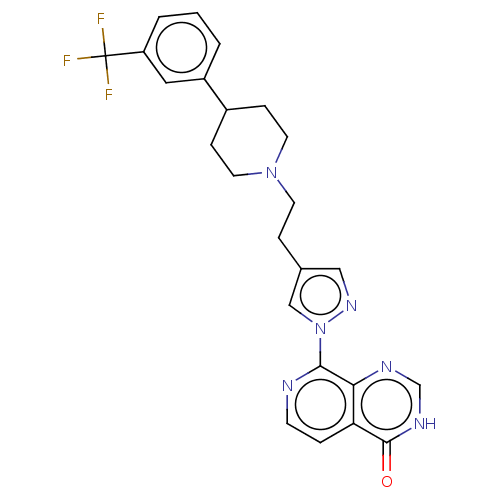
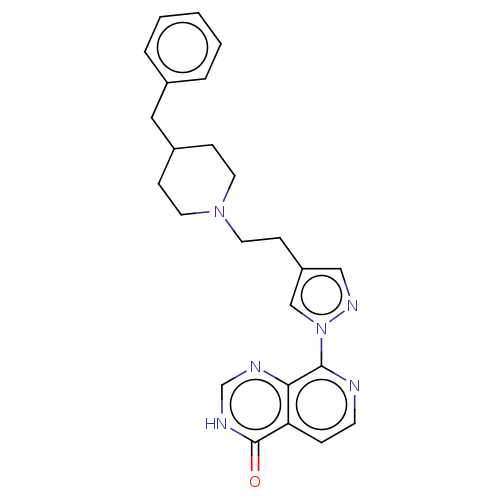
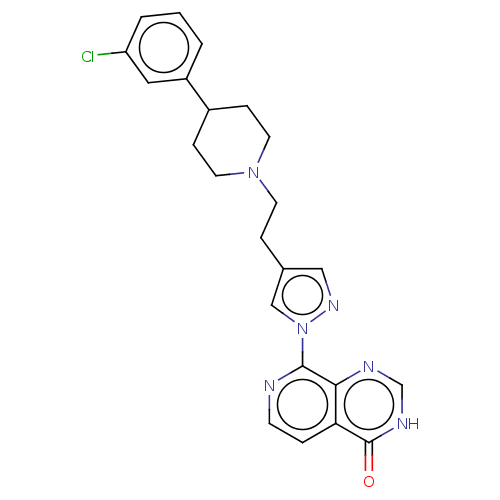
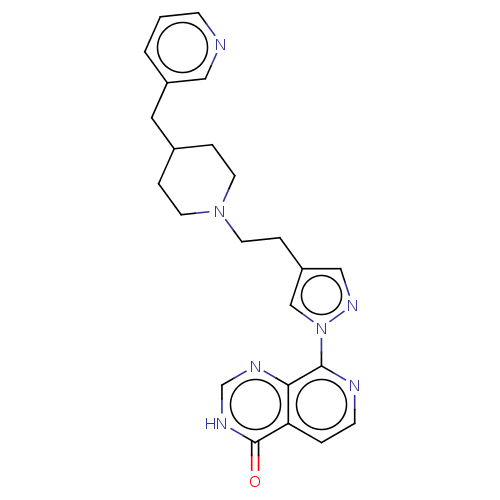
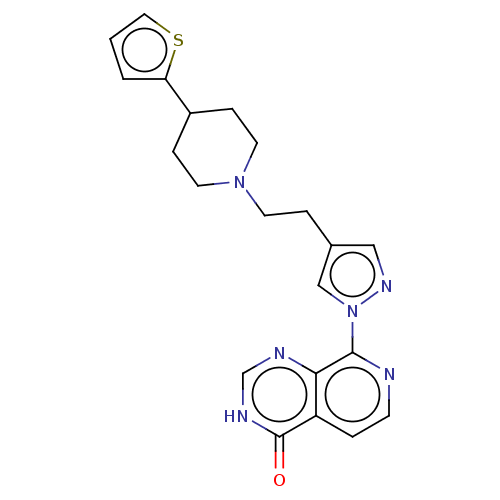
Affinity DataKi: 21nM ΔG°: -43.8kJ/molepH: 7.5 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA catalysed CO2 hydration activity. Phenol red (at a concentration of...More data for this Ligand-Target Pair
Affinity DataKi: 23nM ΔG°: -43.6kJ/molepH: 7.5 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA catalysed CO2 hydration activity. Phenol red (at a concentration of...More data for this Ligand-Target Pair
Affinity DataKi: 26nM ΔG°: -43.3kJ/molepH: 7.5 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA catalysed CO2 hydration activity. Phenol red (at a concentration of...More data for this Ligand-Target Pair
Affinity DataKi: 38nM ΔG°: -42.4kJ/molepH: 7.5 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA catalysed CO2 hydration activity. Phenol red (at a concentration of...More data for this Ligand-Target Pair
Affinity DataKi: 50nM ΔG°: -41.7kJ/molepH: 7.5 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA catalysed CO2 hydration activity. Phenol red (at a concentration of...More data for this Ligand-Target Pair
Affinity DataKi: 50nM ΔG°: -41.7kJ/molepH: 7.5 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA catalysed CO2 hydration activity. Phenol red (at a concentration of...More data for this Ligand-Target Pair
Affinity DataKi: 79nM ΔG°: -40.5kJ/molepH: 7.5 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA catalysed CO2 hydration activity. Phenol red (at a concentration of...More data for this Ligand-Target Pair
Affinity DataKi: 340nM ΔG°: -36.9kJ/molepH: 7.5 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA catalysed CO2 hydration activity. Phenol red (at a concentration of...More data for this Ligand-Target Pair
Affinity DataKi: 510nM ΔG°: -35.9kJ/molepH: 7.5 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA catalysed CO2 hydration activity. Phenol red (at a concentration of...More data for this Ligand-Target Pair
Affinity DataKi: 630nM ΔG°: -35.4kJ/molepH: 7.5 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA catalysed CO2 hydration activity. Phenol red (at a concentration of...More data for this Ligand-Target Pair
Affinity DataKi: 690nM ΔG°: -35.2kJ/molepH: 7.5 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA catalysed CO2 hydration activity. Phenol red (at a concentration of...More data for this Ligand-Target Pair
Affinity DataKi: 740nM ΔG°: -35.0kJ/molepH: 7.5 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA catalysed CO2 hydration activity. Phenol red (at a concentration of...More data for this Ligand-Target Pair
Affinity DataKi: 760nM ΔG°: -34.9kJ/molepH: 7.5 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA catalysed CO2 hydration activity. Phenol red (at a concentration of...More data for this Ligand-Target Pair
Affinity DataKi: 860nM ΔG°: -34.6kJ/molepH: 7.5 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA catalysed CO2 hydration activity. Phenol red (at a concentration of...More data for this Ligand-Target Pair
Affinity DataKi: 980nM ΔG°: -34.3kJ/molepH: 7.5 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA catalysed CO2 hydration activity. Phenol red (at a concentration of...More data for this Ligand-Target Pair
Affinity DataKi: 1.14E+3nM ΔG°: -33.9kJ/molepH: 7.5 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA catalysed CO2 hydration activity. Phenol red (at a concentration of...More data for this Ligand-Target Pair
Affinity DataKi: 1.20E+3nM ΔG°: -33.8kJ/molepH: 7.5 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA catalysed CO2 hydration activity. Phenol red (at a concentration of...More data for this Ligand-Target Pair
Affinity DataKi: 1.26E+3nM ΔG°: -33.7kJ/molepH: 7.5 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA catalysed CO2 hydration activity. Phenol red (at a concentration of...More data for this Ligand-Target Pair
Affinity DataKi: 1.47E+3nM ΔG°: -33.3kJ/molepH: 7.5 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA catalysed CO2 hydration activity. Phenol red (at a concentration of...More data for this Ligand-Target Pair
Affinity DataKi: 2.15E+3nM ΔG°: -32.3kJ/molepH: 7.5 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA catalysed CO2 hydration activity. Phenol red (at a concentration of...More data for this Ligand-Target Pair
Affinity DataKi: 6.13E+3nM ΔG°: -29.8kJ/molepH: 7.5 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA catalysed CO2 hydration activity. Phenol red (at a concentration of...More data for this Ligand-Target Pair
Affinity DataKi: 8.01E+3nM ΔG°: -29.1kJ/molepH: 7.5 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA catalysed CO2 hydration activity. Phenol red (at a concentration of...More data for this Ligand-Target Pair
Affinity DataKi: 1.50E+4nM ΔG°: -27.5kJ/molepH: 7.5 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA catalysed CO2 hydration activity. Phenol red (at a concentration of...More data for this Ligand-Target Pair
Affinity DataKi: 6.80E+5nM ΔG°: -18.1kJ/molepH: 7.5 T: 2°CAssay Description:An Applied Photophysics stopped-flow instrument has been used for assaying the CA catalysed CO2 hydration activity. Phenol red (at a concentration of...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 5B(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 7nMAssay Description:Inhibition of KDM5B (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 5B(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Inhibition of KDM5B (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 5B(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Inhibition of KDM5B (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 5B(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 11nMAssay Description:Inhibition of KDM6B (unknown origin) using biotin-H3K27me3 (21 to 44 residues) as substrate preincubated for 15 mins followed by substrate addition m...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 5B(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 12nMAssay Description:Inhibition of KDM5B (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 5C(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 12nMAssay Description:Inhibition of KDM5C (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 5B(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 14nMAssay Description:Inhibition of KDM5B (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 5B(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 14nMAssay Description:Inhibition of KDM5B (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 5B(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 15nMAssay Description:Inhibition of KDM5B (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 5B(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 15nMAssay Description:Inhibition of KDM5B (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 5B(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 15nMAssay Description:Inhibition of KDM5B (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 5B(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 16nMAssay Description:Inhibition of KDM5B (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 5B(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 17nMAssay Description:Inhibition of KDM5B (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 4B(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 17nMAssay Description:Inhibition of recombinant human N-terminal GST-tagged KDM4B (1 to 500 residues) expressed in baculovirus infected sf9 cells using biotin-H3K9me3 as s...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 5B(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 18nMAssay Description:Inhibition of KDM6B (unknown origin) using biotin-H3K27me3 (21 to 44 residues) as substrate preincubated for 15 mins followed by substrate addition m...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 5B(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 18nMAssay Description:Inhibition of KDM5B (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 5C(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 21nMAssay Description:Inhibition of KDM5C (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 5C(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 21nMAssay Description:Inhibition of KDM5C (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 5B(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 21nMAssay Description:Inhibition of KDM5B (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 5C(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 22nMAssay Description:Inhibition of KDM5C (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 5B(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 23nMAssay Description:Inhibition of KDM6B (unknown origin) using biotin-H3K27me3 (21 to 44 residues) as substrate preincubated for 15 mins followed by substrate addition m...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 5C(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 23nMAssay Description:Inhibition of KDM5C (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 5C(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 23nMAssay Description:Inhibition of KDM5C (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 5C(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 24nMAssay Description:Inhibition of KDM5C (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 5B(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 26nMAssay Description:Inhibition of KDM5B (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 5B(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 26nMAssay Description:Inhibition of KDM5B (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair

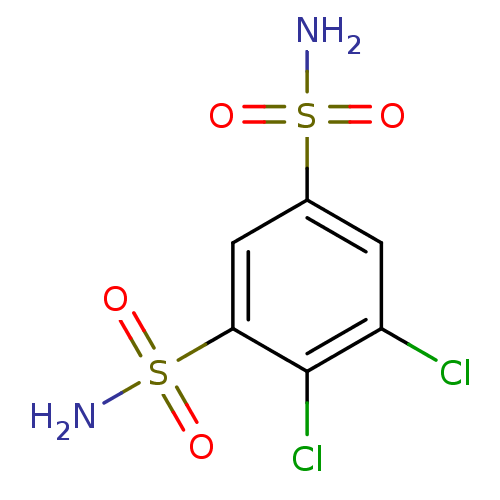
 3D Structure (crystal)
3D Structure (crystal)