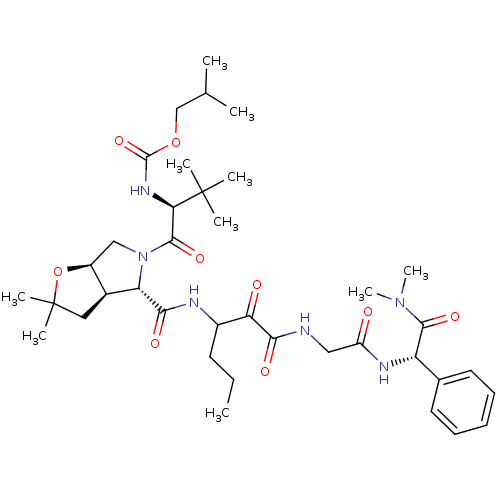
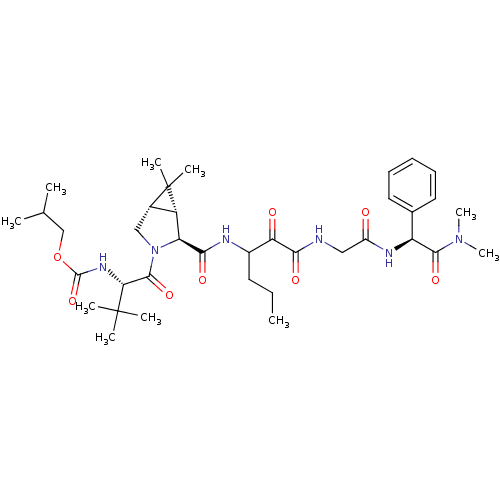
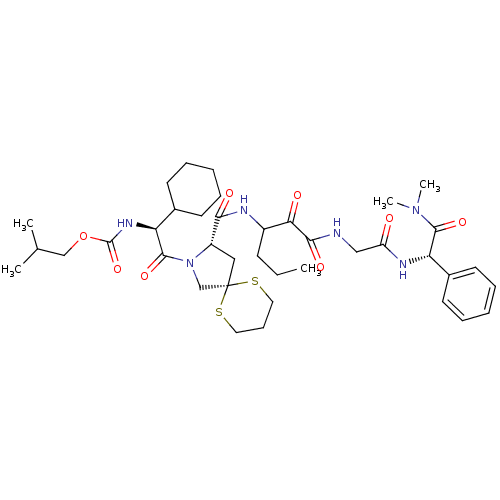
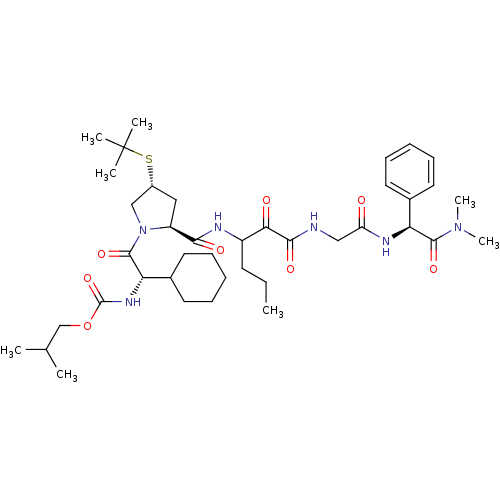
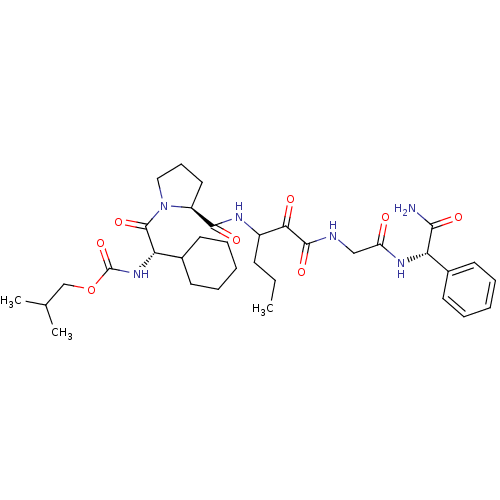
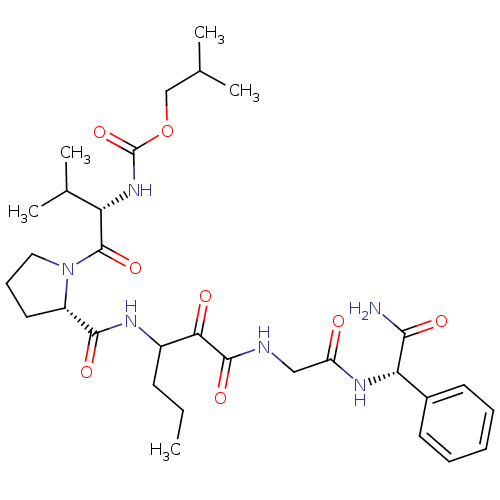
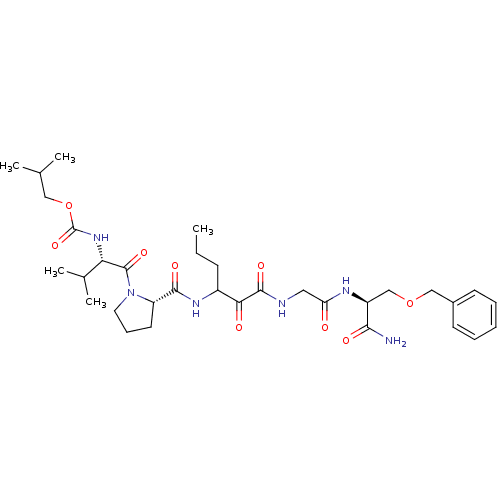
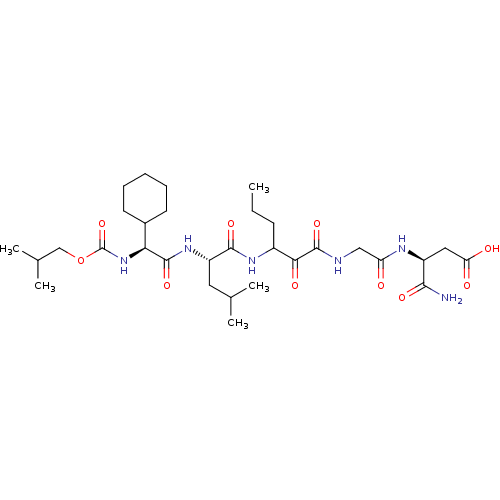
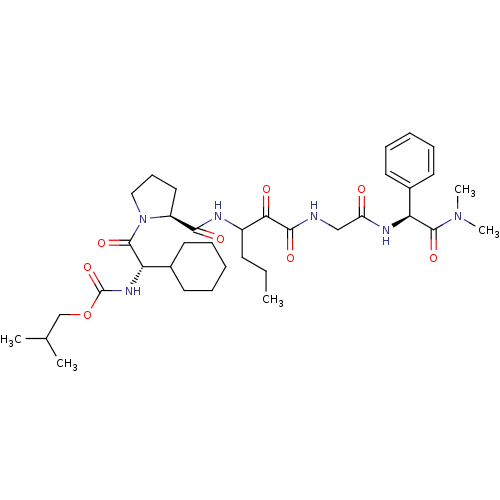
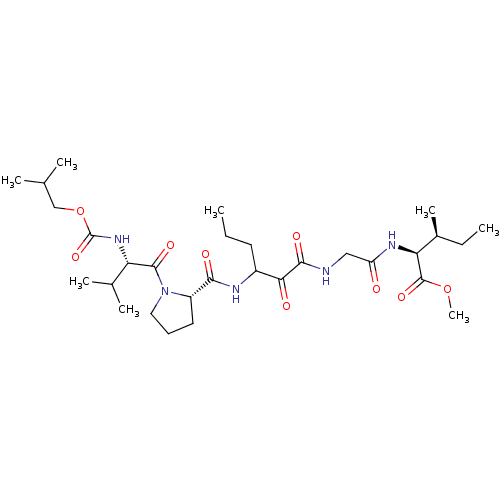
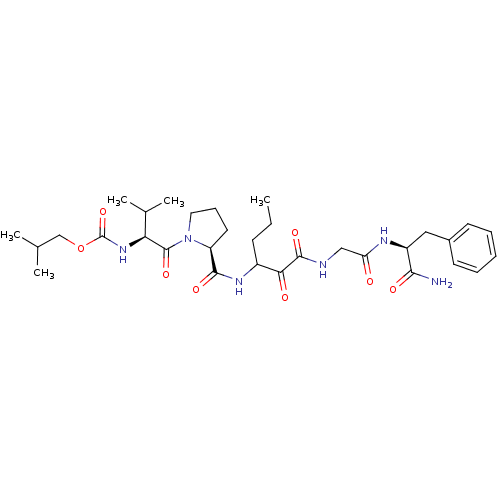
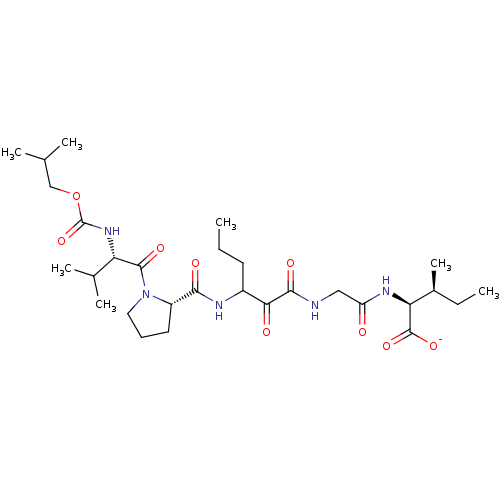
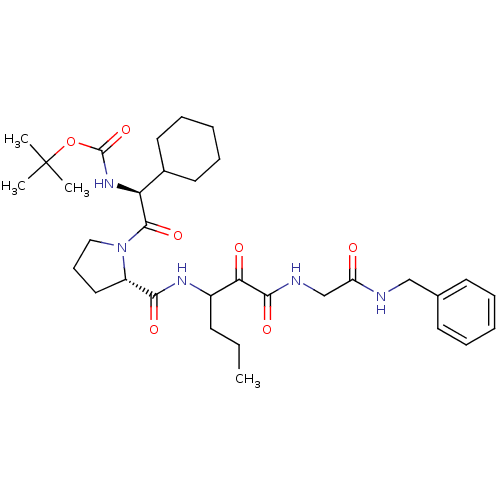
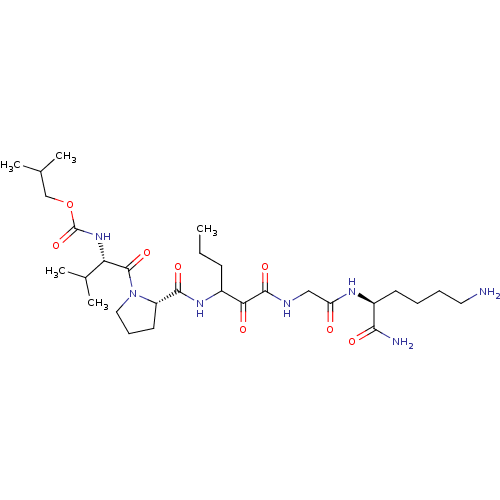
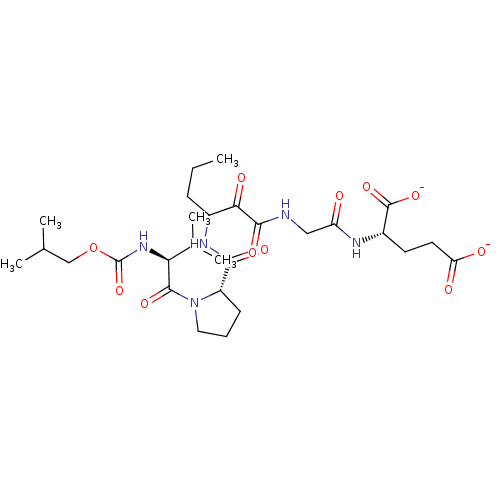
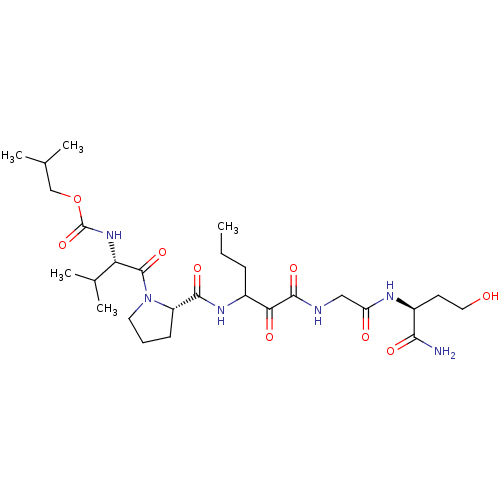
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 2nM ΔG°: -50.5kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
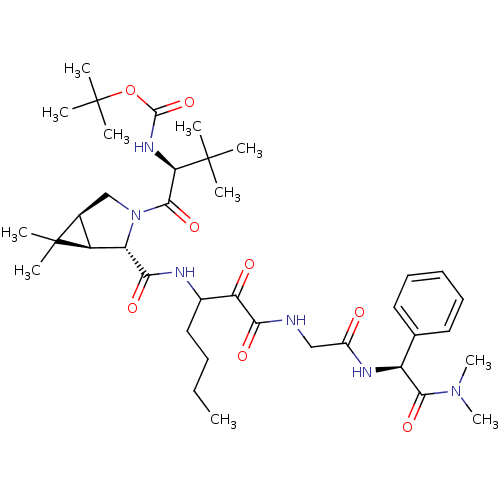
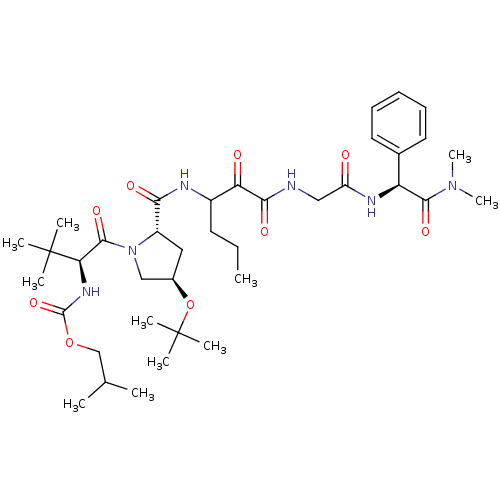
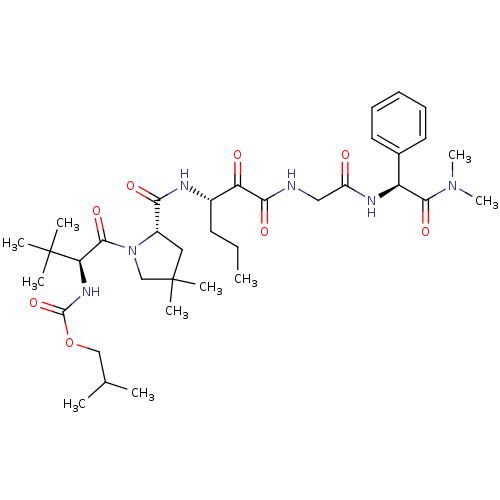
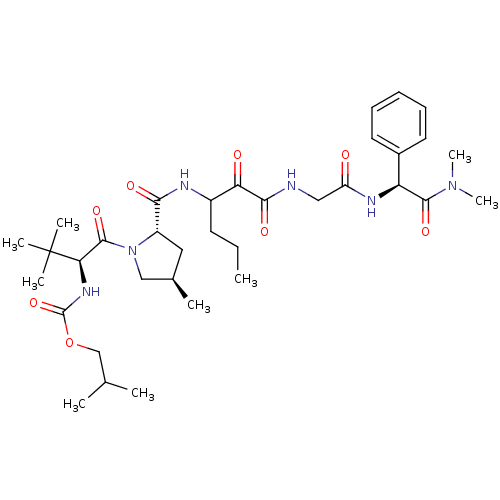
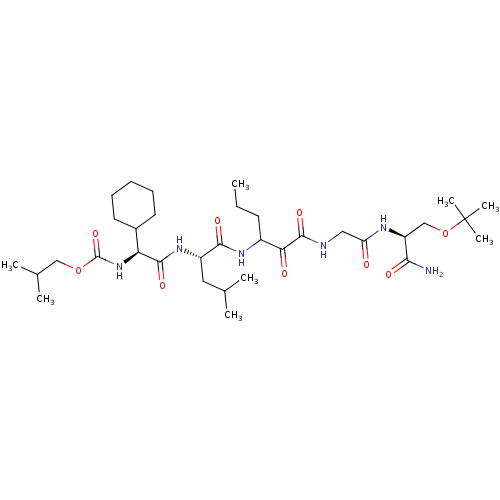
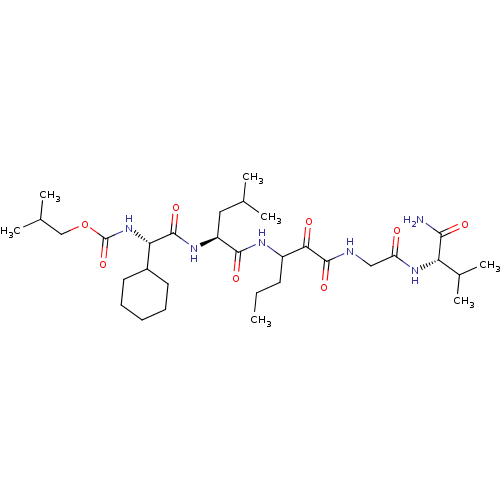
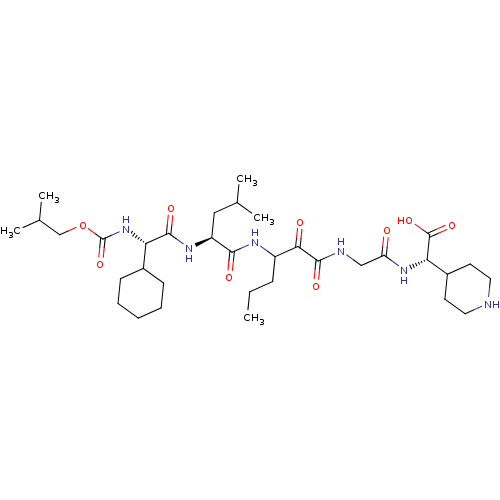
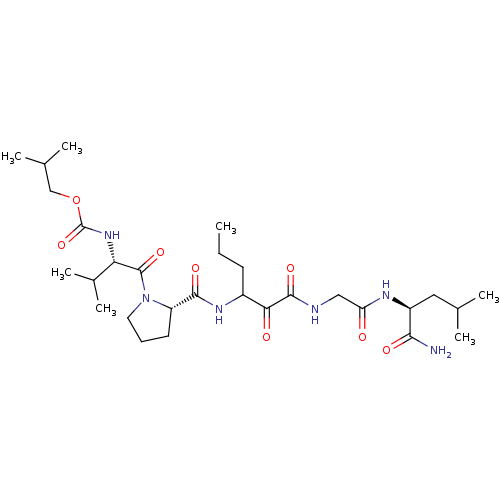
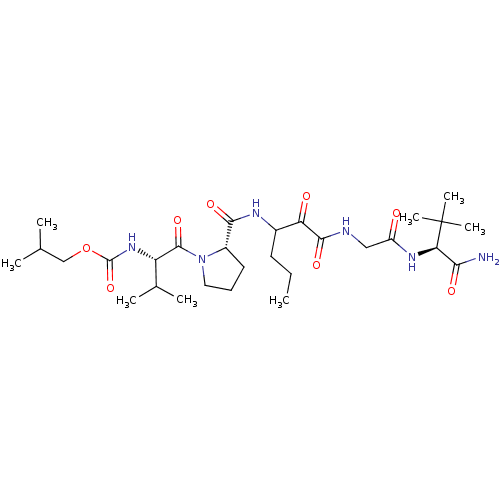
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 3.80nM ΔG°: -48.9kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
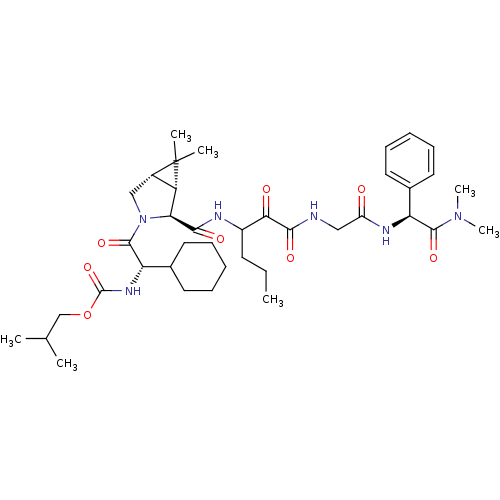
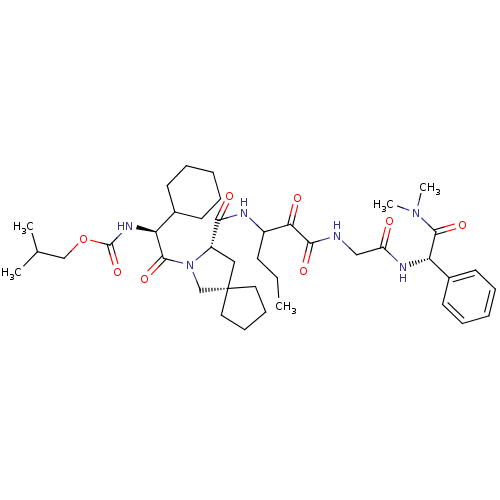
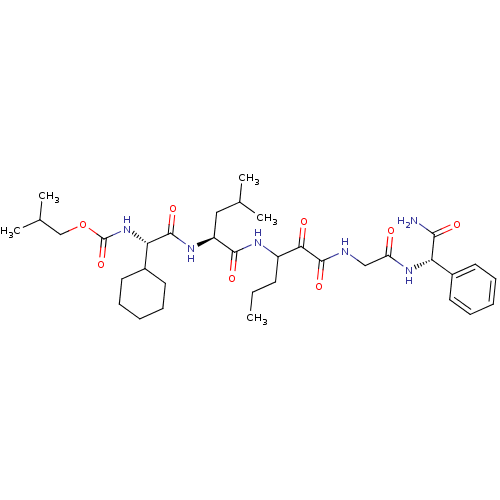
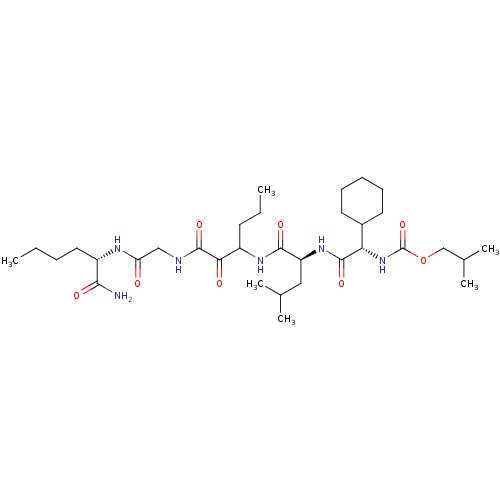
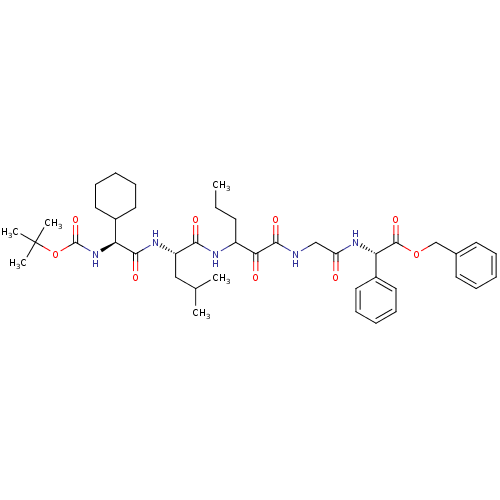
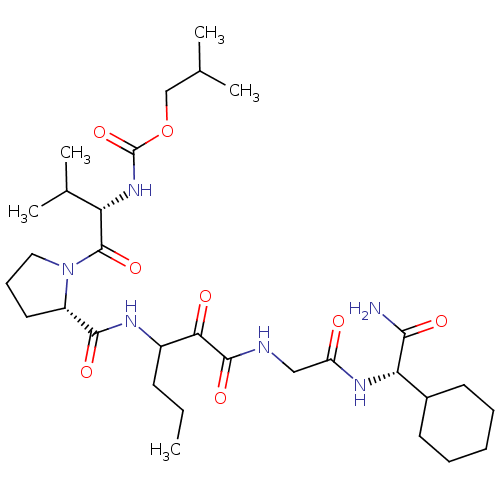
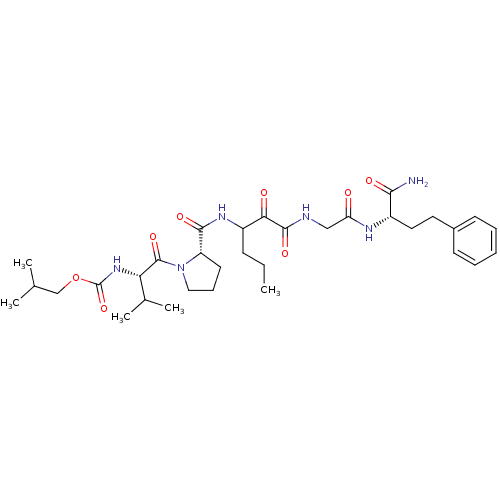
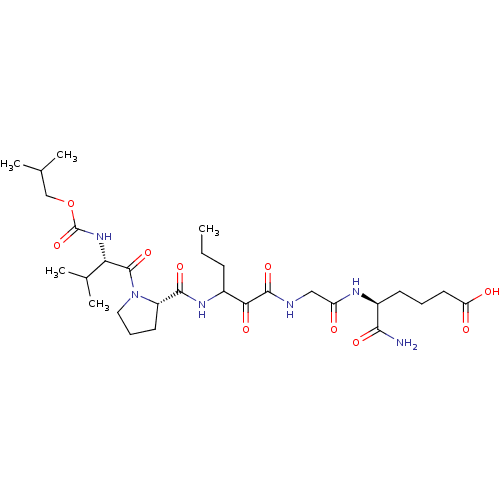
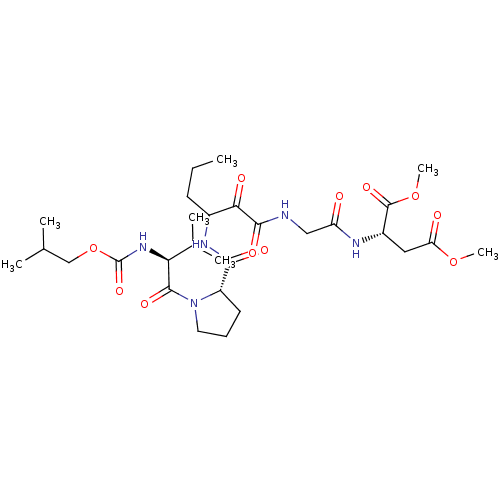
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 10nM ΔG°: -46.4kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
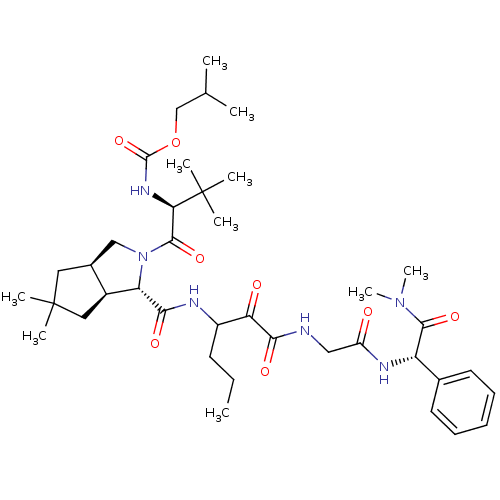
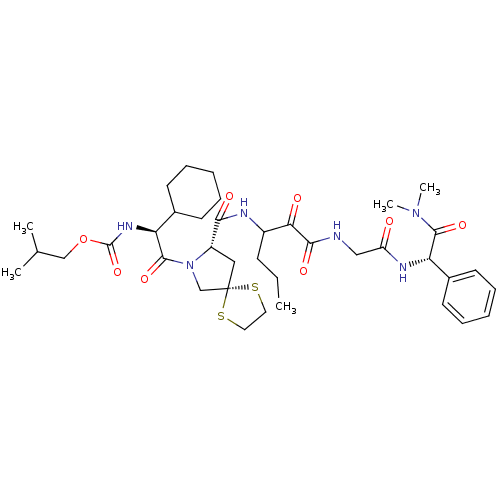
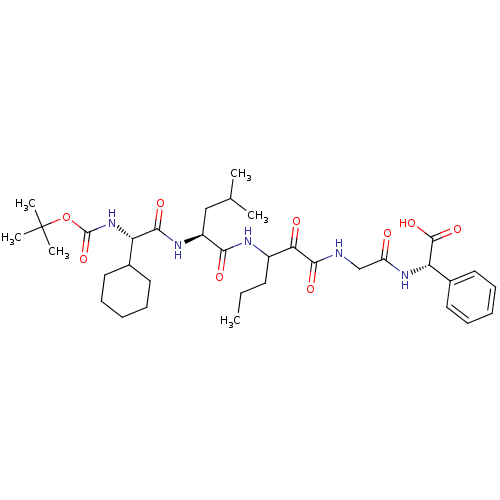
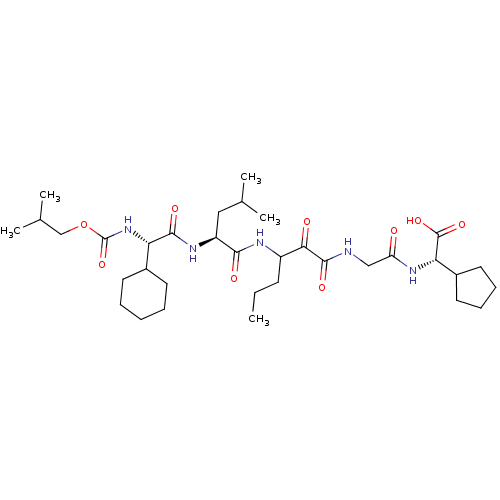
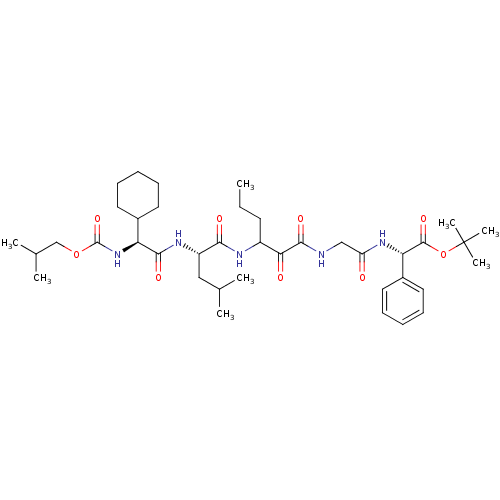
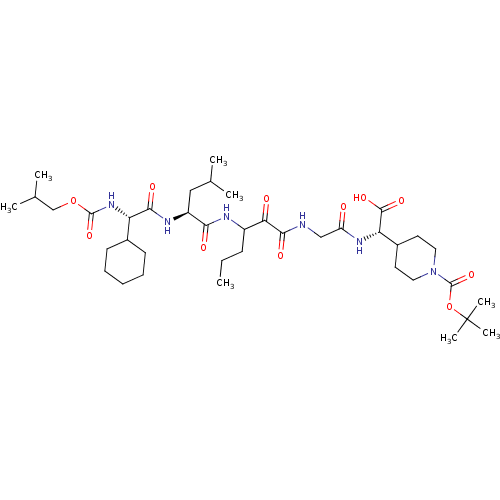
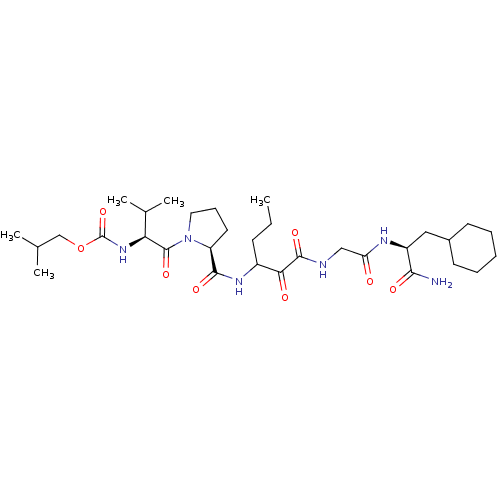
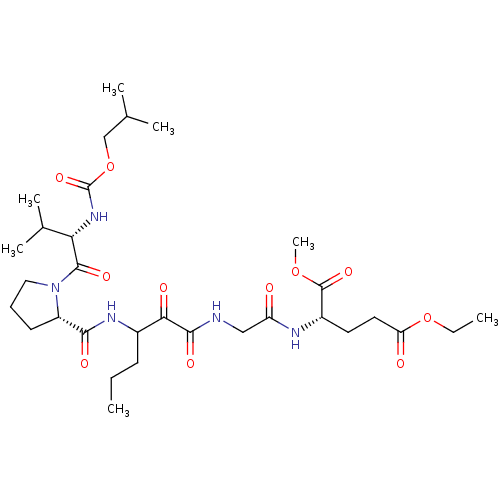
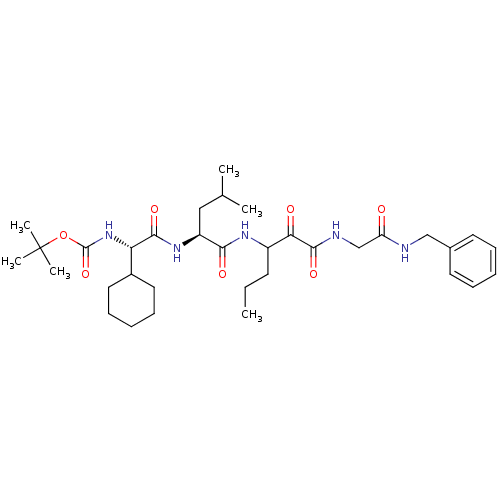
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 10nM ΔG°: -46.4kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 10nM ΔG°: -46.4kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 14nM ΔG°: -45.6kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 15nM ΔG°: -45.4kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 16nM ΔG°: -45.2kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 19nM ΔG°: -44.8kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 26nM ΔG°: -44.0kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 26nM ΔG°: -44.0kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 28nM ΔG°: -43.8kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 36nM ΔG°: -43.2kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 66nMAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 98nMAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 120nM ΔG°: -40.2kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 120nMAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 140nM ΔG°: -39.8kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 280nMAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 330nMAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 360nMAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 460nMAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 600nMAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 650nMAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 2.10E+3nMAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 2.60E+3nMAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 2.60E+3nM ΔG°: -32.4kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 3.30E+3nMAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 3.50E+3nM ΔG°: -31.7kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 6.40E+3nMAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 6.70E+3nM ΔG°: -30.0kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 8.00E+3nM ΔG°: -29.6kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 8.40E+3nMAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 1.00E+4nM ΔG°: -29.0kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 1.20E+4nM ΔG°: -28.6kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 1.50E+4nM ΔG°: -28.0kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 1.50E+4nMAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 1.50E+4nM ΔG°: -28.0kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 1.60E+4nM ΔG°: -27.8kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 1.80E+4nM ΔG°: -27.5kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 2.00E+4nM ΔG°: -27.3kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 4.50E+4nMAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 5.30E+4nM ΔG°: -24.8kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 5.50E+4nM ΔG°: -24.7kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 8.10E+4nMAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 8.20E+4nM ΔG°: -23.7kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: >1.00E+5nM ΔG°: >-23.2kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: >1.00E+5nM ΔG°: >-23.2kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 1.11E+5nM ΔG°: -23.0kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair

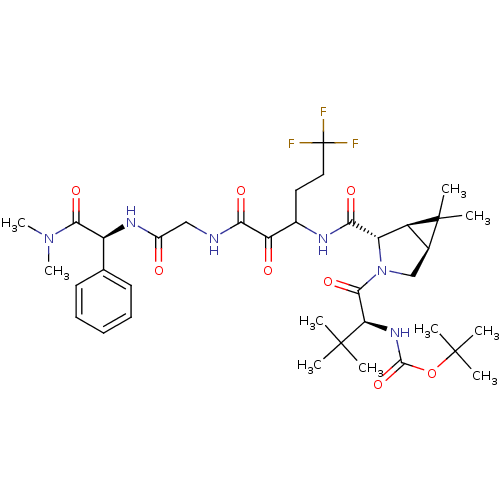
 3D Structure (crystal)
3D Structure (crystal)