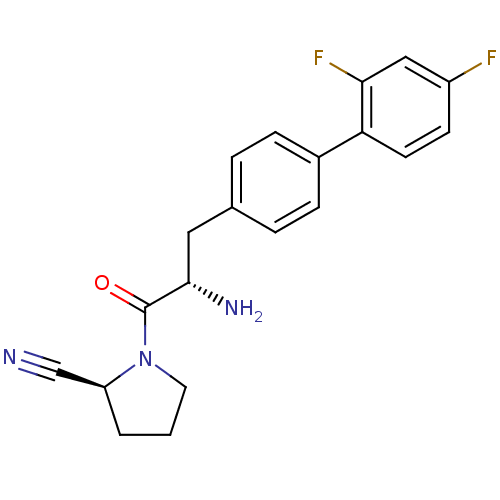
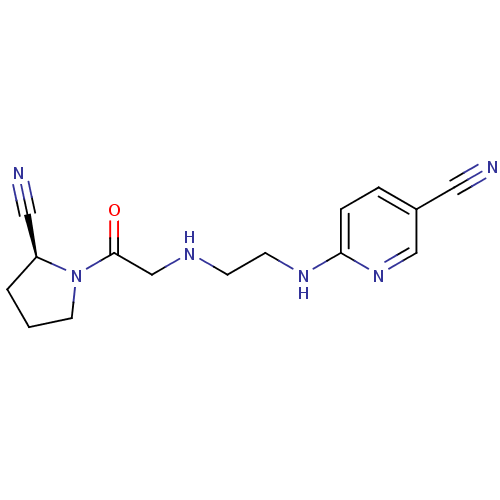
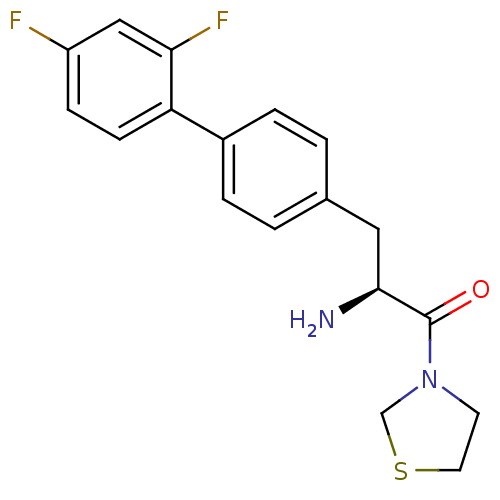
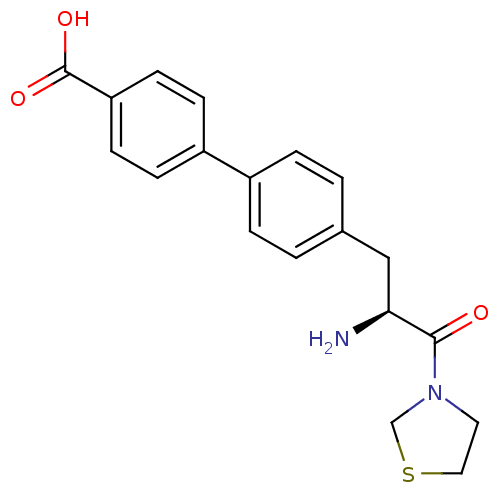
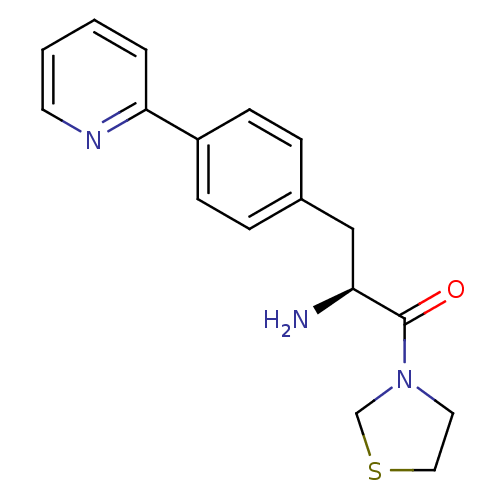
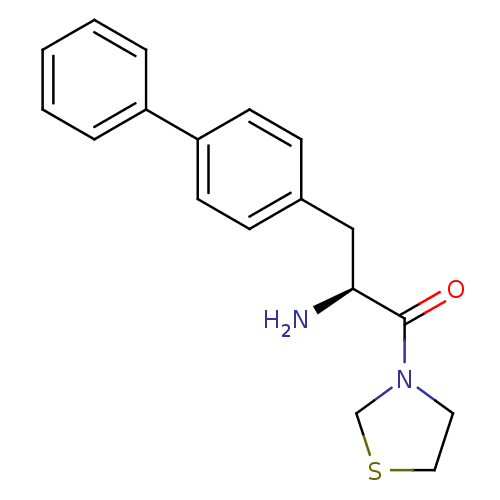
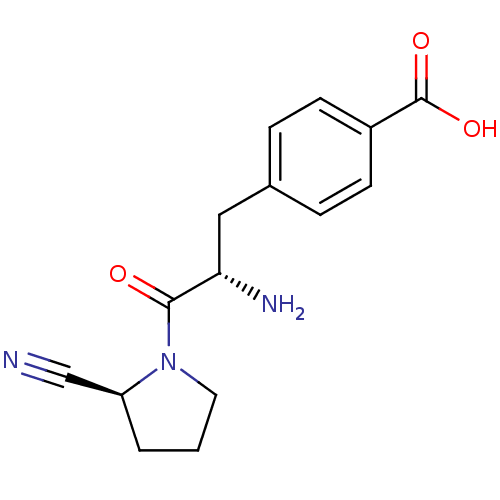
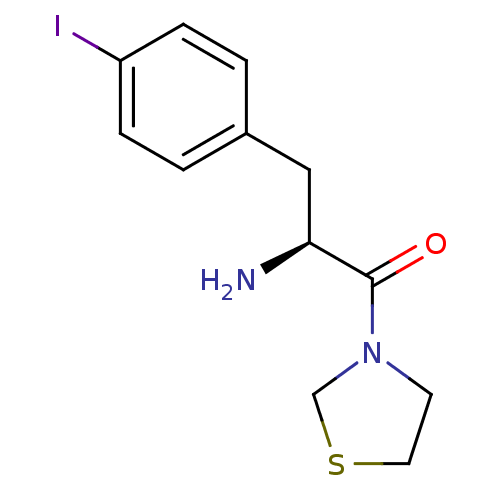
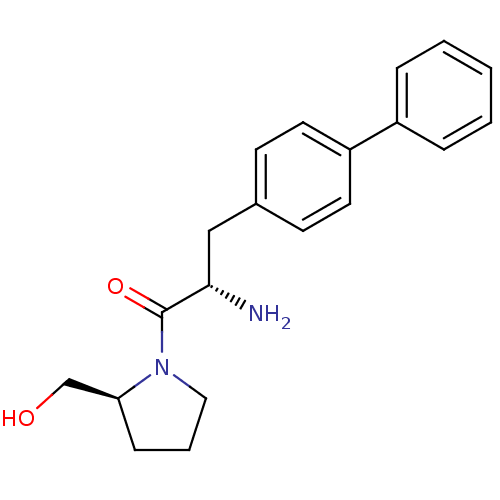
Affinity DataKi: 2.20nM ΔG°: -49.4kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
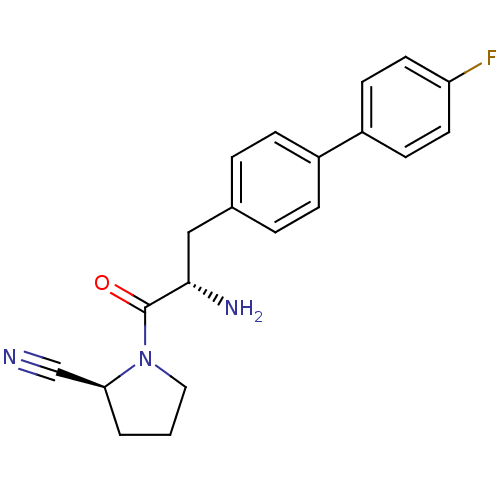
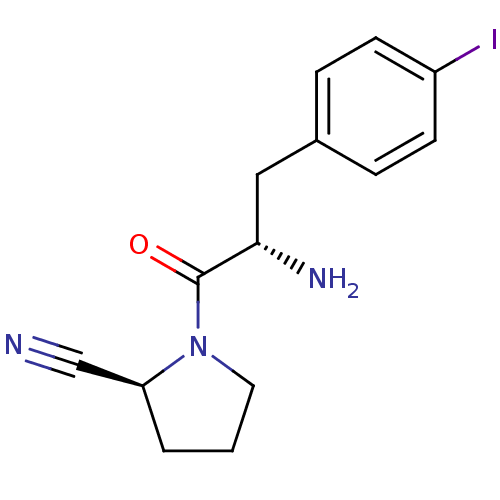
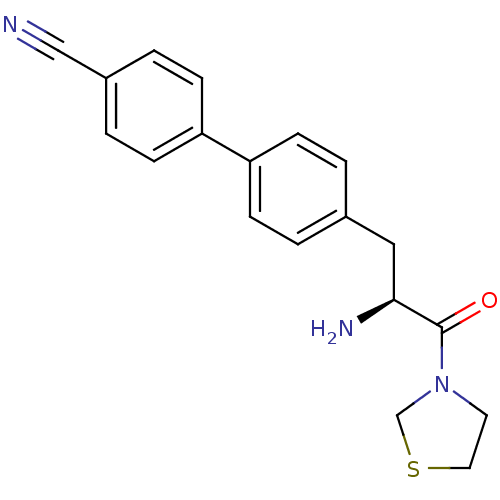
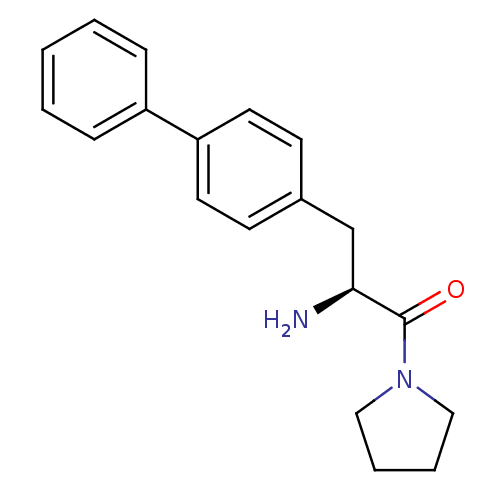
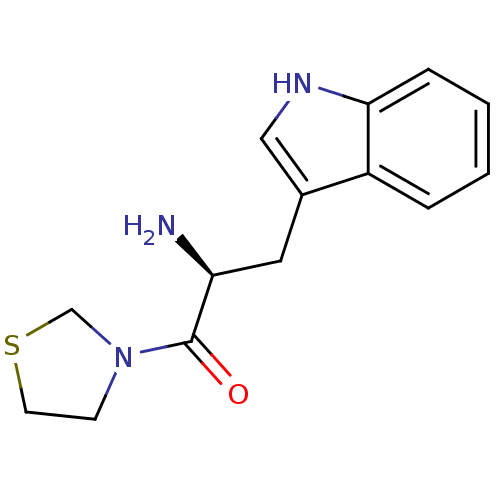
Affinity DataKi: 3.10nM ΔG°: -48.6kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
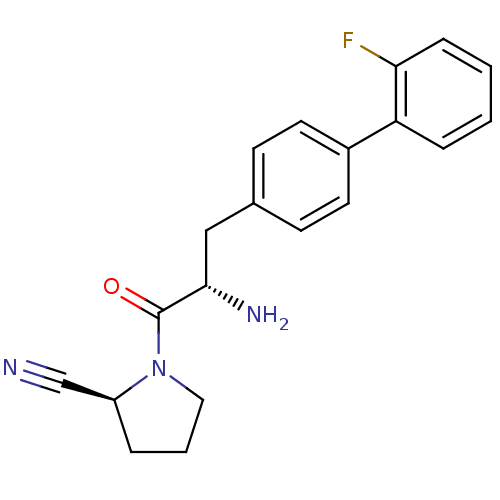
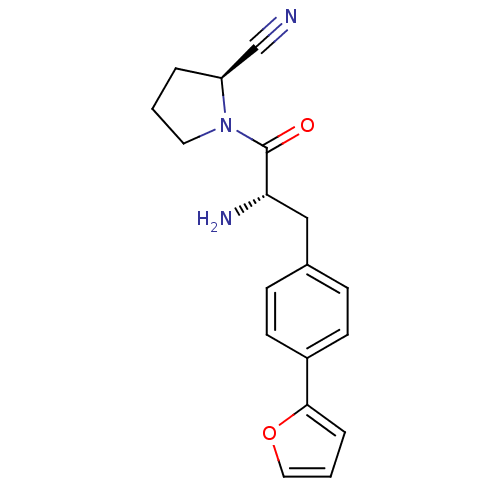
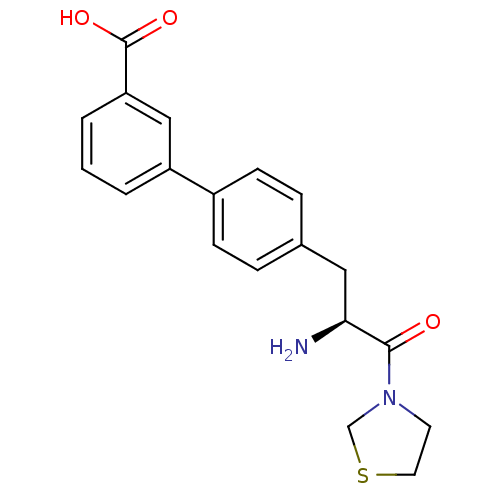
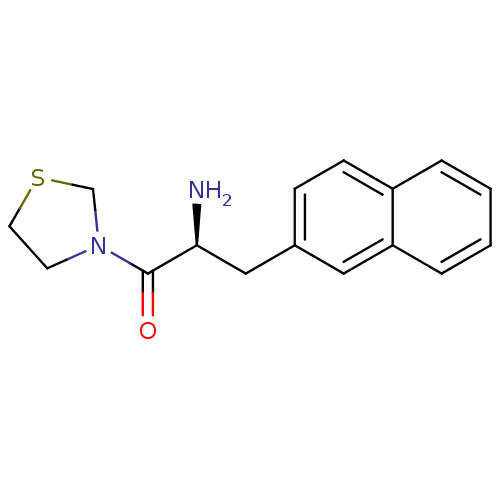
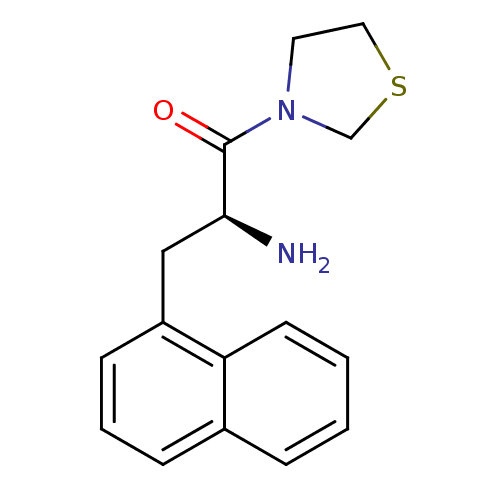
Affinity DataKi: 5.30nM ΔG°: -47.2kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
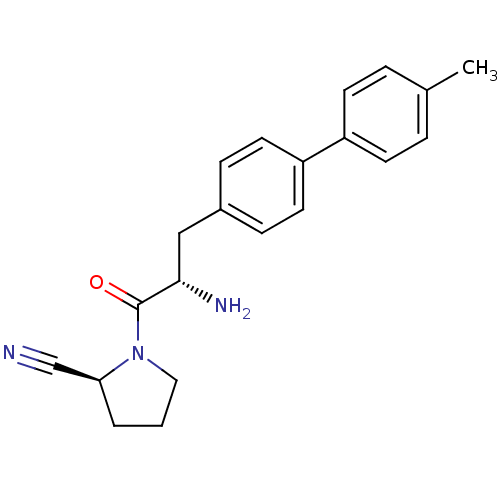
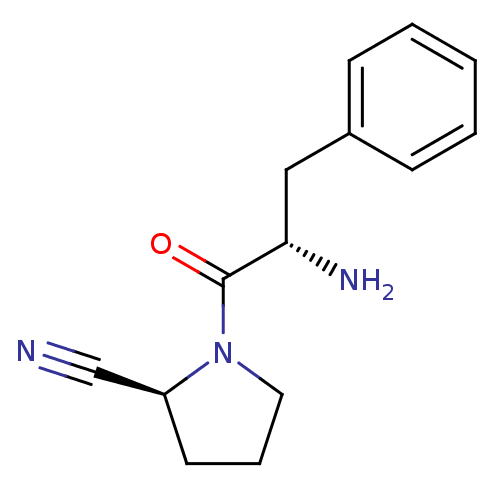
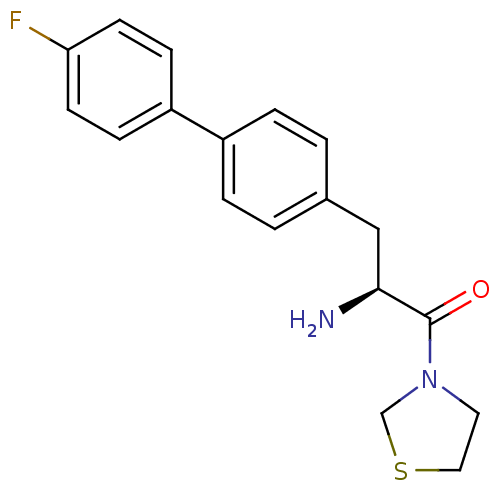
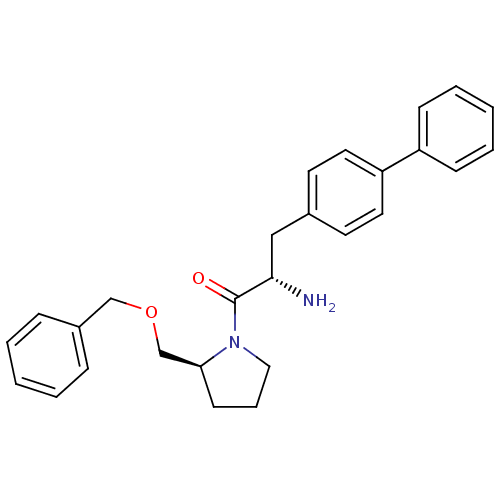
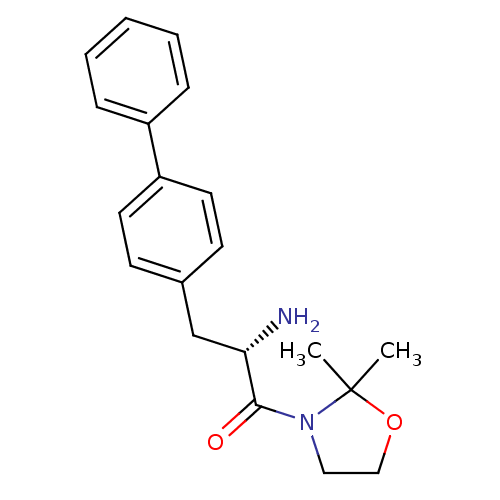
Affinity DataKi: 13nM ΔG°: -45.0kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 20nM ΔG°: -43.9kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 26nM ΔG°: -43.3kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 27nM ΔG°: -43.2kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 34nM ΔG°: -42.6kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 36nM ΔG°: -42.5kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 63nM ΔG°: -41.1kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 96nM ΔG°: -40.1kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 160nM ΔG°: -38.8kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 166nM ΔG°: -38.7kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 170nM ΔG°: -38.6kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 310nM ΔG°: -37.1kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 355nM ΔG°: -36.8kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 360nM ΔG°: -36.8kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 470nM ΔG°: -36.1kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 980nM ΔG°: -34.3kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 1.16E+3nM ΔG°: -33.9kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 8.90E+3nM ΔG°: -28.8kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: >1.28E+4nM ΔG°: >-27.9kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: >1.28E+4nM ΔG°: >-27.9kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: >1.28E+4nM ΔG°: >-27.9kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: >1.28E+4nM ΔG°: >-27.9kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: >1.28E+4nM ΔG°: >-27.9kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
TargetIntegrin alpha-V/beta-5(Homo sapiens (Human))
Johnson & Johnson Pharmaceutical Research And Development
Curated by ChEMBL
Johnson & Johnson Pharmaceutical Research And Development
Curated by ChEMBL
Affinity DataIC50: 0.210nMAssay Description:Binding affinity to Integrin alpha-v-beta-5 receptor by ELISA assayMore data for this Ligand-Target Pair
TargetIntegrin alpha-V/beta-3(Homo sapiens (Human))
Johnson & Johnson Pharmaceutical Research And Development
Curated by ChEMBL
Johnson & Johnson Pharmaceutical Research And Development
Curated by ChEMBL
Affinity DataIC50: 0.25nMAssay Description:Binding affinity to Integrin alpha-v-beta-3 receptor by ELISA assayMore data for this Ligand-Target Pair
TargetAcetylcholinesterase(Electrophorus electricus (Electric eel))
University Of Michigan
Curated by ChEMBL
University Of Michigan
Curated by ChEMBL
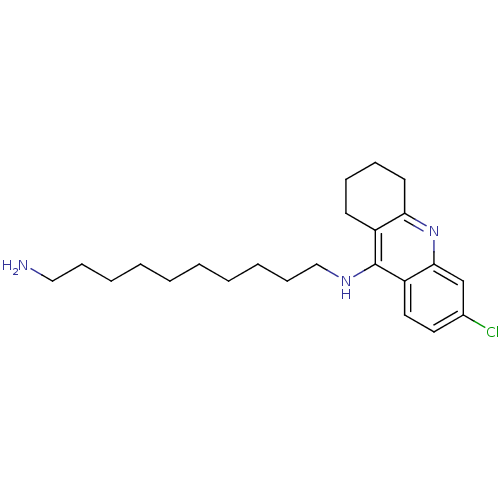
Affinity DataIC50: 0.299nMAssay Description:Inhibition of electric eel AChE preincubated for 30 mins prior to horseradish peroxidase, H2O2 and DETAPAC addition measured for 20 mins under ROS co...More data for this Ligand-Target Pair
TargetIntegrin alpha-V/beta-3(Homo sapiens (Human))
Johnson & Johnson Pharmaceutical Research And Development
Curated by ChEMBL
Johnson & Johnson Pharmaceutical Research And Development
Curated by ChEMBL
Affinity DataIC50: 0.380nMAssay Description:Binding affinity to Integrin alpha-v-beta-3 receptor by ELISA assayMore data for this Ligand-Target Pair
TargetAcetylcholinesterase(Electrophorus electricus (Electric eel))
University Of Michigan
Curated by ChEMBL
University Of Michigan
Curated by ChEMBL
Affinity DataIC50: 0.495nMAssay Description:Inhibition of electric eel AChE using acetylthiocholine as substrate preincubated for 10 mins prior to substrate addition measured every 30 seconds f...More data for this Ligand-Target Pair
TargetIntegrin alpha-V/beta-5(Homo sapiens (Human))
Johnson & Johnson Pharmaceutical Research And Development
Curated by ChEMBL
Johnson & Johnson Pharmaceutical Research And Development
Curated by ChEMBL
Affinity DataIC50: 0.5nMAssay Description:Binding affinity to Integrin alpha-v-beta-5 receptor by ELISA assayMore data for this Ligand-Target Pair
TargetIntegrin alpha-V/beta-3(Homo sapiens (Human))
Johnson & Johnson Pharmaceutical Research And Development
Curated by ChEMBL
Johnson & Johnson Pharmaceutical Research And Development
Curated by ChEMBL
Affinity DataIC50: 0.5nMAssay Description:Binding affinity to Integrin alpha-v-beta-3 receptor by ELISA assayMore data for this Ligand-Target Pair
TargetAcetylcholinesterase(Electrophorus electricus (Electric eel))
University Of Michigan
Curated by ChEMBL
University Of Michigan
Curated by ChEMBL
Affinity DataIC50: 0.604nMAssay Description:Inhibition of electric eel AChE using acetylthiocholine as substrate preincubated for 10 mins prior to substrate addition measured every 30 seconds f...More data for this Ligand-Target Pair
TargetAcetylcholinesterase(Electrophorus electricus (Electric eel))
University Of Michigan
Curated by ChEMBL
University Of Michigan
Curated by ChEMBL
Affinity DataIC50: 0.648nMAssay Description:Inhibition of electric eel AChE using acetylthiocholine as substrate preincubated for 10 mins prior to substrate addition measured every 30 seconds f...More data for this Ligand-Target Pair
TargetIntegrin alpha-V/beta-5(Homo sapiens (Human))
Johnson & Johnson Pharmaceutical Research And Development
Curated by ChEMBL
Johnson & Johnson Pharmaceutical Research And Development
Curated by ChEMBL
Affinity DataIC50: 0.680nMAssay Description:Binding affinity to Integrin alpha-v-beta-5 receptor by ELISA assayMore data for this Ligand-Target Pair
TargetAcetylcholinesterase(Electrophorus electricus (Electric eel))
University Of Michigan
Curated by ChEMBL
University Of Michigan
Curated by ChEMBL
Affinity DataIC50: 0.776nMAssay Description:Inhibition of electric eel AChE using acetylthiocholine as substrate preincubated for 10 mins prior to substrate addition measured every 30 seconds f...More data for this Ligand-Target Pair
TargetIntegrin alpha-V/beta-3(Homo sapiens (Human))
Johnson & Johnson Pharmaceutical Research And Development
Curated by ChEMBL
Johnson & Johnson Pharmaceutical Research And Development
Curated by ChEMBL
Affinity DataIC50: 0.860nMAssay Description:Binding affinity to Integrin alpha-v-beta-3 receptor by ELISA assayMore data for this Ligand-Target Pair
TargetAcetylcholinesterase(Electrophorus electricus (Electric eel))
University Of Michigan
Curated by ChEMBL
University Of Michigan
Curated by ChEMBL
Affinity DataIC50: 0.932nMAssay Description:Inhibition of electric eel AChE using acetylthiocholine as substrate preincubated for 10 mins prior to substrate addition measured every 30 seconds f...More data for this Ligand-Target Pair
TargetIntegrin alpha-V/beta-5(Homo sapiens (Human))
Johnson & Johnson Pharmaceutical Research And Development
Curated by ChEMBL
Johnson & Johnson Pharmaceutical Research And Development
Curated by ChEMBL
Affinity DataIC50: 0.960nMAssay Description:Binding affinity to Integrin alpha-v-beta-5 receptor by ELISA assayMore data for this Ligand-Target Pair
TargetIntegrin alpha-V/beta-3(Homo sapiens (Human))
Johnson & Johnson Pharmaceutical Research And Development
Curated by ChEMBL
Johnson & Johnson Pharmaceutical Research And Development
Curated by ChEMBL
Affinity DataIC50: 0.960nMAssay Description:Binding affinity to Integrin alpha-v-beta-3 receptor by ELISA assayMore data for this Ligand-Target Pair
TargetIntegrin alpha-V/beta-3(Homo sapiens (Human))
Johnson & Johnson Pharmaceutical Research And Development
Curated by ChEMBL
Johnson & Johnson Pharmaceutical Research And Development
Curated by ChEMBL
Affinity DataIC50: 1nMAssay Description:Binding affinity to Integrin alpha-v-beta-3 receptor by ELISA assayMore data for this Ligand-Target Pair
TargetAcetylcholinesterase(Electrophorus electricus (Electric eel))
University Of Michigan
Curated by ChEMBL
University Of Michigan
Curated by ChEMBL
Affinity DataIC50: 1nMAssay Description:Inhibition of electric eel AChE preincubated for 30 mins prior to horseradish peroxidase, H2O2 and DETAPAC addition measured for 20 mins under ROS co...More data for this Ligand-Target Pair
TargetAcetylcholinesterase(Electrophorus electricus (Electric eel))
University Of Michigan
Curated by ChEMBL
University Of Michigan
Curated by ChEMBL
Affinity DataIC50: 1.10nMAssay Description:Inhibition of electric eel AChE using acetylthiocholine as substrate preincubated for 10 mins prior to substrate addition measured every 30 seconds f...More data for this Ligand-Target Pair
TargetIntegrin alpha-V/beta-5(Homo sapiens (Human))
Johnson & Johnson Pharmaceutical Research And Development
Curated by ChEMBL
Johnson & Johnson Pharmaceutical Research And Development
Curated by ChEMBL
Affinity DataIC50: 1.10nMAssay Description:Binding affinity to Integrin alpha-v-beta-5 receptor by ELISA assayMore data for this Ligand-Target Pair
TargetAcetylcholinesterase(Electrophorus electricus (Electric eel))
University Of Michigan
Curated by ChEMBL
University Of Michigan
Curated by ChEMBL
Affinity DataIC50: 1.10nMAssay Description:Inhibition of electric eel AChE using acetylthiocholine as substrate preincubated for 10 mins prior to substrate addition measured every 30 seconds f...More data for this Ligand-Target Pair
TargetAcetylcholinesterase(Electrophorus electricus (Electric eel))
University Of Michigan
Curated by ChEMBL
University Of Michigan
Curated by ChEMBL
Affinity DataIC50: 1.20nMAssay Description:Inhibition of electric eel AChE preincubated for 30 mins prior to horseradish peroxidase, H2O2 and DETAPAC addition measured for 20 mins under ROS co...More data for this Ligand-Target Pair
TargetAcetylcholinesterase(Electrophorus electricus (Electric eel))
University Of Michigan
Curated by ChEMBL
University Of Michigan
Curated by ChEMBL
Affinity DataIC50: 1.40nMAssay Description:Inhibition of electric eel AChE preincubated for 30 mins prior to horseradish peroxidase, H2O2 and DETAPAC addition measured for 20 mins under ROS co...More data for this Ligand-Target Pair
TargetAcetylcholinesterase(Electrophorus electricus (Electric eel))
University Of Michigan
Curated by ChEMBL
University Of Michigan
Curated by ChEMBL
Affinity DataIC50: 1.5nMAssay Description:Inhibition of electric eel AChE preincubated for 30 mins prior to horseradish peroxidase, H2O2 and DETAPAC addition measured for 20 mins under ROS co...More data for this Ligand-Target Pair
TargetAcetylcholinesterase(Electrophorus electricus (Electric eel))
University Of Michigan
Curated by ChEMBL
University Of Michigan
Curated by ChEMBL
Affinity DataIC50: 1.90nMAssay Description:Inhibition of electric eel AChE preincubated for 30 mins prior to horseradish peroxidase, H2O2 and DETAPAC addition measured for 20 mins under ROS co...More data for this Ligand-Target Pair