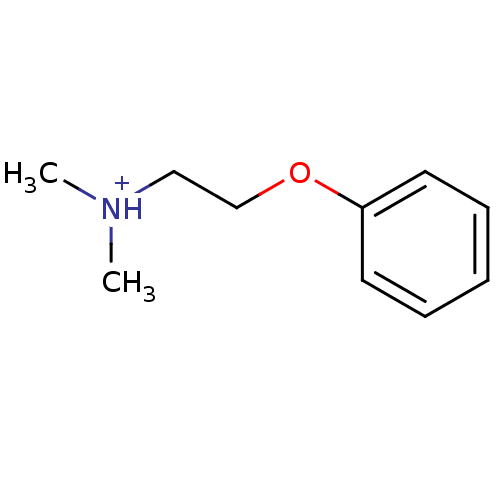
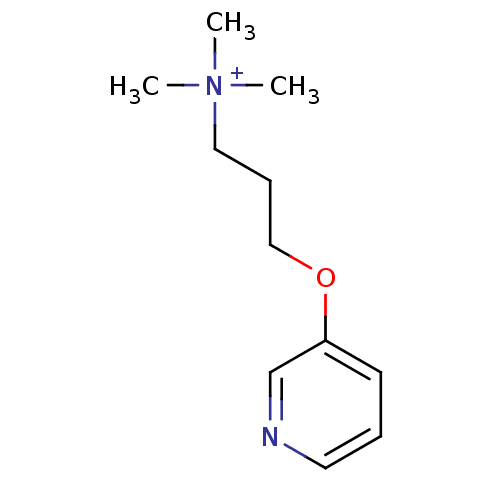
TargetAmyloid-beta precursor protein(Homo sapiens (Human))
Seoul National University Bundang Hospital
Curated by ChEMBL
Seoul National University Bundang Hospital
Curated by ChEMBL
Affinity DataKi: 1.60nMAssay Description:Displacement of [3H]PIB from amyloid beta (1 to 42) aggregates in Alzheimer's disease-human brain homogenate after 3 hrs by gamma countingMore data for this Ligand-Target Pair
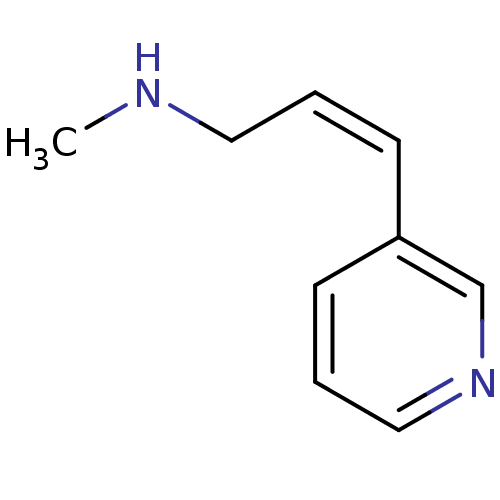
TargetAmyloid-beta precursor protein(Homo sapiens (Human))
Seoul National University Bundang Hospital
Curated by ChEMBL
Seoul National University Bundang Hospital
Curated by ChEMBL
Affinity DataKi: 5.5nMAssay Description:Displacement of [125I]TZDM from amyloid beta (1 to 42) aggregates in Alzheimer's disease-human brain homogenate after 3 hrs by gamma countingMore data for this Ligand-Target Pair
TargetAmyloid-beta precursor protein(Homo sapiens (Human))
Seoul National University Bundang Hospital
Curated by ChEMBL
Seoul National University Bundang Hospital
Curated by ChEMBL
Affinity DataKi: 5.80nMAssay Description:Displacement of [125I]TZDM from amyloid beta (1 to 42) aggregates in Alzheimer's disease-human brain homogenate after 3 hrs by gamma countingMore data for this Ligand-Target Pair
TargetAmyloid-beta precursor protein(Homo sapiens (Human))
Seoul National University Bundang Hospital
Curated by ChEMBL
Seoul National University Bundang Hospital
Curated by ChEMBL
Affinity DataKi: 5.90nMAssay Description:Displacement of [125I]TZDM from amyloid beta (1 to 42) aggregates in Alzheimer's disease-human brain homogenate after 3 hrs by gamma countingMore data for this Ligand-Target Pair
TargetAmyloid-beta precursor protein(Homo sapiens (Human))
Seoul National University Bundang Hospital
Curated by ChEMBL
Seoul National University Bundang Hospital
Curated by ChEMBL
Affinity DataKi: 26nMAssay Description:Displacement of [125I]TZDM from amyloid beta (1 to 42) aggregates in Alzheimer's disease-human brain homogenate after 3 hrs by gamma countingMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-7(Rattus norvegicus (Rat))
Virginia Commonwealth University
Curated by ChEMBL
Virginia Commonwealth University
Curated by ChEMBL
Affinity DataKi: 280nMAssay Description:Displacement of [125I]iodo-MLA from alpha-7 nAChR in rat cerebral cortexMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-7(Rattus norvegicus (Rat))
Virginia Commonwealth University
Curated by ChEMBL
Virginia Commonwealth University
Curated by ChEMBL
Affinity DataKi: 750nMAssay Description:Displacement of [125I]iodo-MLA from alpha-7 nAChR in rat cerebral cortexMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-7(Rattus norvegicus (Rat))
Virginia Commonwealth University
Curated by ChEMBL
Virginia Commonwealth University
Curated by ChEMBL
Affinity DataKi: 1.10E+3nMAssay Description:Displacement of [125I]iodo-MLA from alpha-7 nAChR in rat cerebral cortexMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-7(Rattus norvegicus (Rat))
Virginia Commonwealth University
Curated by ChEMBL
Virginia Commonwealth University
Curated by ChEMBL
Affinity DataKi: 1.25E+3nMAssay Description:Displacement of [125I]iodo-MLA from alpha-7 nAChR in rat cerebral cortexMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-7(Rattus norvegicus (Rat))
Virginia Commonwealth University
Curated by ChEMBL
Virginia Commonwealth University
Curated by ChEMBL
Affinity DataKi: 2.17E+3nMAssay Description:Displacement of [125I]iodo-MLA from alpha-7 nAChR in rat cerebral cortexMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-7(Rattus norvegicus (Rat))
Virginia Commonwealth University
Curated by ChEMBL
Virginia Commonwealth University
Curated by ChEMBL
Affinity DataKi: 4.32E+3nMAssay Description:Displacement of [125I]iodo-MLA from alpha-7 nAChR in rat cerebral cortexMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-7(Rattus norvegicus (Rat))
Virginia Commonwealth University
Curated by ChEMBL
Virginia Commonwealth University
Curated by ChEMBL
Affinity DataKi: 7.50E+3nMAssay Description:Displacement of [125I]iodo-MLA from alpha-7 nAChR in rat cerebral cortexMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-7(Rattus norvegicus (Rat))
Virginia Commonwealth University
Curated by ChEMBL
Virginia Commonwealth University
Curated by ChEMBL
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [125I]iodo-MLA from alpha-7 nAChR in rat cerebral cortexMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-7(Rattus norvegicus (Rat))
Virginia Commonwealth University
Curated by ChEMBL
Virginia Commonwealth University
Curated by ChEMBL
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [125I]iodo-MLA from alpha-7 nAChR in rat cerebral cortexMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-7(Rattus norvegicus (Rat))
Virginia Commonwealth University
Curated by ChEMBL
Virginia Commonwealth University
Curated by ChEMBL
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [125I]iodo-MLA from alpha-7 nAChR in rat cerebral cortexMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-7(Rattus norvegicus (Rat))
Virginia Commonwealth University
Curated by ChEMBL
Virginia Commonwealth University
Curated by ChEMBL
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [125I]iodo-MLA from alpha-7 nAChR in rat cerebral cortexMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-7(Rattus norvegicus (Rat))
Virginia Commonwealth University
Curated by ChEMBL
Virginia Commonwealth University
Curated by ChEMBL
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [125I]iodo-MLA from alpha-7 nAChR in rat cerebral cortexMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-7(Rattus norvegicus (Rat))
Virginia Commonwealth University
Curated by ChEMBL
Virginia Commonwealth University
Curated by ChEMBL
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [125I]iodo-MLA from alpha-7 nAChR in rat cerebral cortexMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-7(Rattus norvegicus (Rat))
Virginia Commonwealth University
Curated by ChEMBL
Virginia Commonwealth University
Curated by ChEMBL
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [125I]iodo-MLA from alpha-7 nAChR in rat cerebral cortexMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-7(Rattus norvegicus (Rat))
Virginia Commonwealth University
Curated by ChEMBL
Virginia Commonwealth University
Curated by ChEMBL
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [125I]iodo-MLA from alpha-7 nAChR in rat cerebral cortexMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-7(Rattus norvegicus (Rat))
Virginia Commonwealth University
Curated by ChEMBL
Virginia Commonwealth University
Curated by ChEMBL
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [125I]iodo-MLA from alpha-7 nAChR in rat cerebral cortexMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-7(Rattus norvegicus (Rat))
Virginia Commonwealth University
Curated by ChEMBL
Virginia Commonwealth University
Curated by ChEMBL
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [125I]iodo-MLA from alpha-7 nAChR in rat cerebral cortexMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-7(Rattus norvegicus (Rat))
Virginia Commonwealth University
Curated by ChEMBL
Virginia Commonwealth University
Curated by ChEMBL
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [125I]iodo-MLA from alpha-7 nAChR in rat cerebral cortexMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-7(Rattus norvegicus (Rat))
Virginia Commonwealth University
Curated by ChEMBL
Virginia Commonwealth University
Curated by ChEMBL
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [125I]iodo-MLA from alpha-7 nAChR in rat cerebral cortexMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-7(Rattus norvegicus (Rat))
Virginia Commonwealth University
Curated by ChEMBL
Virginia Commonwealth University
Curated by ChEMBL
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [125I]iodo-MLA from alpha-7 nAChR in rat cerebral cortexMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-7(Rattus norvegicus (Rat))
Virginia Commonwealth University
Curated by ChEMBL
Virginia Commonwealth University
Curated by ChEMBL
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [125I]iodo-MLA from alpha-7 nAChR in rat cerebral cortexMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-7(Rattus norvegicus (Rat))
Virginia Commonwealth University
Curated by ChEMBL
Virginia Commonwealth University
Curated by ChEMBL
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [125I]iodo-MLA from alpha-7 nAChR in rat cerebral cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 0.290nMAssay Description:Displacement of [125I]D-Trp6-LHRH from human GnRH receptor expressed in CHO-K1 cells after 1 hr by microbeta scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataIC50: 0.330nMAssay Description:Binding affinity of compound to human recombinant Ribonuclease L was evaluatedMore data for this Ligand-Target Pair
Affinity DataIC50: 0.380nMAssay Description:Displacement of [125I]D-Trp6-LHRH from human GnRH receptor expressed in CHO-K1 cells after 1 hr by microbeta scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataIC50: 0.400nMAssay Description:The inhibitory activity of some of the compounds described above against FLT3-ITD, FLT3 wild type (WT), VEGFR2 (KDR), and SYK kinase was measured. Th...More data for this Ligand-Target Pair
Affinity DataIC50: 0.400nMAssay Description:The inhibitory activity of some of the compounds described above against FLT3-ITD, FLT3 wild type (WT), VEGFR2 (KDR), and SYK kinase was measured. Th...More data for this Ligand-Target Pair
Affinity DataIC50: 0.400nMAssay Description:The inhibitory activity of some of the compounds described above against FLT3-ITD, FLT3 wild type (WT), VEGFR2 (KDR), and SYK kinase was measured.The...More data for this Ligand-Target Pair
Affinity DataIC50: 0.400nMAssay Description:The inhibitory activity of some of the compounds described above against FLT3-ITD, FLT3 wild type (WT), VEGFR2 (KDR), and SYK kinase was measured.The...More data for this Ligand-Target Pair
Affinity DataIC50: 0.460nMAssay Description:Displacement of [125I]D-Trp6-LHRH from human GnRH receptor expressed in CHO-K1 cells after 1 hr by microbeta scintillation counting methodMore data for this Ligand-Target Pair
TargetDimer of Receptor-type tyrosine-protein kinase FLT3(Homo sapiens (Human))
Hanmi Pharm.
US Patent
Hanmi Pharm.
US Patent
Affinity DataIC50: 0.5nMAssay Description:The inhibitory activity of some of the compounds described above against FLT3-ITD, FLT3 wild type (WT), VEGFR2 (KDR), and SYK kinase was measured.The...More data for this Ligand-Target Pair
Affinity DataIC50: <0.5nMAssay Description:Inhibition of human AXL using EAIYAAPFAKKK as substrate by [gamma-33P]-ATP assayMore data for this Ligand-Target Pair
TargetDimer of Receptor-type tyrosine-protein kinase FLT3(Homo sapiens (Human))
Hanmi Pharm.
US Patent
Hanmi Pharm.
US Patent
Affinity DataIC50: 0.5nMAssay Description:The inhibitory activity of some of the compounds described above against FLT3-ITD, FLT3 wild type (WT), VEGFR2 (KDR), and SYK kinase was measured. Th...More data for this Ligand-Target Pair
Affinity DataIC50: <0.5nMAssay Description:Inhibition of N-terminal His-tagged recombinant human AXL (473 to end amino acids) expressed by baculovirus in Sf9 cells using axltide substrate and ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.580nMAssay Description:Displacement of [125I]D-Trp6-LHRH from human GnRH receptor expressed in CHO-K1 cells after 1 hr by microbeta scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataIC50: 0.590nMAssay Description:Displacement of [125I]D-Trp6-LHRH from human GnRH receptor expressed in CHO-K1 cells after 1 hr by microbeta scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataIC50: 0.740nMAssay Description:Displacement of [125I]D-Trp6-LHRH from human GnRH receptor expressed in CHO-K1 cells after 1 hr by microbeta scintillation counting methodMore data for this Ligand-Target Pair
Target11-beta-hydroxysteroid dehydrogenase 1(Homo sapiens (Human))
Seoul National University
Curated by ChEMBL
Seoul National University
Curated by ChEMBL
Affinity DataIC50: 0.900nMAssay Description:Inhibition of human 11beta-HSD1 expressed in HEK293 cells using NADPH assessed as conversion of cortisone to cortisol by cell-based assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.940nMAssay Description:Inhibition of N-terminal His-tagged recombinant human AXL (473 to end amino acids) expressed by baculovirus in Sf9 cells using axltide substrate and ...More data for this Ligand-Target Pair
TargetProteinase-activated receptor 1(Homo sapiens (Human))
Korea Research Institute Of Technology
Curated by ChEMBL
Korea Research Institute Of Technology
Curated by ChEMBL
Affinity DataIC50: 1.10nMAssay Description:Displacement of [3H]haTRAP from PAR1 in human platelet membranes after 60 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.10nMAssay Description:The inhibitory activity of some of the compounds described above against FLT3-ITD, FLT3 wild type (WT), VEGFR2 (KDR), and SYK kinase was measured.The...More data for this Ligand-Target Pair
Affinity DataIC50: 1.10nMAssay Description:The inhibitory activity of some of the compounds described above against FLT3-ITD, FLT3 wild type (WT), VEGFR2 (KDR), and SYK kinase was measured. Th...More data for this Ligand-Target Pair

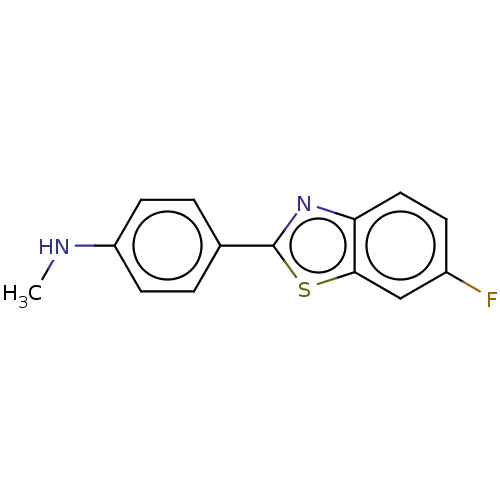
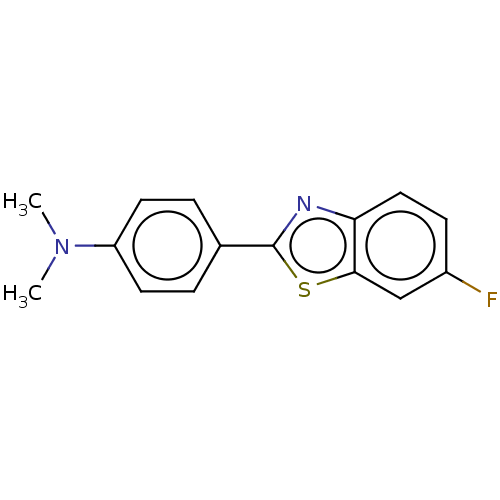
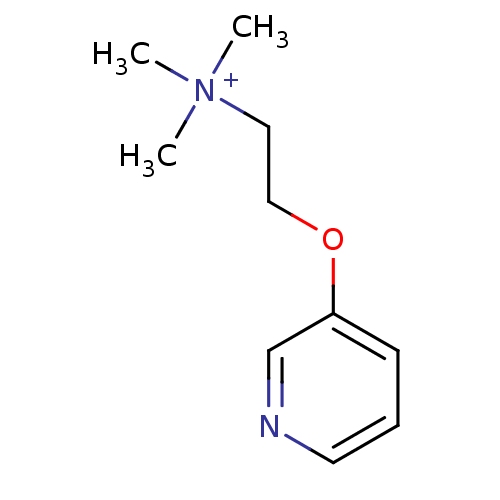
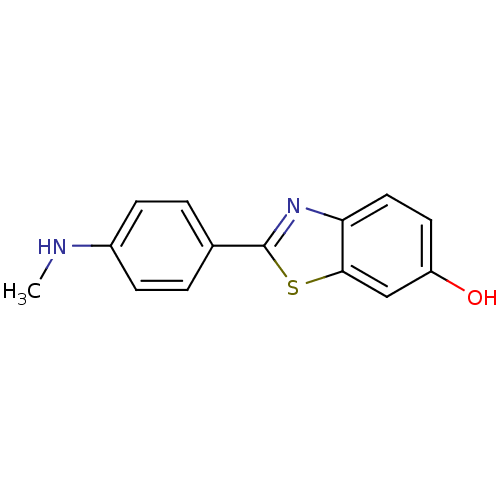
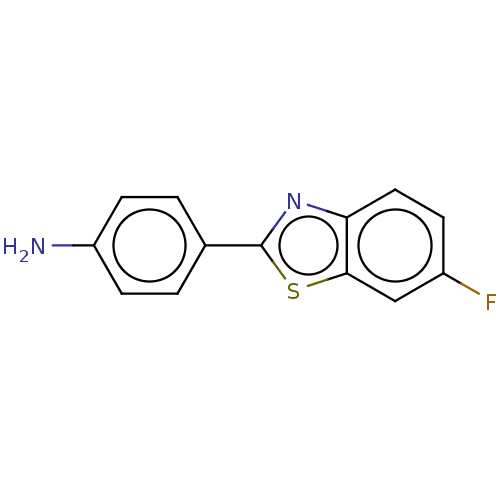
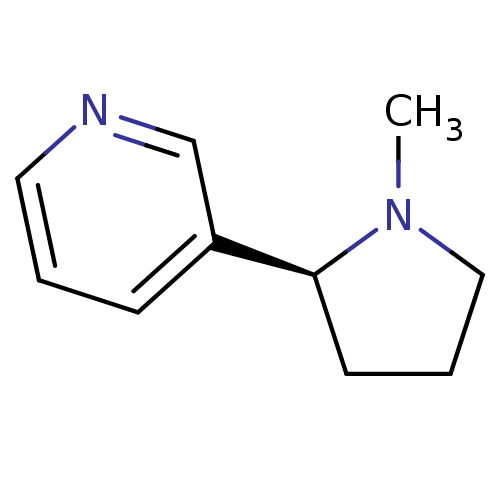
 3D Structure (crystal)
3D Structure (crystal)