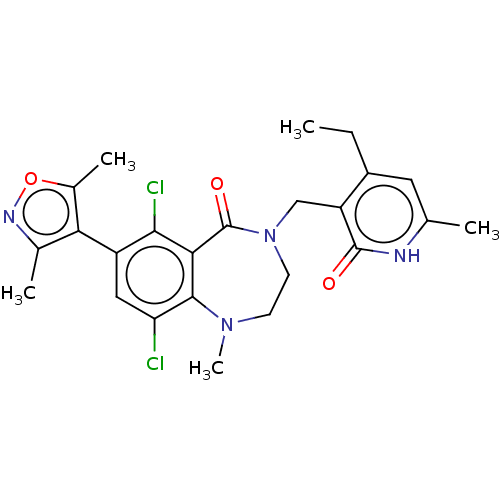
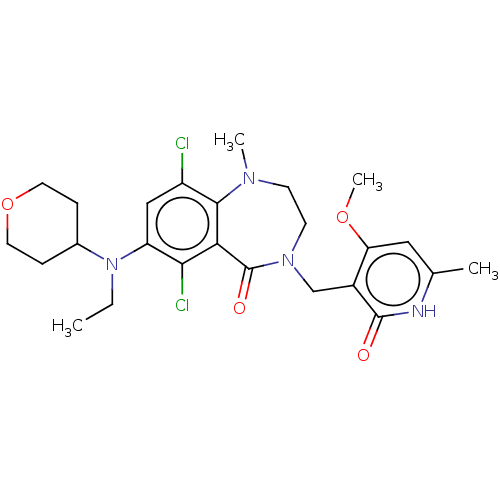
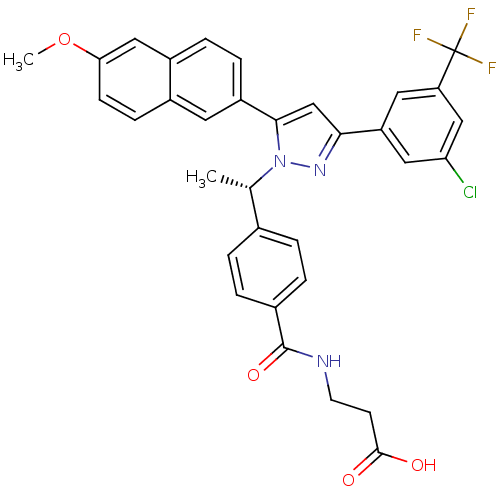
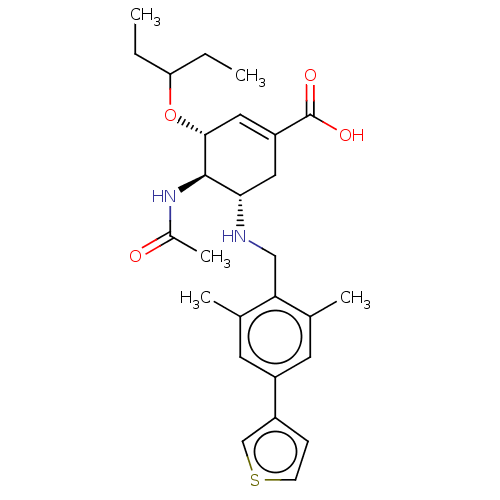
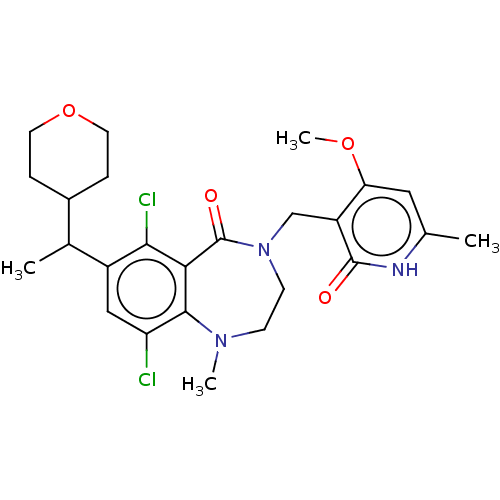
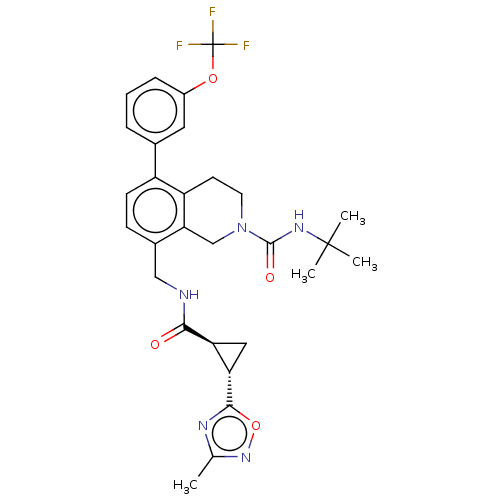
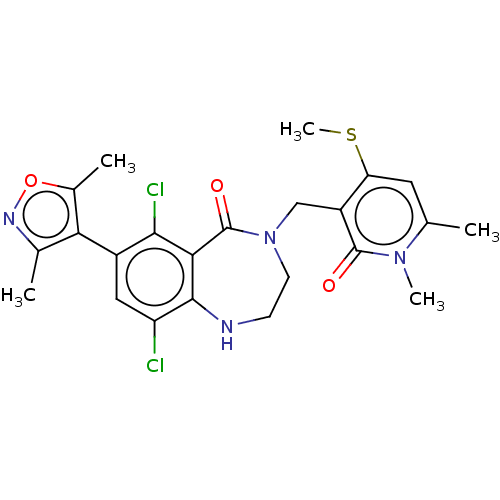
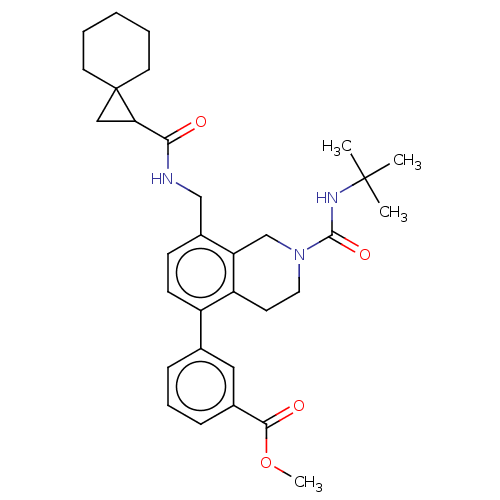
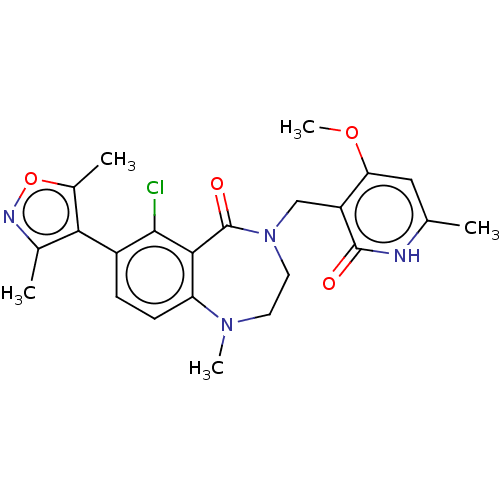
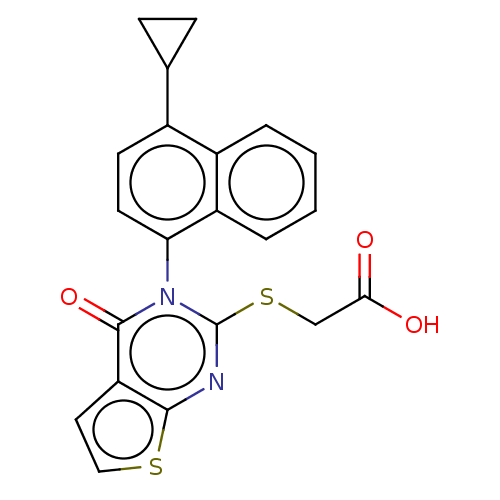
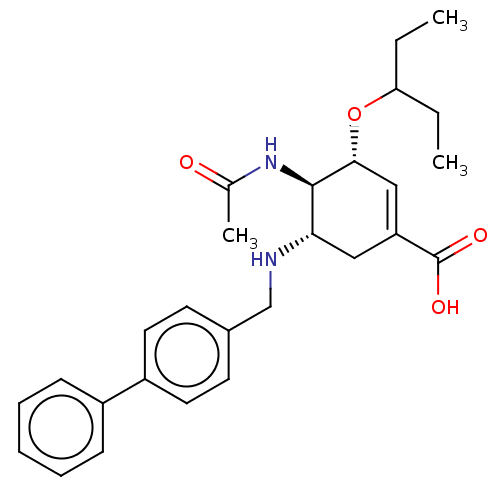
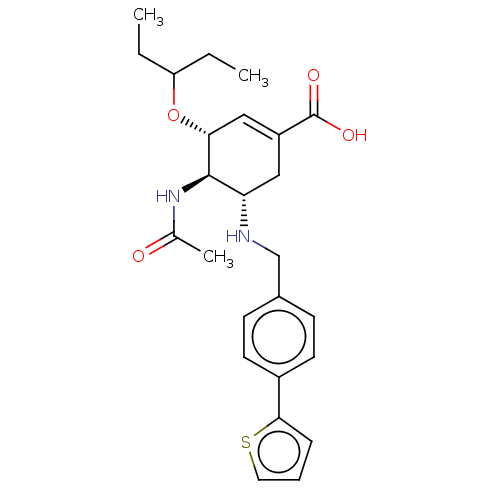
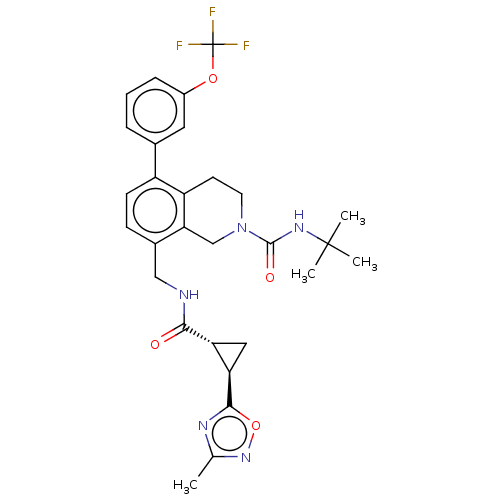
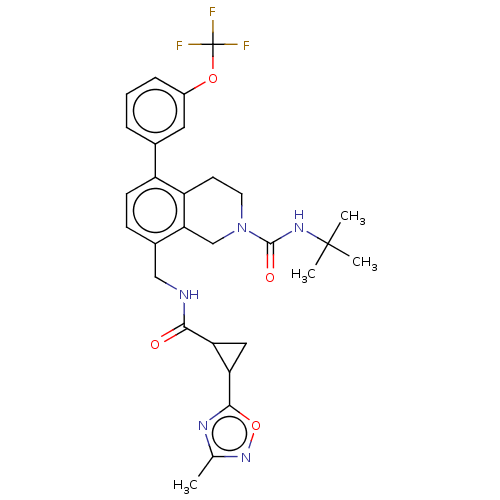
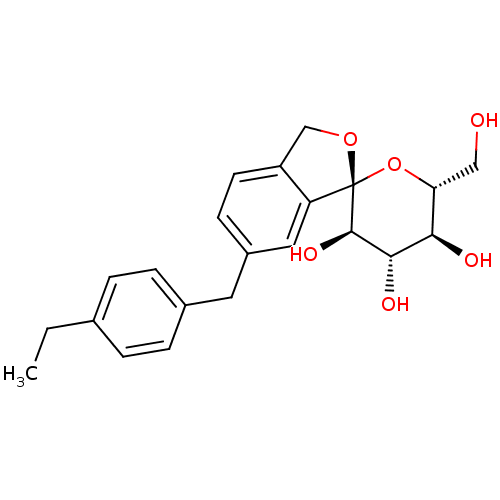
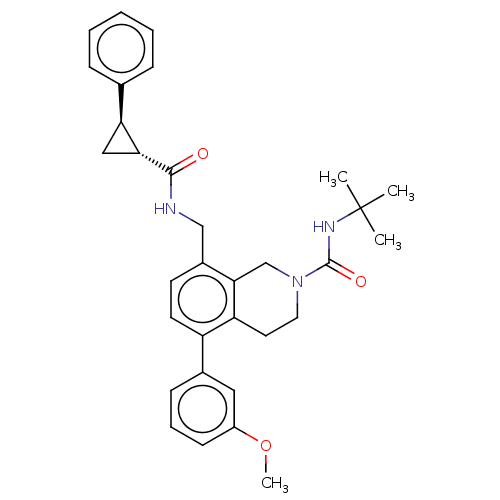
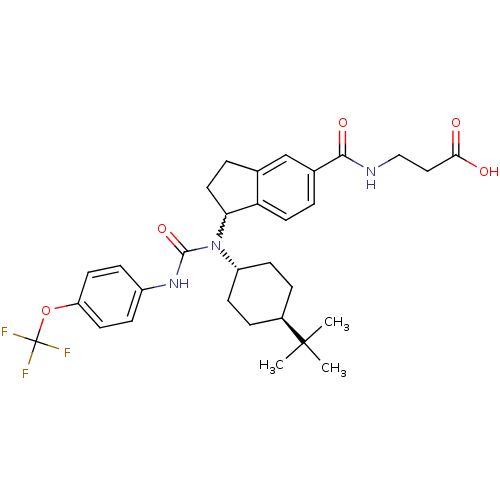
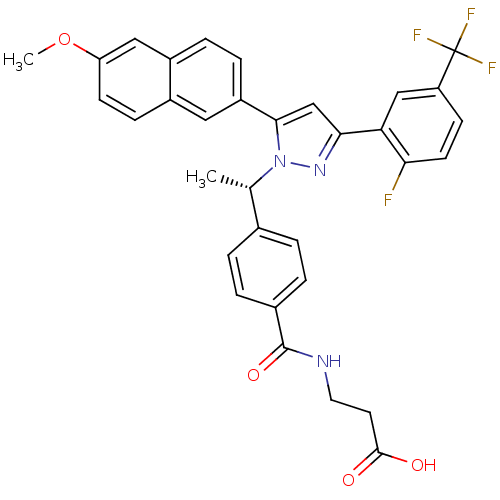
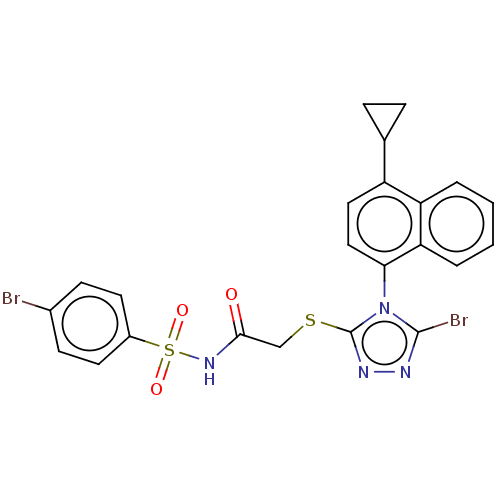
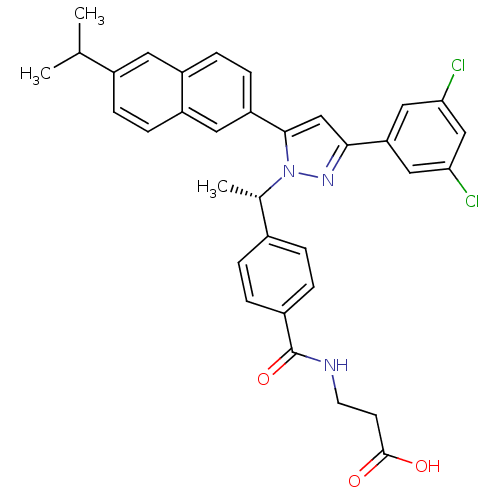
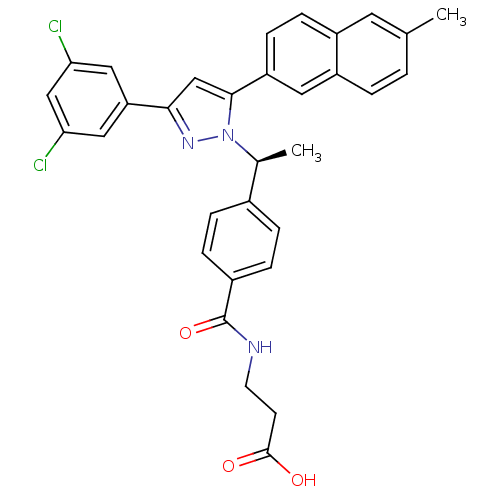
TargetHistone-lysine N-methyltransferase EZH2(Homo sapiens (Human))
Tri-Institutional Therapeutics Discovery Institute
Curated by ChEMBL
Tri-Institutional Therapeutics Discovery Institute
Curated by ChEMBL
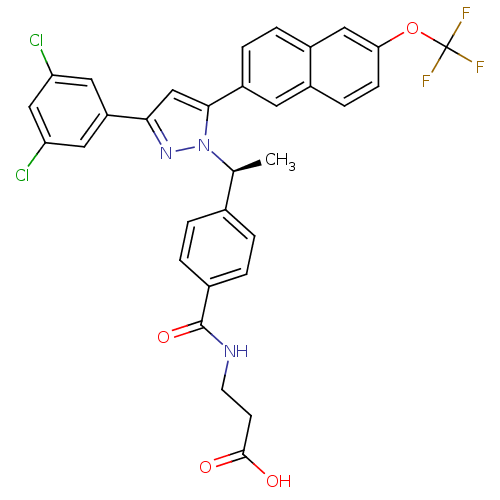
Affinity DataIC50: 0.350nMAssay Description:Inhibition of EZH2 (unknown origin) using 3H-SAM as substrate incubated for 1 hour by filter binding methodMore data for this Ligand-Target Pair
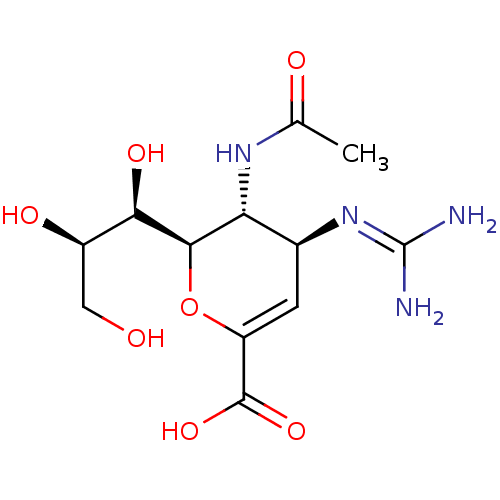
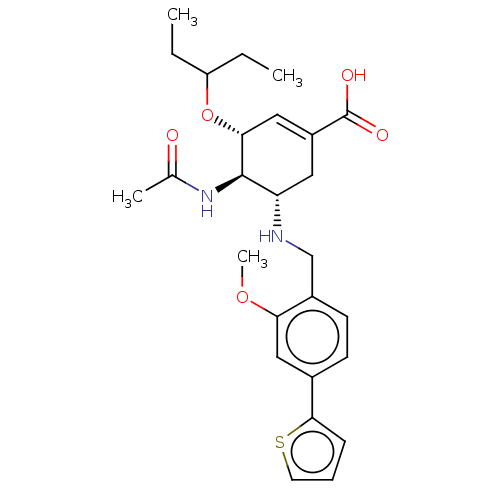
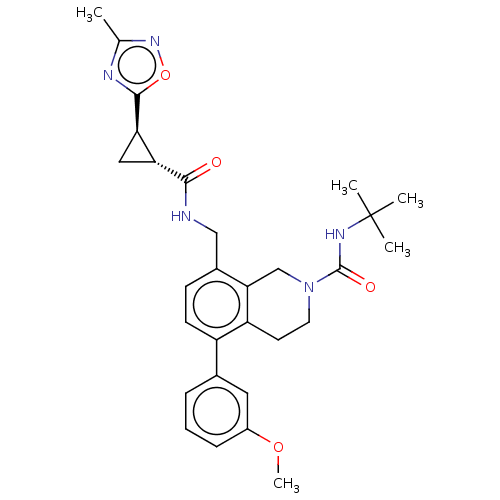
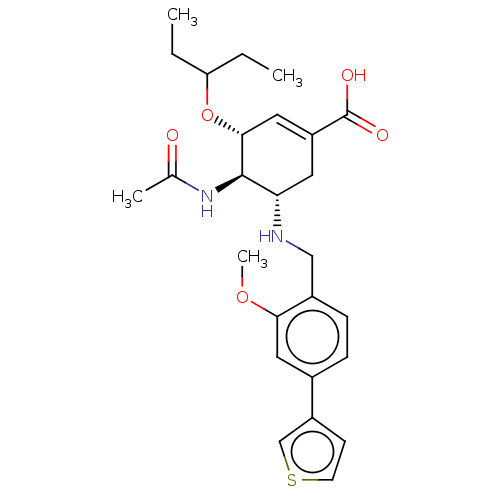
TargetNeuraminidase(Influenza A virus (A/Puerto Rico/8/34/Mount Sinai(...)
Shandong University
Curated by ChEMBL
Shandong University
Curated by ChEMBL
Affinity DataIC50: 0.5nMAssay Description:Inhibition of Influenza A virus A/PR/8/1934 (H1N1) neuraminidase using MUNANA as substrate preincubated for 10 mins followed by substrate addition an...More data for this Ligand-Target Pair
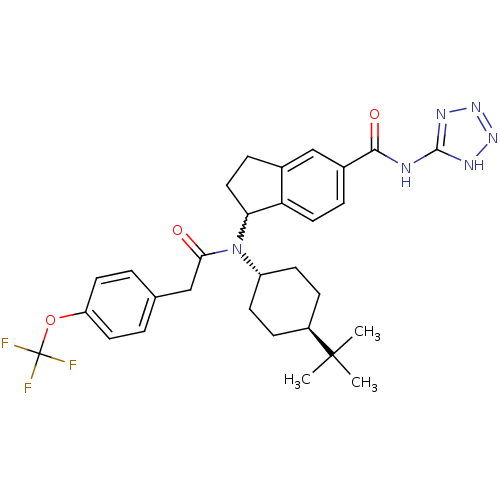
Affinity DataIC50: 0.660nMAssay Description:DGAT2 activity was determined by measuring the amount of enzymatic product triolein (1,2,3-Tri(cis-9-octadecenoyl)glycerol) using the membrane prep m...More data for this Ligand-Target Pair
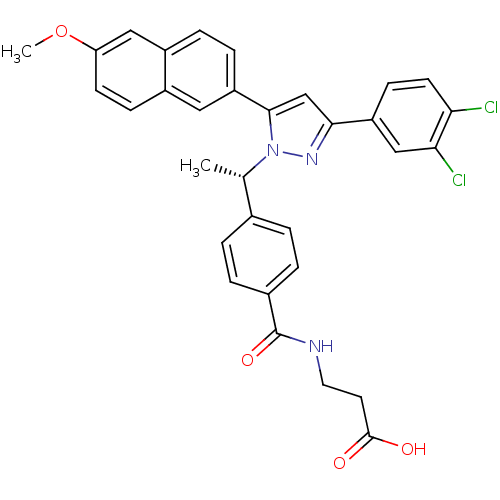
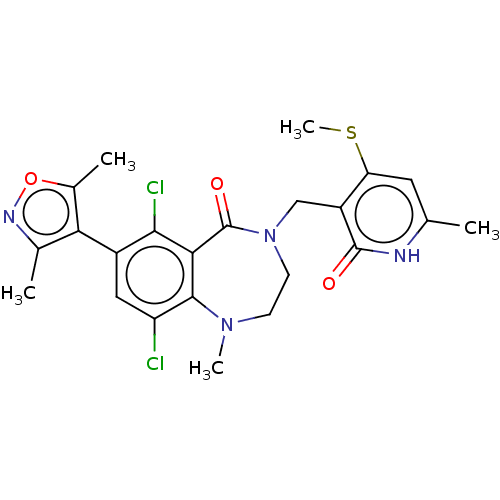
TargetHistone-lysine N-methyltransferase EZH2(Homo sapiens (Human))
Tri-Institutional Therapeutics Discovery Institute
Curated by ChEMBL
Tri-Institutional Therapeutics Discovery Institute
Curated by ChEMBL
Affinity DataIC50: 0.870nMAssay Description:Inhibition of EZH2 (unknown origin) using 3H-SAM as substrate incubated for 1 hour by filter binding methodMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Homo sapiens (Human))
Tri-Institutional Therapeutics Discovery Institute
Curated by ChEMBL
Tri-Institutional Therapeutics Discovery Institute
Curated by ChEMBL
Affinity DataIC50: 0.900nMAssay Description:Inhibition of EZH2 (unknown origin) using 3H-SAM as substrate incubated for 1 hour by filter binding methodMore data for this Ligand-Target Pair
TargetNeuraminidase(Influenza A virus (A/Puerto Rico/8/34/Mount Sinai(...)
Shandong University
Curated by ChEMBL
Shandong University
Curated by ChEMBL
Affinity DataIC50: 0.950nMAssay Description:Inhibition of Influenza A virus A/PR/8/1934 (H1N1) neuraminidase using MUNANA as substrate preincubated for 10 mins followed by substrate addition an...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Homo sapiens (Human))
Tri-Institutional Therapeutics Discovery Institute
Curated by ChEMBL
Tri-Institutional Therapeutics Discovery Institute
Curated by ChEMBL
Affinity DataIC50: 1nMAssay Description:Inhibition of EZH2 (unknown origin) using 3H-SAM as substrate incubated for 1 hour by filter binding methodMore data for this Ligand-Target Pair
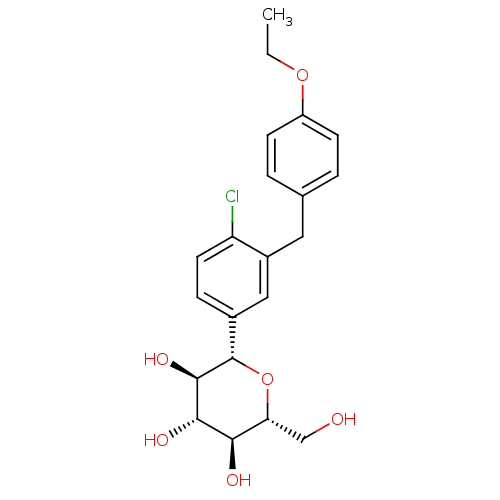
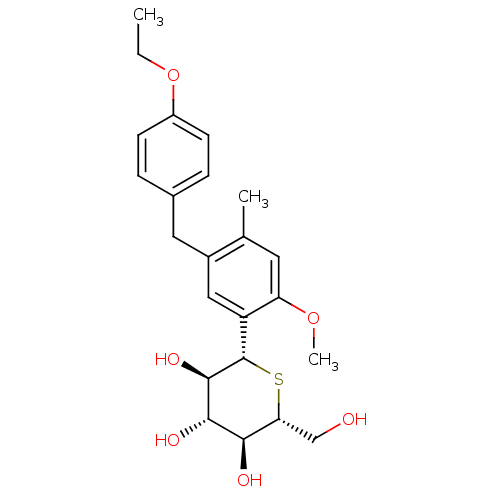
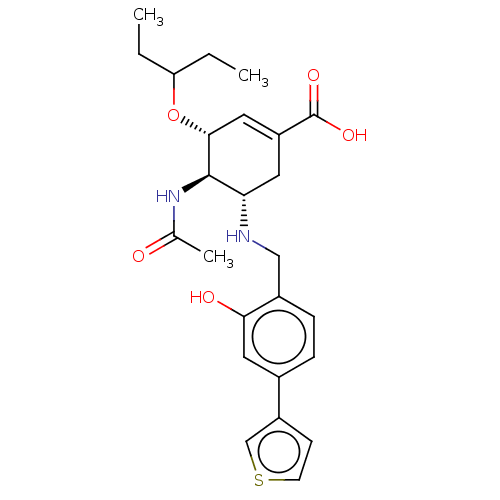
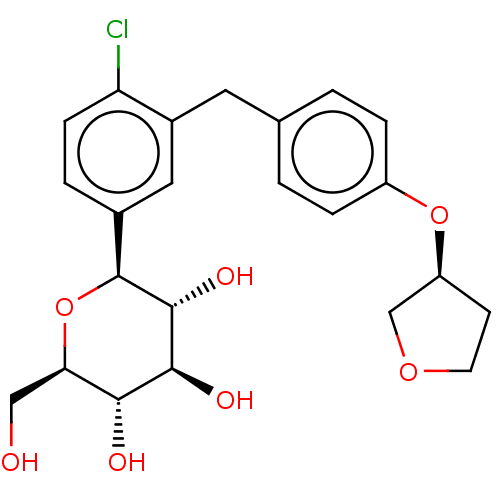
TargetSodium/glucose cotransporter 2(Homo sapiens (Human))
Guangdong Pharmaceutical University
Curated by ChEMBL
Guangdong Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 1.10nMAssay Description:Inhibition of SGLT2 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 1.13nMAssay Description:DGAT2 activity was determined by measuring the amount of enzymatic product triolein (1,2,3-Tri(cis-9-octadecenoyl)glycerol) using the membrane prep m...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Homo sapiens (Human))
Tri-Institutional Therapeutics Discovery Institute
Curated by ChEMBL
Tri-Institutional Therapeutics Discovery Institute
Curated by ChEMBL
Affinity DataIC50: 1.30nMAssay Description:Inhibition of EZH2 (unknown origin) using 3H-SAM as substrate incubated for 1 hour by filter binding methodMore data for this Ligand-Target Pair
TargetSodium/glucose cotransporter 2(Homo sapiens (Human))
Guangdong Pharmaceutical University
Curated by ChEMBL
Guangdong Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 1.40nMAssay Description:Inhibition of human SGLT2 expressed in HEK293 cells assessed as decrease in [14C]-AMG uptake preincubated for 10 mins followed by [14C]-AMG addition ...More data for this Ligand-Target Pair
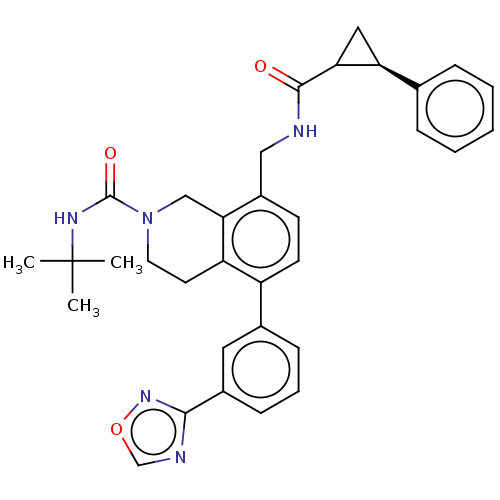
Affinity DataIC50: 1.80nMAssay Description:Displacement of [125]glucagon from human glucagon receptor expressed in CHO cellsMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Homo sapiens (Human))
Tri-Institutional Therapeutics Discovery Institute
Curated by ChEMBL
Tri-Institutional Therapeutics Discovery Institute
Curated by ChEMBL
Affinity DataIC50: 1.80nMAssay Description:Inhibition of EZH2 (unknown origin) using 3H-SAM as substrate incubated for 1 hour by filter binding methodMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Homo sapiens (Human))
Tri-Institutional Therapeutics Discovery Institute
Curated by ChEMBL
Tri-Institutional Therapeutics Discovery Institute
Curated by ChEMBL
Affinity DataIC50: 1.90nMAssay Description:Inhibition of EZH2 (unknown origin) using 3H-SAM as substrate incubated for 1 hour by filter binding methodMore data for this Ligand-Target Pair
Affinity DataIC50: 1.90nMAssay Description:Displacement of [125I]glucagon from human GCGR expressed in CHO cells after 3 hrs by scintillation proximity assayMore data for this Ligand-Target Pair
TargetNeuraminidase(Influenza A virus (A/Puerto Rico/8/34/Mount Sinai(...)
Shandong University
Curated by ChEMBL
Shandong University
Curated by ChEMBL
Affinity DataIC50: 1.90nMAssay Description:Inhibition of Influenza A virus A/PR/8/1934 (H1N1) neuraminidase using MUNANA as substrate preincubated for 10 mins followed by substrate addition an...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Homo sapiens (Human))
Tri-Institutional Therapeutics Discovery Institute
Curated by ChEMBL
Tri-Institutional Therapeutics Discovery Institute
Curated by ChEMBL
Affinity DataIC50: 2nMAssay Description:Inhibition of EZH2 (unknown origin) using 3H-SAM as substrate incubated for 1 hour by filter binding methodMore data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:Inhibition of human DGAT2 expressed in Sf9 cell membranes assessed as triolein formation by LC/MS/MS analysis using oleoyl-CoA as substrateMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Homo sapiens (Human))
Tri-Institutional Therapeutics Discovery Institute
Curated by ChEMBL
Tri-Institutional Therapeutics Discovery Institute
Curated by ChEMBL
Affinity DataIC50: 2.10nMAssay Description:Inhibition of EZH2 (unknown origin) using 3H-SAM as substrate incubated for 1 hour by filter binding methodMore data for this Ligand-Target Pair
TargetSodium/glucose cotransporter 2(Homo sapiens (Human))
Guangdong Pharmaceutical University
Curated by ChEMBL
Guangdong Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 2.20nMAssay Description:Inhibition of SGLT2 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 2.20nMAssay Description:DGAT2 activity was determined by measuring the amount of enzymatic product triolein (1,2,3-Tri(cis-9-octadecenoyl)glycerol) using the membrane prep m...More data for this Ligand-Target Pair
TargetSodium/glucose cotransporter 2(Homo sapiens (Human))
Guangdong Pharmaceutical University
Curated by ChEMBL
Guangdong Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 2.30nMAssay Description:Inhibition of SGLT2 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 2.5nMAssay Description:Displacement of [125I]glucagon from human GCGR expressed in CHO cells after 3 hrs by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.5nMAssay Description:Inhibition of Influenza A virus A/California/04/2009 (H1N1) Neuraminidase H274Y mutant using MUNANA as substrate preincubated for 10 mins followed by...More data for this Ligand-Target Pair
TargetNeuraminidase(Influenza A virus (A/Puerto Rico/8/34/Mount Sinai(...)
Shandong University
Curated by ChEMBL
Shandong University
Curated by ChEMBL
Affinity DataIC50: 2.60nMAssay Description:Inhibition of Influenza A virus A/PR/8/1934 (H1N1) neuraminidase using MUNANA as substrate preincubated for 10 mins followed by substrate addition an...More data for this Ligand-Target Pair
TargetNeuraminidase(Influenza A virus (A/Puerto Rico/8/34/Mount Sinai(...)
Shandong University
Curated by ChEMBL
Shandong University
Curated by ChEMBL
Affinity DataIC50: 2.70nMAssay Description:Inhibition of Influenza A virus A/PR/8/1934 (H1N1) neuraminidase using MUNANA as substrate preincubated for 10 mins followed by substrate addition an...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Homo sapiens (Human))
Tri-Institutional Therapeutics Discovery Institute
Curated by ChEMBL
Tri-Institutional Therapeutics Discovery Institute
Curated by ChEMBL
Affinity DataIC50: 2.90nMAssay Description:Inhibition of EZH2 (unknown origin) using 3H-SAM as substrate incubated for 1 hour by filter binding methodMore data for this Ligand-Target Pair
Affinity DataIC50: 3nMAssay Description:DGAT2 activity was determined by measuring the amount of enzymatic product triolein (1,2,3-Tri(cis-9-octadecenoyl)glycerol) using the membrane prep m...More data for this Ligand-Target Pair
Affinity DataIC50: 3nMAssay Description:Inhibition of human DGAT2 expressed in Sf9 cell membranes assessed as triolein formation by LC/MS/MS analysis using oleoyl-CoA as substrateMore data for this Ligand-Target Pair
TargetSodium/glucose cotransporter 2(Homo sapiens (Human))
Guangdong Pharmaceutical University
Curated by ChEMBL
Guangdong Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 3.10nMAssay Description:Inhibition of SGLT2 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 3.30nMAssay Description:Inhibition of human URAT1 mediated 14C-uric acid uptake expressed in HEK293 cells using 14C-uric acid as substrate incubated for 30 mins by liquid sc...More data for this Ligand-Target Pair
Ligand Info
TargetNeuraminidase(Influenza A virus (A/Puerto Rico/8/34/Mount Sinai(...)
Shandong University
Curated by ChEMBL
Shandong University
Curated by ChEMBL
Affinity DataIC50: 3.40nMAssay Description:Inhibition of Influenza A virus A/PR/8/1934 (H1N1) neuraminidase using MUNANA as substrate preincubated for 10 mins followed by substrate addition an...More data for this Ligand-Target Pair
TargetNeuraminidase(Influenza A virus (A/Puerto Rico/8/34/Mount Sinai(...)
Shandong University
Curated by ChEMBL
Shandong University
Curated by ChEMBL
Affinity DataIC50: 3.40nMAssay Description:Inhibition of Influenza A virus A/PR/8/1934 (H1N1) neuraminidase using MUNANA as substrate preincubated for 10 mins followed by substrate addition an...More data for this Ligand-Target Pair
Affinity DataIC50: 3.60nMAssay Description:DGAT2 activity was determined by measuring the amount of enzymatic product triolein (1,2,3-Tri(cis-9-octadecenoyl)glycerol) using the membrane prep m...More data for this Ligand-Target Pair
Affinity DataIC50: 3.60nMAssay Description:DGAT2 activity was determined by measuring the amount of enzymatic product triolein (1,2,3-Tri(cis-9-octadecenoyl)glycerol) using the membrane prep m...More data for this Ligand-Target Pair
TargetSodium/glucose cotransporter 2(Homo sapiens (Human))
Guangdong Pharmaceutical University
Curated by ChEMBL
Guangdong Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 4nMAssay Description:Inhibition of SGLT2 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Inhibition of human DGAT2 expressed in Sf9 cell membranes assessed as triolein formation by LC/MS/MS analysis using oleoyl-CoA as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 4.10nMAssay Description:Displacement of [125]glucagon from human glucagon receptor expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 4.10nMAssay Description:Displacement of [125I]glucagon from human GCGR expressed in CHO cells after 3 hrs by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4.10nMAssay Description:Displacement of [125]glucagon from human glucagon receptor expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 4.20nMAssay Description:Inhibition of human URAT1 mediated 14C-uric acid uptake expressed in HEK293 cells using 14C-uric acid as substrate incubated for 30 mins by liquid sc...More data for this Ligand-Target Pair
Affinity DataIC50: 4.30nMAssay Description:Displacement of [125I]glucagon from human GCGR expressed in CHO cells after 3 hrs by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4.60nMAssay Description:Displacement of [125I]glucagon from human GCGR expressed in CHO cells after 3 hrs by scintillation proximity assayMore data for this Ligand-Target Pair
TargetNeuraminidase(Influenza A virus (A/Puerto Rico/8/34/Mount Sinai(...)
Shandong University
Curated by ChEMBL
Shandong University
Curated by ChEMBL
Affinity DataIC50: 4.70nMAssay Description:Inhibition of Influenza A virus A/PR/8/1934 (H1N1) neuraminidase using MUNANA as substrate preincubated for 10 mins followed by substrate addition an...More data for this Ligand-Target Pair
Affinity DataIC50: 4.80nMAssay Description:Displacement of [125]glucagon from human glucagon receptor expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 4.90nMAssay Description:Displacement of [125]glucagon from human glucagon receptor expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 5.20nMAssay Description:Displacement of [125I]glucagon from human GCGR expressed in CHO cells after 3 hrs by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5.70nMAssay Description:Antagonist activity at human GCGR expressed in CHO cells assessed as inhibition of glucagon-induced cAMP accumulation preincubated for 30 mins prior ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.78nMAssay Description:DGAT2 activity was determined by measuring the amount of enzymatic product triolein (1,2,3-Tri(cis-9-octadecenoyl)glycerol) using the membrane prep m...More data for this Ligand-Target Pair
Affinity DataIC50: 5.80nMAssay Description:DGAT2 activity was determined by measuring the amount of enzymatic product triolein (1,2,3-Tri(cis-9-octadecenoyl)glycerol) using the membrane prep m...More data for this Ligand-Target Pair


 3D Structure (crystal)
3D Structure (crystal)