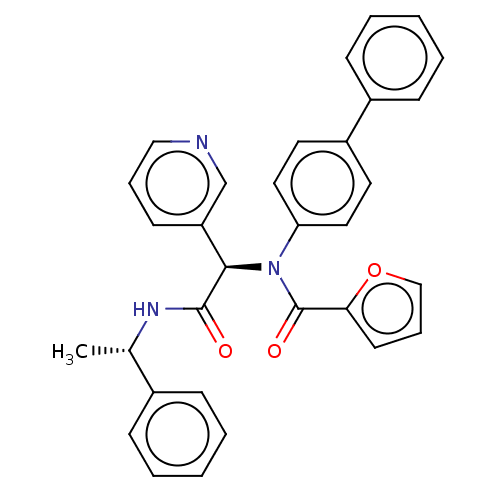
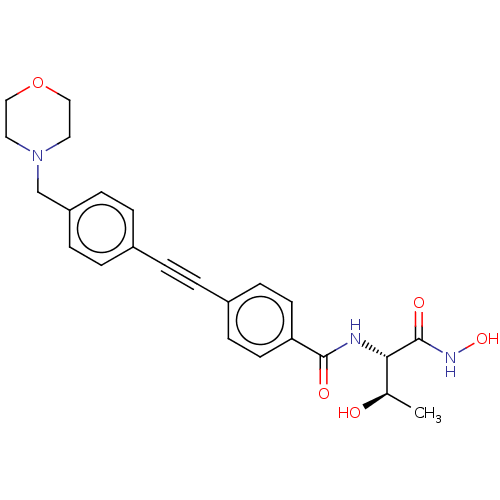
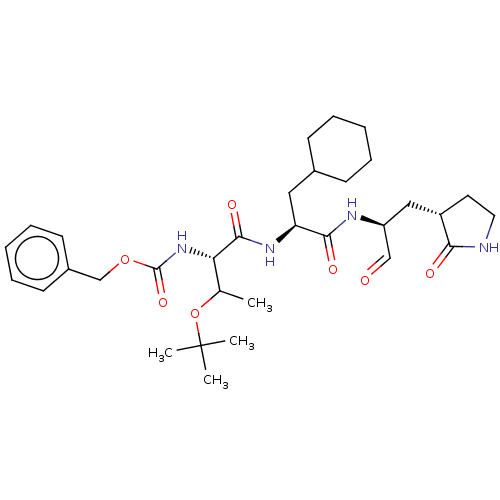
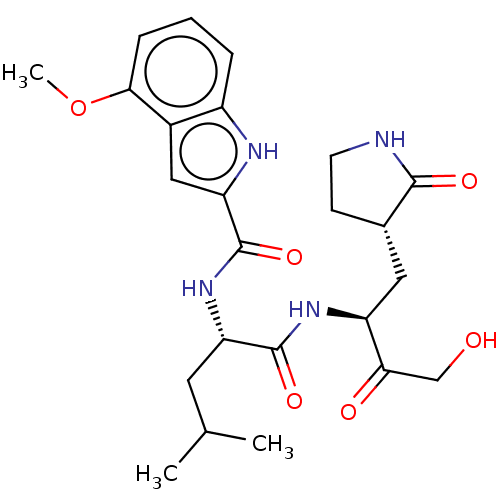
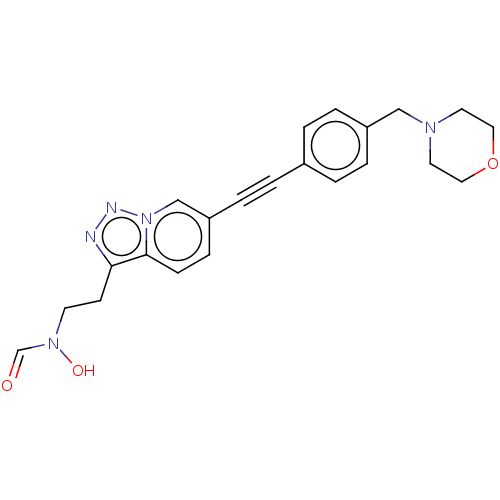
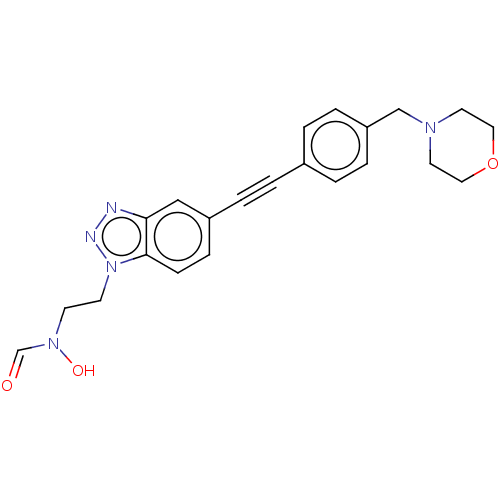
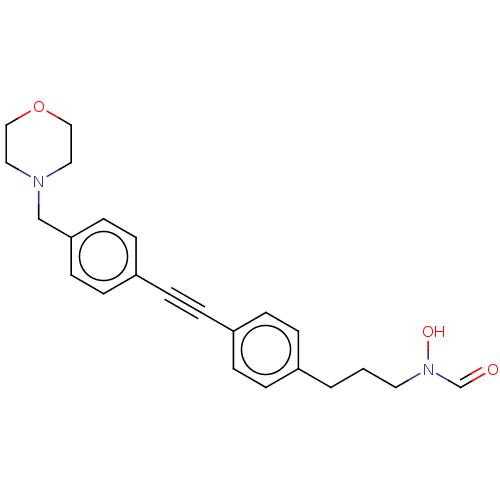
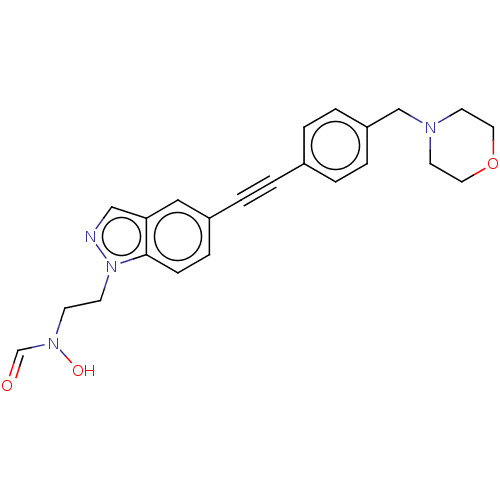
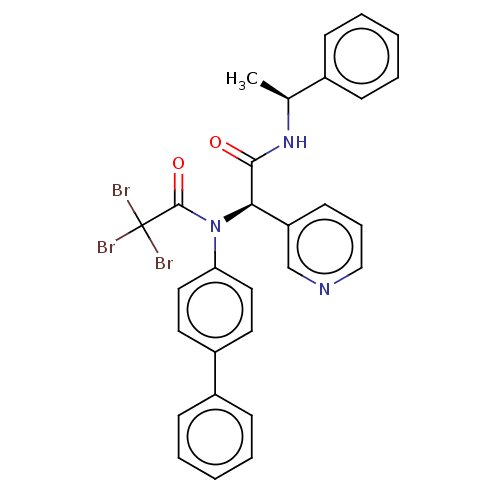
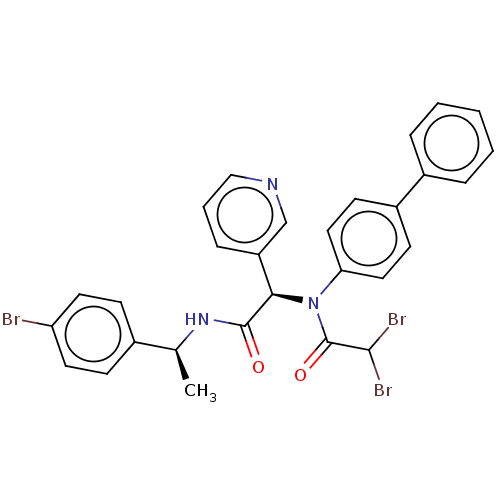
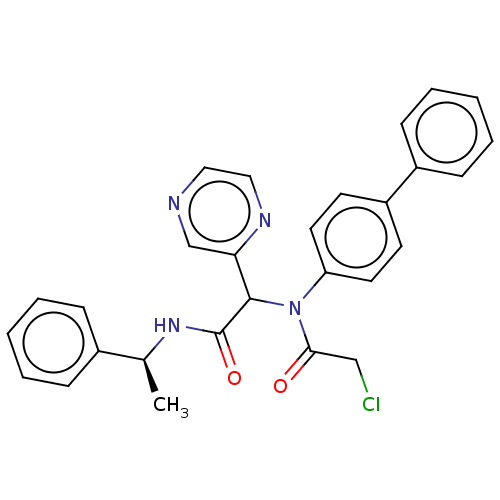
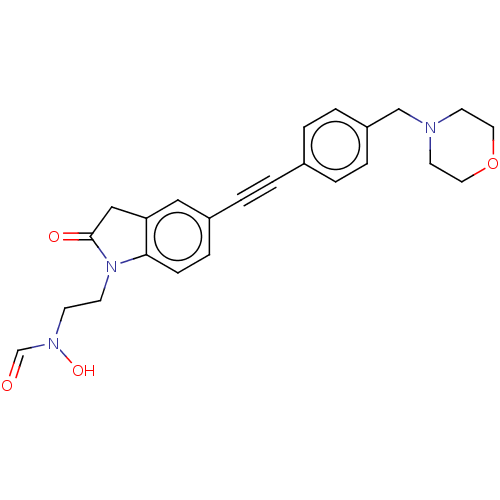
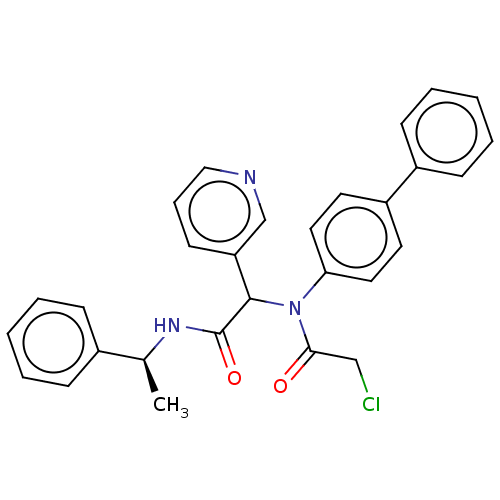
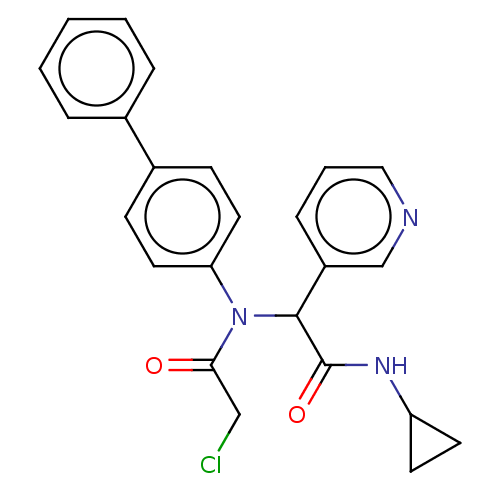
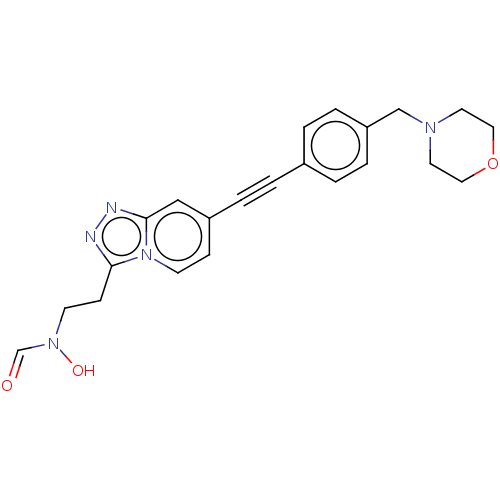
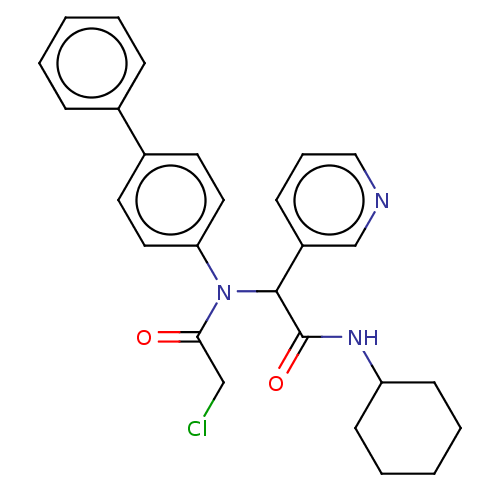
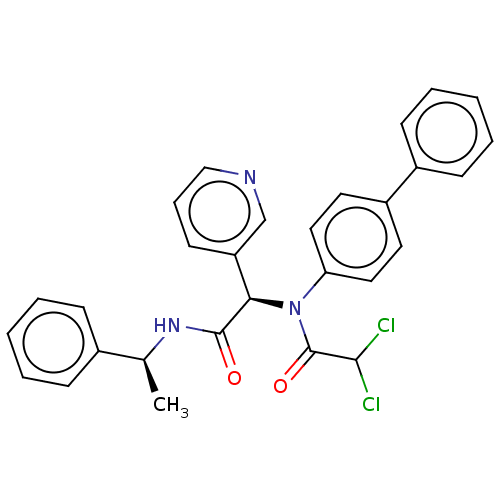
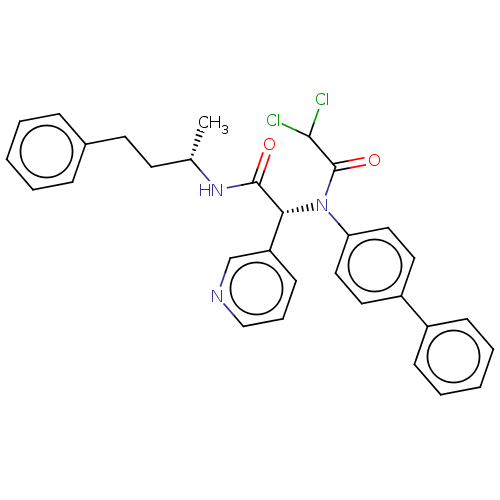
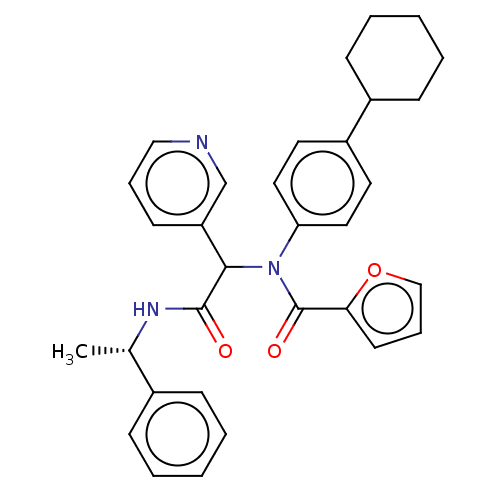
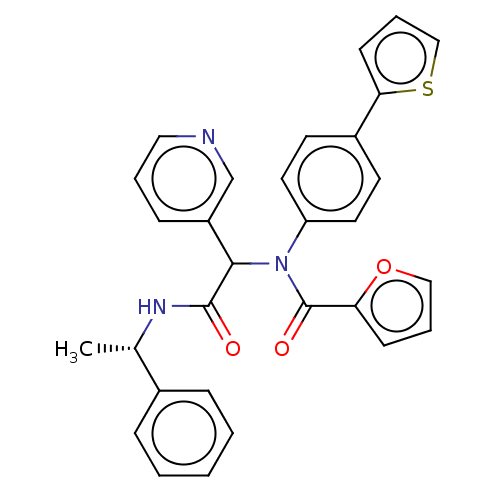
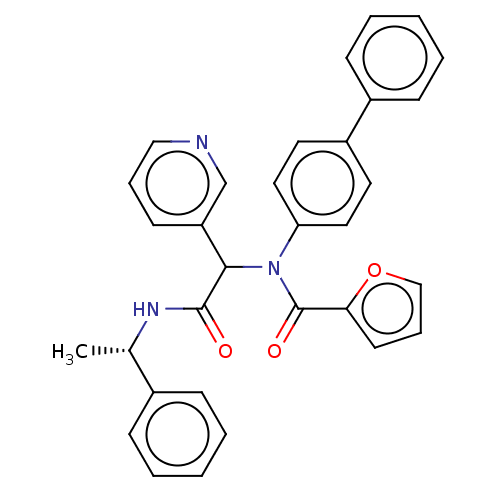
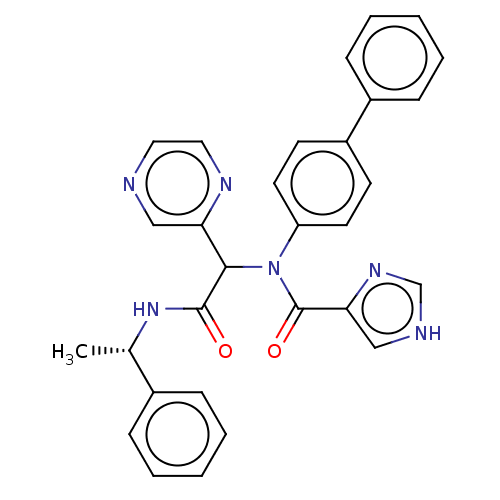
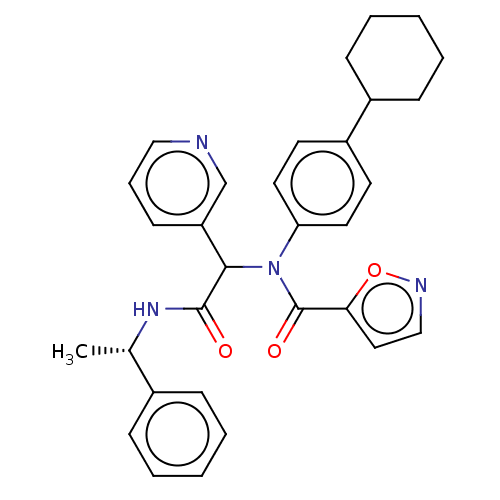
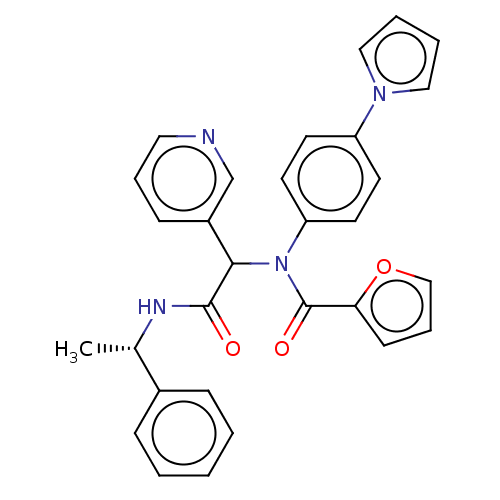
Affinity DataKi: 70nMAssay Description:Non-covalent inhibition to SARS-CoV-2 -Mpro assessed as inhibition constant by native mass spectrometric analysisMore data for this Ligand-Target Pair
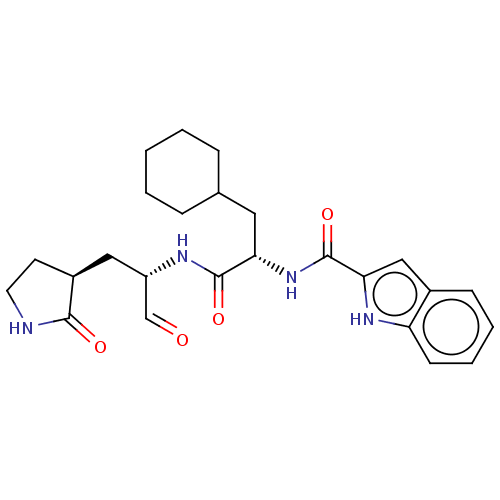
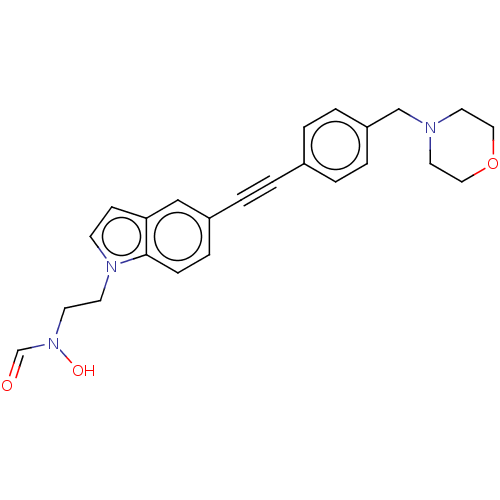
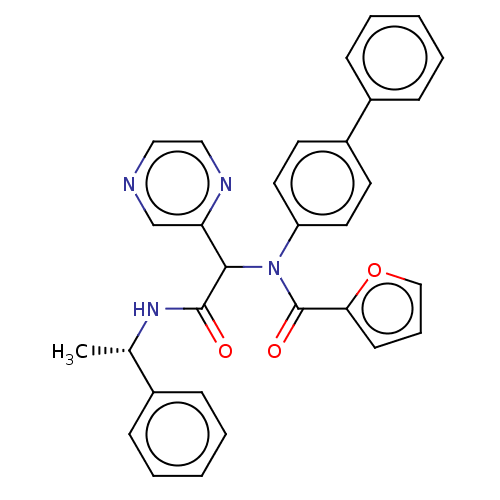
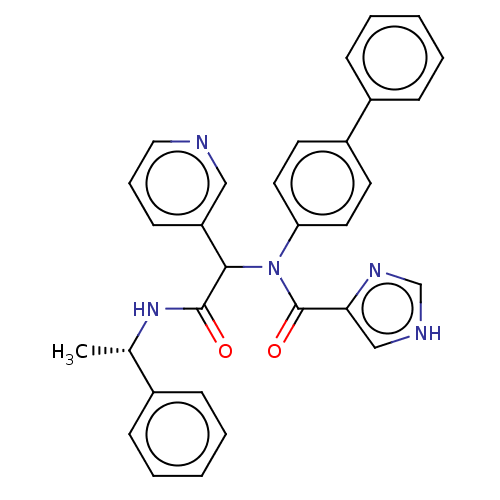
Affinity DataKi: 2.29E+3nMAssay Description:Non-covalent inhibition to SARS-CoV-2 -Mpro assessed as inhibition constant by native mass spectrometric analysisMore data for this Ligand-Target Pair
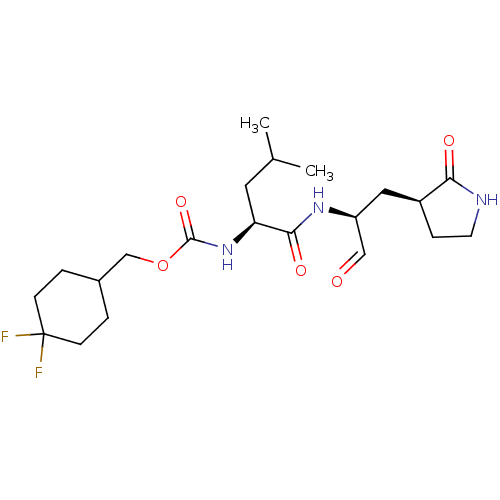
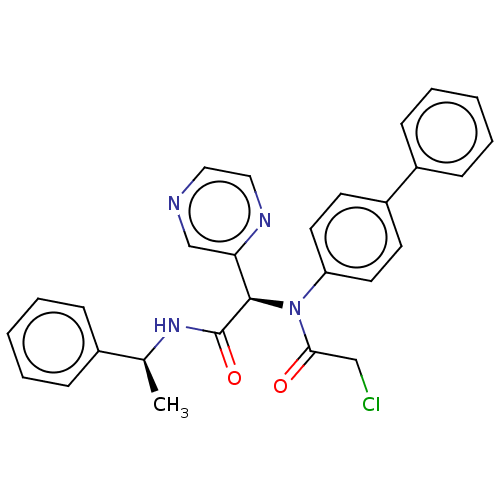
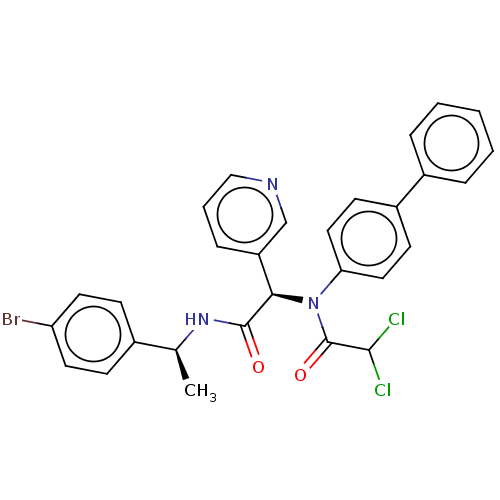
Affinity DataIC50: 0.210nMAssay Description:FRET-based enzymatic assay.More data for this Ligand-Target Pair
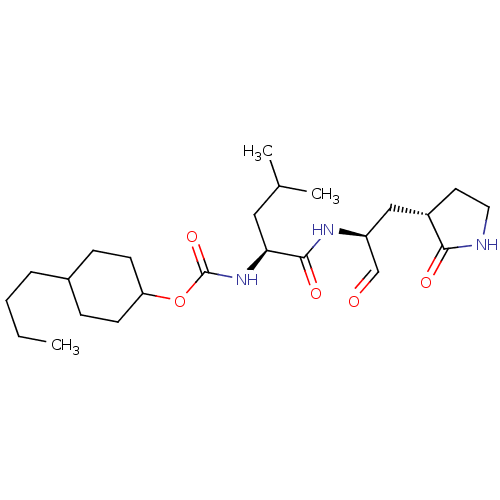
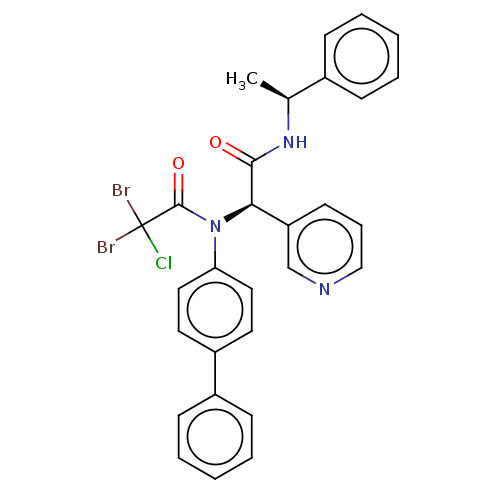
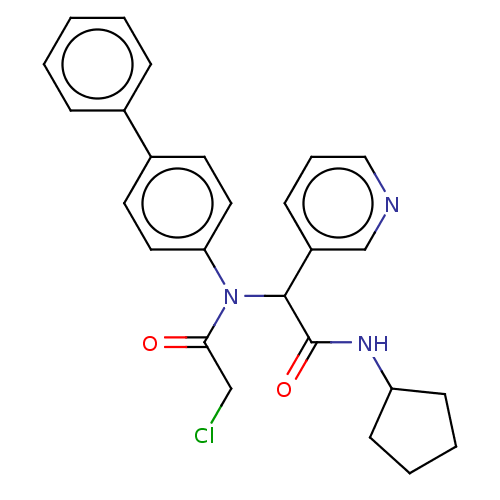
Affinity DataIC50: <0.5nMAssay Description:FRET-based enzymatic assay.More data for this Ligand-Target Pair
Affinity DataIC50: <0.5nMAssay Description:FRET-based enzymatic assay.More data for this Ligand-Target Pair
Affinity DataIC50: 0.560nMAssay Description:FRET-based enzymatic assay.More data for this Ligand-Target Pair
Affinity DataIC50: 0.990nMAssay Description:FRET-based enzymatic assay.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: <1nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.20nMAssay Description:FRET-based enzymatic assay.More data for this Ligand-Target Pair
Affinity DataIC50: 5nMAssay Description:FRET-based enzymatic assay.More data for this Ligand-Target Pair
Affinity DataIC50: 7nMAssay Description:FRET-based enzymatic assay.More data for this Ligand-Target Pair
Affinity DataIC50: 8nMAssay Description:FRET-based enzymatic assay.More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:FRET-based enzymatic assay.More data for this Ligand-Target Pair
Affinity DataIC50: 33nMAssay Description:FRET-based enzymatic assay.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 37nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 44nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 45nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
Affinity DataIC50: 50nMAssay Description:FRET-based enzymatic assay.More data for this Ligand-Target Pair
Affinity DataIC50: 50nMAssay Description:FRET-based enzymatic assay.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 63nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 64nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
Affinity DataIC50: 74nMAssay Description:FRET-based enzymatic assay.More data for this Ligand-Target Pair
Affinity DataIC50: 80nMAssay Description:FRET-based enzymatic assay.More data for this Ligand-Target Pair
Affinity DataIC50: 80nMAssay Description:FRET-based enzymatic assay.More data for this Ligand-Target Pair
Affinity DataIC50: 105nMAssay Description:FRET-based enzymatic assay.More data for this Ligand-Target Pair
Affinity DataIC50: 120nMAssay Description:FRET-based enzymatic assay.More data for this Ligand-Target Pair
Affinity DataIC50: 146nMAssay Description:FRET-based enzymatic assay.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 170nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
Affinity DataIC50: 170nMAssay Description:FRET-based enzymatic assay.More data for this Ligand-Target Pair
Affinity DataIC50: 180nMAssay Description:FRET-based enzymatic assay.More data for this Ligand-Target Pair
Affinity DataIC50: 200nMAssay Description:Non-covalent inhibition of SARS-Cov-2 -Mpro incubated for 30 mins using FRET substrate measured for 1 hr by FRET-based enzymatic assayMore data for this Ligand-Target Pair
Affinity DataIC50: 220nMAssay Description:FRET-based enzymatic assay.More data for this Ligand-Target Pair
Affinity DataIC50: 230nMAssay Description:FRET-based enzymatic assay.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 280nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
Affinity DataIC50: 280nMAssay Description:Non-covalent inhibition of SARS-Cov-2 -Mpro incubated for 30 mins using FRET substrate measured for 1 hr by FRET-based enzymatic assayMore data for this Ligand-Target Pair
Affinity DataIC50: 300nMAssay Description:FRET-based enzymatic assay.More data for this Ligand-Target Pair
Affinity DataIC50: 300nMAssay Description:FRET-based enzymatic assay.More data for this Ligand-Target Pair
Affinity DataIC50: 330nMAssay Description:FRET-based enzymatic assay.More data for this Ligand-Target Pair
Affinity DataIC50: 430nMAssay Description:FRET-based enzymatic assay.More data for this Ligand-Target Pair
Affinity DataIC50: 460nMAssay Description:FRET-based enzymatic assay.More data for this Ligand-Target Pair
Affinity DataIC50: 510nMAssay Description:Non-covalent inhibition of SARS-Cov-2 -Mpro incubated for 30 mins using FRET substrate measured for 1 hr by FRET-based enzymatic assayMore data for this Ligand-Target Pair
Affinity DataIC50: 560nMAssay Description:Non-covalent inhibition of SARS-Cov-2 -Mpro incubated for 30 mins using FRET substrate measured for 1 hr by FRET-based enzymatic assayMore data for this Ligand-Target Pair
Affinity DataIC50: 660nMAssay Description:Non-covalent inhibition of SARS-Cov-2 -Mpro incubated for 30 mins using FRET substrate measured for 1 hr by FRET-based enzymatic assayMore data for this Ligand-Target Pair
Affinity DataIC50: 670nMAssay Description:Non-covalent inhibition of SARS-Cov-2 -Mpro incubated for 30 mins using FRET substrate measured for 1 hr by FRET-based enzymatic assayMore data for this Ligand-Target Pair
Affinity DataIC50: 810nMAssay Description:Non-covalent inhibition of SARS-Cov-2 -Mpro incubated for 30 mins using FRET substrate measured for 1 hr by FRET-based enzymatic assayMore data for this Ligand-Target Pair
Affinity DataIC50: 940nMAssay Description:Non-covalent inhibition of SARS-Cov-2 -Mpro incubated for 30 mins using FRET substrate measured for 1 hr by FRET-based enzymatic assayMore data for this Ligand-Target Pair
Affinity DataIC50: 960nMAssay Description:Non-covalent inhibition of SARS-Cov-2 -Mpro incubated for 30 mins using FRET substrate measured for 1 hr by FRET-based enzymatic assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 1.00E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: <1.00E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: <1.00E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair

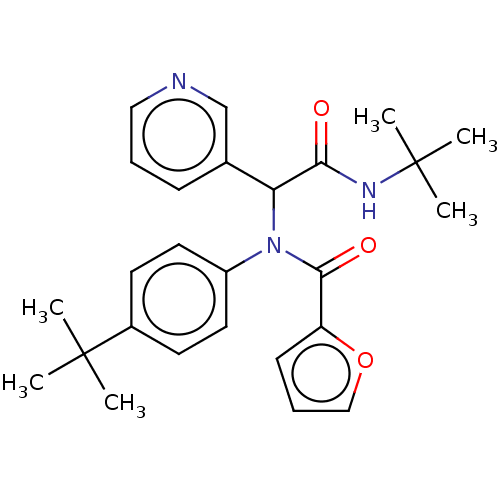
 3D Structure (crystal)
3D Structure (crystal)